research
Professor Jae Kyoung Kim and his research team from the Department of Mathematical Sciences at KAIST found that the circadian clock drives changes in circadian rhythms of p53 which functions as a tumor suppressor. Using a differential equation, he applied a model-driven mathematical approach to learn the mechanism and role of p53.
Kim’s mathematical modeling has been validated by experimental studies conducted by a research team at Virginia Polytechnic Institute and State University (Virginia Tech) in the United State, which is led by Professor Carla Finkielstein. As a result, the researchers revealed that there is an important link existed between the circadian clock and cancer.
The findings of this research were published online in Proceedings of the National Academy of Sciences of the United States of the America (PNAS) on November 9, 2016.
The circadian clock in our brain controls behavioral and physiological processes within a period of 24 hours, including making us fall asleep at a certain time by triggering the release of the sleep hormone melatonin in our brain, for example, around 9 pm. The clock is also involved in various physiological processes such as cell division, movement, and development.
Disruptions caused by the mismatch of the circadian clock and real time due to chronic late night work, shiftwork, and other similar issues may lead to various diseases such as diabetes, cancer, and heart disease.
In 2014, when Kim met with Finkielstein, her research team succeeded in observing the changes of p53 over a period of 24 hours, but could not understand how the circadian clock controls the 24-hour rhythm of p53. It was difficult to determine p53’s mechanism since its cell regulation system is far more complex than other cells
To solve the problem, Kim set up a computer simulation using mathematical modeling and ran millions of simulations. Instead of the traditional method based on trial and error experiments, mathematical modeling allowed to save a great deal of time, cost, and manpower.
During this process, Kim proved that the biorhythm of p53 and Period2, an important protein in the circadian clock, are closely related. Cells usually consist of a cell nucleus and cytoplasm. While p53 exists in both nucleus and cytoplasm, it becomes more stable and its degradation slows down when it is in the nucleus.
Kim predicted that the Period2 protein, which plays a key role in the functioning of the circadian clock, could influence the nucleus entry of the p53 protein.
Kim’s predictions based on mathematical modeling have been validated by the Virginia team, thereby revealing a strong connection between the circadian clock and cancer.
Researchers said that this research will help explain the cause of different results from numerous anticancer drugs, which are used to normalize the level of p53, when they are administrated at different times and find the most effective dosing times for the drugs.
They also believe that this study will play an important role in identifying the cause of increasing cancer rates in shift-workers whose circadian clocks are unstable and will contribute to the development of more effective treatments for cancer.
Professor Kim said, “This is an exciting thing that my research can contribute to improving the healthy lives of nurses, police officers, firefighters, and the like, who work in shifts against their circadian rhythms. Taking these findings as an opportunity, I hope to see more active interchanges of ideas between biological sciences and mathematical science in Korea.”
This research has been jointly conducted between KAIST and Virginia Tech and supported by the T. J. Park Science Fellowship of POSCO, the National Science Foundation of the United States, and the Young Researcher Program of the National Research Foundation of Korea.
Picture 1. The complex interaction between tumor antigen p53 and Period2 (Per2) which plays a major role in the circadian clock as revealed by mathematical simulations and experiments
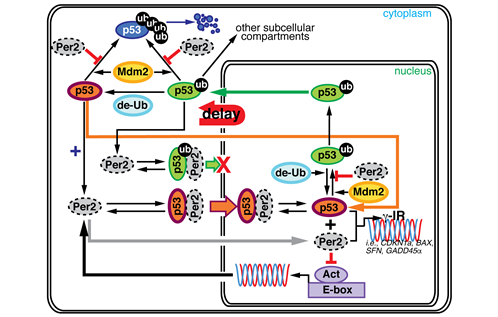
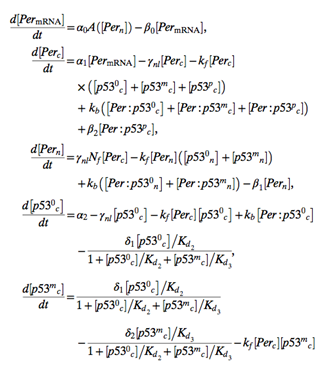
Picture 2. A portion of the mathematical model used in the research

Picture 3. Professor Jae Kyoung Kim (third from left) and the Virginia Tech Research Team

-
research KAIST Researchers Suggest an Extraordinary Alternative to Petroleum-based PET - Bacteria!
< (From left) Dr. Cindy Pricilia, Ph.D. Candidate Cheon Woo Moon, Distinguished Professor Sang Yup Lee > Currently, the world is suffering from environmental problems caused by plastic waste. The KAIST research team has succeeded in producing a microbial-based plastic that is biodegradable and can replace existing PET bottles, making it a hot topic. Our university announced on the 7th of November that the research team of Distinguished Professor Sang Yup Lee of the Department of Ch
2024-11-08 -
research X-ray Scattering Shines Light on Protein Folding
- Multiple forms of a non-functional, unfolded protein follow different pathways and timelines to reach its folded, functional state, a study reveals. - KAIST researchers have used an X-ray method to track how proteins fold, which could improve computer simulations of this process, with implications for understanding diseases and improving drug discovery. Their findings were reported in the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on June 30. Whe
2020-07-09 -
research A Novel Biosensor to Advance Diverse High-Level Production of Microbial Cell Factories
A research group at KAIST presented a novel biosensor which can produce diverse, high-level microbial cell factories. The biosensor monitors the concentration of products and even intermediates when new strains are being developed. This strategy provides a new platform for manufacturing diverse natural products from renewable resources. The team succeeded in creating four natural products of high-level pharmaceutical importance with this strategy. Malonyl-CoA is a major building block for m
2018-10-11 -
research Recombinant E. Coli As a Biofactory for the Biosynthesis of Diverse Nanomaterials
(Distinguished Professor Lee and PhD candidate Choi) A metabolic research group at KAIST and Chung-Ang University in Korea has developed a recombinant E. coli strain that biosynthesizes 60 different nanomaterials covering 35 elements on the periodic table. Among the elements, the team could biosynthesize 33 novel nanomaterials for the first time, advancing the forward design of nanomaterials through the biosynthesis of various single and multi-elements. The study analy
2018-05-23 -
research Deep Learning Predicts Drug-Drug and Drug-Food Interactions
A Korean research team from KAIST developed a computational framework, DeepDDI, that accurately predicts and generates 86 types of drug-drug and drug-food interactions as outputs of human-readable sentences, which allows in-depth understanding of the drug-drug and drug-food interactions. Drug interactions, including drug-drug interactions (DDIs) and drug-food constituent interactions (DFIs), can trigger unexpected pharmacological effects, including adverse drug events (ADEs), wi
2018-04-18