NDA
-
 Highly Efficient and Stable Double Layer Solar Cell Developed
Solar cells convert light into energy, but they can be inefficient and vulnerable to the environment, degrading with, ironically, too much light or other factors, including moisture and low temperature. An international research team has developed a new type of solar cell that can both withstand environmental hazards and is 26.7% efficient in power conversion.
They published their results on March 26 in Science.
The researchers, led by Byungha Shin, a professor from the Department of Materials Science and Engineering at KAIST, focused on developing a new class of light-absorbing material, called a wide bandgap perovskite. The material has a highly effective crystal structure that can process the power needs, but it can become problematic when exposed to environmental hazards, such as moisture. Researchers have made some progress increasing the efficiency of solar cells based on perovskite, but the material has greater potential than what was previously achieved.
To achieve better performance, Shin and his team built a double layer solar cell, called tandem, in which two or more light absorbers are stacked together to better utilize solar energy. To use perovskite in these tandem devices, the scientists modified the material’s optical property, which allows it to absorb a wider range of solar energy. Without the adjustment, the material is not as useful in achieving high performing tandem solar cells. The modification of the optical property of perovskite, however, comes with a penalty — the material becomes hugely vulnerable to the environment, in particular, to light.
To counteract the wide bandgap perovskite’s delicate nature, the researchers engineered combinations of molecules composing a two-dimensional layer in the perovskite, stabilizing the solar cells.
“We developed a high-quality wide bandgap perovskite material and, in combination with silicon solar cells, achieved world-class perovskite-silicon tandem cells,” Shin said.
The development was only possible due to the engineering method, in which the mixing ratio of the molecules building the two-dimensional layer are carefully controlled. In this case, the perovskite material not only improved efficiency of the resulting solar cell but also gained durability, retaining 80% of its initial power conversion capability even after 1,000 hours of continuous illumination. This is the first time such a high efficiency has been achieved with a wide bandgap perovskite single layer alone, according to Shin.
“Such high-efficiency wide bandgap perovskite is an essential technology for achieving ultra-high efficiency of perovskite-silicon tandem (double layer) solar cells,” Shin said. “The results also show the importance of bandgap matching of upper and lower cells in these tandem solar cells.”
The researchers, having stabilized the wide bandgap perovskite material, are now focused on developing even more efficient tandem solar cells that are expected to have more than 30% of power conversion efficiency, something that no one has achieved yet,
“Our ultimate goal is to develop ultra-high-efficiency tandem solar cells that contribute to the increase of shared solar energy among all energy sources,” Shin said. “We want to contribute to making the planet healthier.”
This work was supported by the National Research Foundation of Korea, the Korea Institute of Energy Technology Evaluation and Planning, the Ministry of Trade Industry and Energy of Korea, and the U.S. Department of Energy.
Other contributors include Daehan Kim, Jekyung Kim, Passarut Boonmongkolras, Seong Ryul Pae and Minkyu Kim, all of whom affiliated with the Department of Materials Science and Engineering at KAIST. Other authors include Byron W. Larson, Sean P. Dunfield, Chuanxiao Xiao, Jinhui Tong, Fei Zhang, Joseph J. Berry, Kai Zhu and Dong Hoe Kim, all of who are affiliated with the National Renewable Energy Laboratory in Colorado. Dunfield is also affiliated with the Materials Science and Engineering Program at the University of Colorado; Berry is also affiliated with the Department of Physics and the Renewable and Sustainable Energy Institute at the University of Colorado Boulder; and Kim is also affiliated with the Department of Nanotechnology and Advanced Materials Engineering at Sejong University. Hee Joon Jung and Vinayak Dravid of the Department of Materials Science and Engineering at Northwestern University; Ik Jae Park, Su Geun Ji and Jin Young Kim of the Department of Materials Science and Engineering at Seoul National University; and Seok Beom Kang of the Department of Nanotechnology and Advanced Materials Engineering of Sejong University also contributed.
Image credit: Professor Byungha Shin, KAIST
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication: Kim et al. (2020) “Efficient, stable silicon tandem cells enabled by anion-engineered wide band gap perovskites”. Science. Available online at https://doi.org/10.1126/science.aba3433
Profile:
Byungha Shin
Professor
byungha@kaist.ac.kr
http://energymatlab.kaist.ac.kr/
Department of Materials Science and Engineering
KAIST
Profile:
Daehan Kim
Ph.D. Candidate
zxzx4592@kaist.ac.kr
http://energymatlab.kaist.ac.kr/
Department of Materials Science and Engineering
KAIST
(END)
2020.03.27 View 22904
Highly Efficient and Stable Double Layer Solar Cell Developed
Solar cells convert light into energy, but they can be inefficient and vulnerable to the environment, degrading with, ironically, too much light or other factors, including moisture and low temperature. An international research team has developed a new type of solar cell that can both withstand environmental hazards and is 26.7% efficient in power conversion.
They published their results on March 26 in Science.
The researchers, led by Byungha Shin, a professor from the Department of Materials Science and Engineering at KAIST, focused on developing a new class of light-absorbing material, called a wide bandgap perovskite. The material has a highly effective crystal structure that can process the power needs, but it can become problematic when exposed to environmental hazards, such as moisture. Researchers have made some progress increasing the efficiency of solar cells based on perovskite, but the material has greater potential than what was previously achieved.
To achieve better performance, Shin and his team built a double layer solar cell, called tandem, in which two or more light absorbers are stacked together to better utilize solar energy. To use perovskite in these tandem devices, the scientists modified the material’s optical property, which allows it to absorb a wider range of solar energy. Without the adjustment, the material is not as useful in achieving high performing tandem solar cells. The modification of the optical property of perovskite, however, comes with a penalty — the material becomes hugely vulnerable to the environment, in particular, to light.
To counteract the wide bandgap perovskite’s delicate nature, the researchers engineered combinations of molecules composing a two-dimensional layer in the perovskite, stabilizing the solar cells.
“We developed a high-quality wide bandgap perovskite material and, in combination with silicon solar cells, achieved world-class perovskite-silicon tandem cells,” Shin said.
The development was only possible due to the engineering method, in which the mixing ratio of the molecules building the two-dimensional layer are carefully controlled. In this case, the perovskite material not only improved efficiency of the resulting solar cell but also gained durability, retaining 80% of its initial power conversion capability even after 1,000 hours of continuous illumination. This is the first time such a high efficiency has been achieved with a wide bandgap perovskite single layer alone, according to Shin.
“Such high-efficiency wide bandgap perovskite is an essential technology for achieving ultra-high efficiency of perovskite-silicon tandem (double layer) solar cells,” Shin said. “The results also show the importance of bandgap matching of upper and lower cells in these tandem solar cells.”
The researchers, having stabilized the wide bandgap perovskite material, are now focused on developing even more efficient tandem solar cells that are expected to have more than 30% of power conversion efficiency, something that no one has achieved yet,
“Our ultimate goal is to develop ultra-high-efficiency tandem solar cells that contribute to the increase of shared solar energy among all energy sources,” Shin said. “We want to contribute to making the planet healthier.”
This work was supported by the National Research Foundation of Korea, the Korea Institute of Energy Technology Evaluation and Planning, the Ministry of Trade Industry and Energy of Korea, and the U.S. Department of Energy.
Other contributors include Daehan Kim, Jekyung Kim, Passarut Boonmongkolras, Seong Ryul Pae and Minkyu Kim, all of whom affiliated with the Department of Materials Science and Engineering at KAIST. Other authors include Byron W. Larson, Sean P. Dunfield, Chuanxiao Xiao, Jinhui Tong, Fei Zhang, Joseph J. Berry, Kai Zhu and Dong Hoe Kim, all of who are affiliated with the National Renewable Energy Laboratory in Colorado. Dunfield is also affiliated with the Materials Science and Engineering Program at the University of Colorado; Berry is also affiliated with the Department of Physics and the Renewable and Sustainable Energy Institute at the University of Colorado Boulder; and Kim is also affiliated with the Department of Nanotechnology and Advanced Materials Engineering at Sejong University. Hee Joon Jung and Vinayak Dravid of the Department of Materials Science and Engineering at Northwestern University; Ik Jae Park, Su Geun Ji and Jin Young Kim of the Department of Materials Science and Engineering at Seoul National University; and Seok Beom Kang of the Department of Nanotechnology and Advanced Materials Engineering of Sejong University also contributed.
Image credit: Professor Byungha Shin, KAIST
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication: Kim et al. (2020) “Efficient, stable silicon tandem cells enabled by anion-engineered wide band gap perovskites”. Science. Available online at https://doi.org/10.1126/science.aba3433
Profile:
Byungha Shin
Professor
byungha@kaist.ac.kr
http://energymatlab.kaist.ac.kr/
Department of Materials Science and Engineering
KAIST
Profile:
Daehan Kim
Ph.D. Candidate
zxzx4592@kaist.ac.kr
http://energymatlab.kaist.ac.kr/
Department of Materials Science and Engineering
KAIST
(END)
2020.03.27 View 22904 -
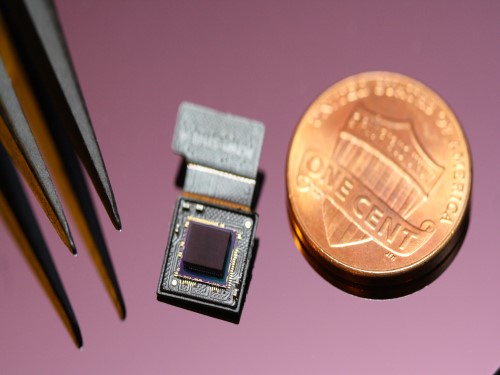 Ultrathin but Fully Packaged High-Resolution Camera
- Biologically inspired ultrathin arrayed camera captures super-resolution images. -
The unique structures of biological vision systems in nature inspired scientists to design ultracompact imaging systems. A research group led by Professor Ki-Hun Jeong have made an ultracompact camera that captures high-contrast and high-resolution images. Fully packaged with micro-optical elements such as inverted micro-lenses, multilayered pinhole arrays, and gap spacers on the image sensor, the camera boasts a total track length of 740 μm and a field of view of 73°.
Inspired by the eye structures of the paper wasp species Xenos peckii, the research team completely suppressed optical noise between micro-lenses while reducing camera thickness. The camera has successfully demonstrated high-contrast clear array images acquired from tiny micro lenses. To further enhance the image quality of the captured image, the team combined the arrayed images into one image through super-resolution imaging.
An insect’s compound eye has superior visual characteristics, such as a wide viewing angle, high motion sensitivity, and a large depth of field while maintaining a small volume of visual structure with a small focal length. Among them, the eyes of Xenos peckii and an endoparasite found on paper wasps have hundreds of photoreceptors in a single lens unlike conventional compound eyes. In particular, the eye structures of an adult Xenos peckii exhibit hundreds of photoreceptors on an individual eyelet and offer engineering inspiration for ultrathin cameras or imaging applications because they have higher visual acuity than other compound eyes.
For instance, Xenos peckii’s eye-inspired cameras provide a 50 times higher spatial resolution than those based on arthropod eyes. In addition, the effective image resolution of the Xenos peckii’s eye can be further improved using the image overlaps between neighboring eyelets. This unique structure offers higher visual resolution than other insect eyes.
The team achieved high-contrast and super-resolution imaging through a novel arrayed design of micro-optical elements comprising multilayered aperture arrays and inverted micro-lens arrays directly stacked over an image sensor. This optical component was integrated with a complementary metal oxide semiconductor image sensor.
This is first demonstration of super-resolution imaging which acquires a single integrated image with high contrast and high resolving power reconstructed from high-contrast array images. It is expected that this ultrathin arrayed camera can be applied for further developing mobile devices, advanced surveillance vehicles, and endoscopes.
Professor Jeong said, “This research has led to technological advances in imaging technology. We will continue to strive to make significant impacts on multidisciplinary research projects in the fields of microtechnology and nanotechnology, seeking inspiration from natural photonic structures.”
This work was featured in Light Science & Applications last month and was supported by the National Research Foundation (NRF) of and the Ministry of Health and Welfare (MOHW) of Korea.
Image credit: Professor Ki-Hun Jeong, KAIST
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication:
Kisoo Kim, Kyung-Won Jang, Jae-Kwan Ryu, and Ki-Hun Jeong. (2020) “Biologically inspired ultrathin arrayed camera for high-contrast and high-resolution imaging”. Light Science & Applications. Volume 9. Article 28. Available online at https://doi.org/10.1038/s41377-020-0261-8
Profile:
Ki-Hun Jeong
Professor
kjeong@kaist.ac.kr
http://biophotonics.kaist.ac.kr/
Department of Bio and Brain Engineering
KAIST
Profile:
Kisoo Kim
Ph.D. Candidate
kisoo.kim1@kaist.ac.kr
http://biophotonics.kaist.ac.kr/
Department of Bio and Brain Engineering
KAIST
(END)
2020.03.23 View 20558
Ultrathin but Fully Packaged High-Resolution Camera
- Biologically inspired ultrathin arrayed camera captures super-resolution images. -
The unique structures of biological vision systems in nature inspired scientists to design ultracompact imaging systems. A research group led by Professor Ki-Hun Jeong have made an ultracompact camera that captures high-contrast and high-resolution images. Fully packaged with micro-optical elements such as inverted micro-lenses, multilayered pinhole arrays, and gap spacers on the image sensor, the camera boasts a total track length of 740 μm and a field of view of 73°.
Inspired by the eye structures of the paper wasp species Xenos peckii, the research team completely suppressed optical noise between micro-lenses while reducing camera thickness. The camera has successfully demonstrated high-contrast clear array images acquired from tiny micro lenses. To further enhance the image quality of the captured image, the team combined the arrayed images into one image through super-resolution imaging.
An insect’s compound eye has superior visual characteristics, such as a wide viewing angle, high motion sensitivity, and a large depth of field while maintaining a small volume of visual structure with a small focal length. Among them, the eyes of Xenos peckii and an endoparasite found on paper wasps have hundreds of photoreceptors in a single lens unlike conventional compound eyes. In particular, the eye structures of an adult Xenos peckii exhibit hundreds of photoreceptors on an individual eyelet and offer engineering inspiration for ultrathin cameras or imaging applications because they have higher visual acuity than other compound eyes.
For instance, Xenos peckii’s eye-inspired cameras provide a 50 times higher spatial resolution than those based on arthropod eyes. In addition, the effective image resolution of the Xenos peckii’s eye can be further improved using the image overlaps between neighboring eyelets. This unique structure offers higher visual resolution than other insect eyes.
The team achieved high-contrast and super-resolution imaging through a novel arrayed design of micro-optical elements comprising multilayered aperture arrays and inverted micro-lens arrays directly stacked over an image sensor. This optical component was integrated with a complementary metal oxide semiconductor image sensor.
This is first demonstration of super-resolution imaging which acquires a single integrated image with high contrast and high resolving power reconstructed from high-contrast array images. It is expected that this ultrathin arrayed camera can be applied for further developing mobile devices, advanced surveillance vehicles, and endoscopes.
Professor Jeong said, “This research has led to technological advances in imaging technology. We will continue to strive to make significant impacts on multidisciplinary research projects in the fields of microtechnology and nanotechnology, seeking inspiration from natural photonic structures.”
This work was featured in Light Science & Applications last month and was supported by the National Research Foundation (NRF) of and the Ministry of Health and Welfare (MOHW) of Korea.
Image credit: Professor Ki-Hun Jeong, KAIST
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication:
Kisoo Kim, Kyung-Won Jang, Jae-Kwan Ryu, and Ki-Hun Jeong. (2020) “Biologically inspired ultrathin arrayed camera for high-contrast and high-resolution imaging”. Light Science & Applications. Volume 9. Article 28. Available online at https://doi.org/10.1038/s41377-020-0261-8
Profile:
Ki-Hun Jeong
Professor
kjeong@kaist.ac.kr
http://biophotonics.kaist.ac.kr/
Department of Bio and Brain Engineering
KAIST
Profile:
Kisoo Kim
Ph.D. Candidate
kisoo.kim1@kaist.ac.kr
http://biophotonics.kaist.ac.kr/
Department of Bio and Brain Engineering
KAIST
(END)
2020.03.23 View 20558 -
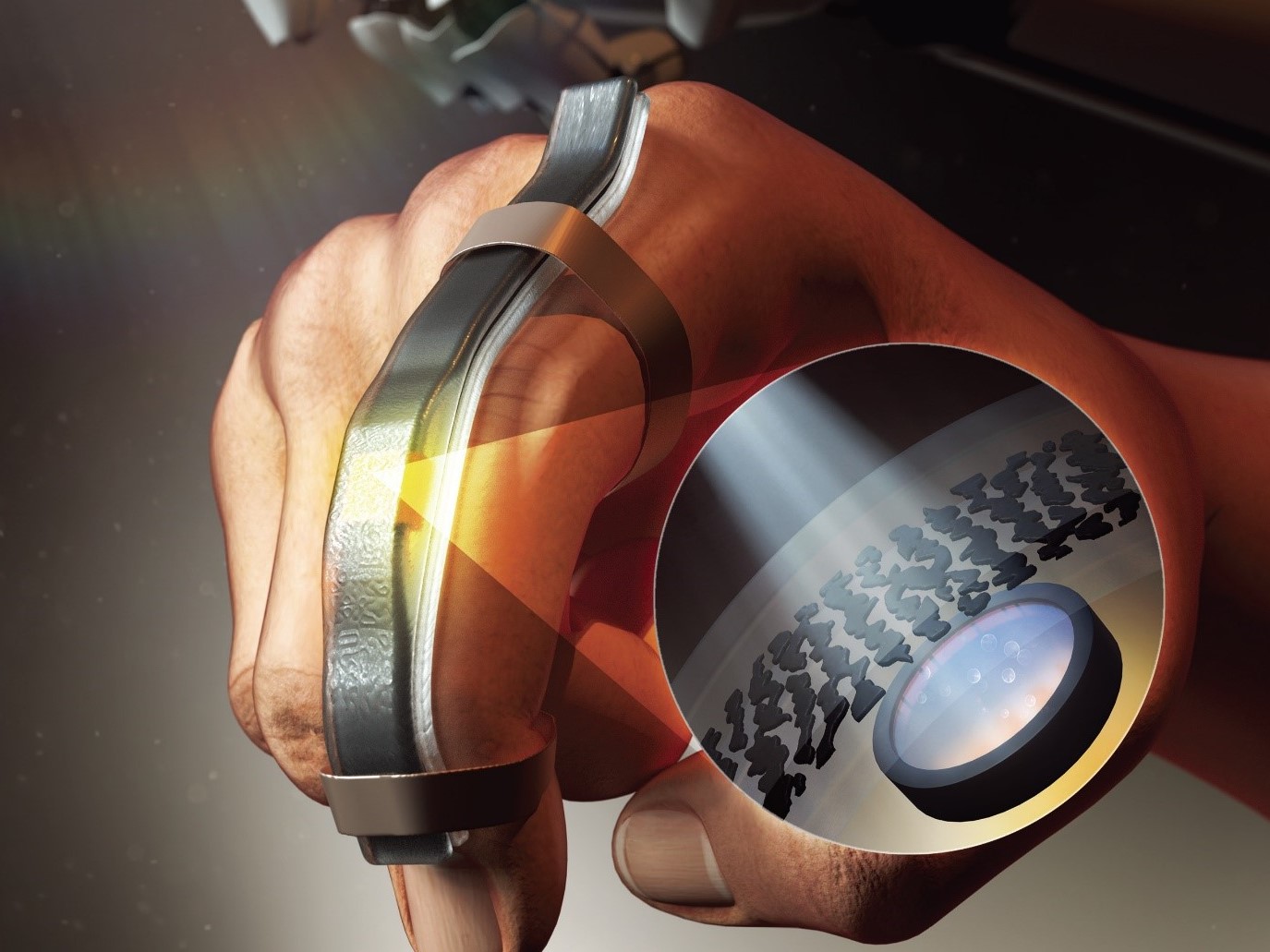 Wearable Strain Sensor Using Light Transmittance Helps Measure Physical Signals Better
KAIST researchers have developed a novel wearable strain sensor based on the modulation of optical transmittance of a carbon nanotube (CNT)-embedded elastomer. The sensor is capable of sensitive, stable, and continuous measurement of physical signals. This technology, featured in the March 4th issue of ACS Applied Materials & Interfaces as a front cover article, shows great potential for the detection of subtle human motions and the real-time monitoring of body postures for healthcare applications.
A wearable strain sensor must have high sensitivity, flexibility, and stretchability, as well as low cost. Those used especially for health monitoring should also be tied to long-term solid performance, and be environmentally stable. Various stretchable strain sensors based on piezo-resistive and capacitive principles have been developed to meet all these requirements.
Conventional piezo-resistive strain sensors using functional nanomaterials, including CNTs as the most common example, have shown high sensitivity and great sensing performance. However, they suffer from poor long-term stability and linearity, as well as considerable signal hysteresis. As an alternative, piezo-capacitive strain sensors with better stability, lower hysteresis, and higher stretchability have been suggested. But due to the fact that piezo-capacitive strain sensors exhibit limited sensitivity and strong electromagnetic interference caused by the conductive objects in the surrounding environment, these conventional stretchable strain sensors are still facing limitations that are yet to be resolved.
A KAIST research team led by Professor Inkyu Park from the Department of Mechanical Engineering suggested that an optical-type stretchable strain sensor can be a good alternative to resolve the limitations of conventional piezo-resistive and piezo-capacitive strain sensors, because they have high stability and are less affected by environmental disturbances. The team then introduced an optical wearable strain sensor based on the light transmittance changes of a CNT-embedded elastomer, which further addresses the low sensitivity problem of conventional optical stretchable strain sensors.
In order to achieve a large dynamic range for the sensor, Professor Park and his researchers chose Ecoflex as an elastomeric substrate with good mechanical durability, flexibility, and attachability on human skin, and the new optical wearable strain sensor developed by the research group actually shows a wide dynamic range of 0 to 400%.
In addition, the researchers propagated the microcracks under tensile strain within the film of multi-walled CNTs embedded in the Ecoflex substrate, changing the optical transmittance of the film. By doing so, it was possible for them to develop a wearable strain sensor having a sensitivity 10 times higher than conventional optical stretchable strain sensors.
The proposed sensor has also passed the durability test with excellent results. The sensor’s response after 13,000 sets of cyclic loading was stable without any noticeable drift. This suggests that the sensor response can be used without degradation, even if the sensor is repeatedly used for a long time and in various environmental conditions.
Using the developed sensor, the research team could measure the finger bending motion and used it for robot control. They also developed a three-axes sensor array for body posture monitoring. The sensor was able to monitor human motions with small strains such as a pulse near the carotid artery and muscle movement around the mouth during pronunciation.
Professor Park said, “In this study, our group developed a new wearable strain sensor platform that overcomes many limitations of previously developed resistive, capacitive, and optical-type stretchable strain sensors. Our sensor could be widely used in a variety of fields including soft robotics, wearable electronics, electronic skin, healthcare, and even entertainment.”
This work was supported by the National Research Foundation (NRF) of Korea.
Publication:
Jimin Gu, Donguk Kwon, Junseong Ahn, and Inkyu Park. (2020) “Wearable Strain sensors Using Light Transmittance Change of Carbon Nanotube-Embedded Elastomers with Microcracks” ACS Applied Materials & Interfaces. Volume 12. Issue 9. Available online at https://doi.org/10.1021/acsami.9b18069
Profile:
Inkyu Park
Professor
inkyu@kaist.ac.kr
http://mintlab1.kaist.ac.kr
Micro/Nano Transducers Laboratory (MINT Lab)
Department of Mechanical Engineering (ME)Korea Advanced Institute of Science and Technology (KAIST)
Profile:
Jimin Gu
Ph.D. Candidate
mint9411@kaist.ac.kr
http://mintlab1.kaist.ac.kr
MINT Lab
KAIST ME
(END)
2020.03.20 View 21412
Wearable Strain Sensor Using Light Transmittance Helps Measure Physical Signals Better
KAIST researchers have developed a novel wearable strain sensor based on the modulation of optical transmittance of a carbon nanotube (CNT)-embedded elastomer. The sensor is capable of sensitive, stable, and continuous measurement of physical signals. This technology, featured in the March 4th issue of ACS Applied Materials & Interfaces as a front cover article, shows great potential for the detection of subtle human motions and the real-time monitoring of body postures for healthcare applications.
A wearable strain sensor must have high sensitivity, flexibility, and stretchability, as well as low cost. Those used especially for health monitoring should also be tied to long-term solid performance, and be environmentally stable. Various stretchable strain sensors based on piezo-resistive and capacitive principles have been developed to meet all these requirements.
Conventional piezo-resistive strain sensors using functional nanomaterials, including CNTs as the most common example, have shown high sensitivity and great sensing performance. However, they suffer from poor long-term stability and linearity, as well as considerable signal hysteresis. As an alternative, piezo-capacitive strain sensors with better stability, lower hysteresis, and higher stretchability have been suggested. But due to the fact that piezo-capacitive strain sensors exhibit limited sensitivity and strong electromagnetic interference caused by the conductive objects in the surrounding environment, these conventional stretchable strain sensors are still facing limitations that are yet to be resolved.
A KAIST research team led by Professor Inkyu Park from the Department of Mechanical Engineering suggested that an optical-type stretchable strain sensor can be a good alternative to resolve the limitations of conventional piezo-resistive and piezo-capacitive strain sensors, because they have high stability and are less affected by environmental disturbances. The team then introduced an optical wearable strain sensor based on the light transmittance changes of a CNT-embedded elastomer, which further addresses the low sensitivity problem of conventional optical stretchable strain sensors.
In order to achieve a large dynamic range for the sensor, Professor Park and his researchers chose Ecoflex as an elastomeric substrate with good mechanical durability, flexibility, and attachability on human skin, and the new optical wearable strain sensor developed by the research group actually shows a wide dynamic range of 0 to 400%.
In addition, the researchers propagated the microcracks under tensile strain within the film of multi-walled CNTs embedded in the Ecoflex substrate, changing the optical transmittance of the film. By doing so, it was possible for them to develop a wearable strain sensor having a sensitivity 10 times higher than conventional optical stretchable strain sensors.
The proposed sensor has also passed the durability test with excellent results. The sensor’s response after 13,000 sets of cyclic loading was stable without any noticeable drift. This suggests that the sensor response can be used without degradation, even if the sensor is repeatedly used for a long time and in various environmental conditions.
Using the developed sensor, the research team could measure the finger bending motion and used it for robot control. They also developed a three-axes sensor array for body posture monitoring. The sensor was able to monitor human motions with small strains such as a pulse near the carotid artery and muscle movement around the mouth during pronunciation.
Professor Park said, “In this study, our group developed a new wearable strain sensor platform that overcomes many limitations of previously developed resistive, capacitive, and optical-type stretchable strain sensors. Our sensor could be widely used in a variety of fields including soft robotics, wearable electronics, electronic skin, healthcare, and even entertainment.”
This work was supported by the National Research Foundation (NRF) of Korea.
Publication:
Jimin Gu, Donguk Kwon, Junseong Ahn, and Inkyu Park. (2020) “Wearable Strain sensors Using Light Transmittance Change of Carbon Nanotube-Embedded Elastomers with Microcracks” ACS Applied Materials & Interfaces. Volume 12. Issue 9. Available online at https://doi.org/10.1021/acsami.9b18069
Profile:
Inkyu Park
Professor
inkyu@kaist.ac.kr
http://mintlab1.kaist.ac.kr
Micro/Nano Transducers Laboratory (MINT Lab)
Department of Mechanical Engineering (ME)Korea Advanced Institute of Science and Technology (KAIST)
Profile:
Jimin Gu
Ph.D. Candidate
mint9411@kaist.ac.kr
http://mintlab1.kaist.ac.kr
MINT Lab
KAIST ME
(END)
2020.03.20 View 21412 -
 3D Hierarchically Porous Nanostructured Catalyst Helps Efficiently Reduce CO2
- This new catalyst will bring CO2 one step closer to serving as a sustainable energy source. -
KAIST researchers developed a three-dimensional (3D) hierarchically porous nanostructured catalyst with carbon dioxide (CO2) to carbon monoxide (CO) conversion rate up to 3.96 times higher than that of conventional nanoporous gold catalysts. This new catalyst helps overcome the existing limitations of the mass transport that has been a major cause of decreases in the CO2 conversion rate, holding a strong promise for the large-scale and cost-effective electrochemical conversion of CO2 into useful chemicals.
As CO2 emissions increase and fossil fuels deplete globally, reducing and converting CO2 to clean energy electrochemically has attracted a great deal of attention as a promising technology. Especially due to the fact that the CO2 reduction reaction occurs competitively with hydrogen evolution reactions (HER) at similar redox potentials, the development of an efficient electrocatalyst for selective and robust CO2 reduction reactions has remained a key technological issue.
Gold (Au) is one of the most commonly used catalysts in CO2 reduction reactions, but the high cost and scarcity of Au pose obstacles for mass commercial applications. The development of nanostructures has been extensively studied as a potential approach to improving the selectivity for target products and maximizing the number of active stable sites, thus enhancing the energy efficiency.
However, the nanopores of the previously reported complex nanostructures were easily blocked by gaseous CO bubbles during aqueous reactions. The CO bubbles hindered mass transport of the reactants through the electrolyte, resulting in low CO2 conversion rates.
In the study published in the Proceedings of the National Academy of Sciences of the USA (PNAS) on March 4, a research group at KAIST led by Professor Seokwoo Jeon and Professor Jihun Oh from the Department of Materials Science and Engineering designed a 3D hierarchically porous Au nanostructure with two different sizes of macropores and nanopores. The team used proximity-field nanopatterning (PnP) and electroplating techniques that are effective for fabricating the 3D well-ordered nanostructures.
The proposed nanostructure, comprised of interconnected macroporous channels 200 to 300 nanometers (nm) wide and 10 nm nanopores, induces efficient mass transport through the interconnected macroporous channels as well as high selectivity by producing highly active stable sites from numerous nanopores.
As a result, its electrodes show a high CO selectivity of 85.8% at a low overpotential of 0.264 V and efficient mass activity that is up to 3.96 times higher than that of de-alloyed nanoporous Au electrodes.
“These results are expected to solve the problem of mass transfer in the field of similar electrochemical reactions and can be applied to a wide range of green energy applications for the efficient utilization of electrocatalysts,” said the researchers.
This work was supported by the National Research Foundation (NRF) of Korea.
Image credit: Professor Seokwoo Jeon and Professor Jihun Oh, KAIST
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication:
Hyun et al. (2020) Hierarchically porous Au nanostructures with interconnected channels for efficient mass transport in electrocatalytic CO2 reduction. Proceedings of the National Academy of Sciences of the USA (PNAS). Available online at https://doi.org/10.1073/pnas.1918837117
Profile:
Seokwoo Jeon, PhD
Professor
jeon39@kaist.ac.kr
http://fdml.kaist.ac.kr
Department of Materials Science and Engineering (MSE)
https://www.kaist.ac.kr
Korea Advanced Institute of Science and Technology (KAIST)Daejeon, Republic of Korea
Profile:
Jihun Oh, PhD
Associate Professor
jihun.oh@kaist.ac.kr
http://les.kaist.ac.kr
Department of Materials Science and Engineering (MSE)
Department of Energy, Environment, Water and Sustainability (EEWS)
KAIST
Profile:
Gayea Hyun
PhD Candidate
cldywkd93@kaist.ac.kr
http://fdml.kaist.ac.kr
Flexible Devices and Metamaterials Laboratory (FDML)
Department of Materials Science and Engineering (MSE)
KAIST
Profile:
Jun Tae Song, PhD
Assistant Professor
song.juntae@cstf.kyushu-u.ac.jp
http://www.cstf.kyushu-u.ac.jp/~ishihara-lab/
Department of Applied Chemistry
https://www.kyushu-u.ac.jp
Kyushu UniversityFukuoka, Japan
(END)
2020.03.13 View 19454
3D Hierarchically Porous Nanostructured Catalyst Helps Efficiently Reduce CO2
- This new catalyst will bring CO2 one step closer to serving as a sustainable energy source. -
KAIST researchers developed a three-dimensional (3D) hierarchically porous nanostructured catalyst with carbon dioxide (CO2) to carbon monoxide (CO) conversion rate up to 3.96 times higher than that of conventional nanoporous gold catalysts. This new catalyst helps overcome the existing limitations of the mass transport that has been a major cause of decreases in the CO2 conversion rate, holding a strong promise for the large-scale and cost-effective electrochemical conversion of CO2 into useful chemicals.
As CO2 emissions increase and fossil fuels deplete globally, reducing and converting CO2 to clean energy electrochemically has attracted a great deal of attention as a promising technology. Especially due to the fact that the CO2 reduction reaction occurs competitively with hydrogen evolution reactions (HER) at similar redox potentials, the development of an efficient electrocatalyst for selective and robust CO2 reduction reactions has remained a key technological issue.
Gold (Au) is one of the most commonly used catalysts in CO2 reduction reactions, but the high cost and scarcity of Au pose obstacles for mass commercial applications. The development of nanostructures has been extensively studied as a potential approach to improving the selectivity for target products and maximizing the number of active stable sites, thus enhancing the energy efficiency.
However, the nanopores of the previously reported complex nanostructures were easily blocked by gaseous CO bubbles during aqueous reactions. The CO bubbles hindered mass transport of the reactants through the electrolyte, resulting in low CO2 conversion rates.
In the study published in the Proceedings of the National Academy of Sciences of the USA (PNAS) on March 4, a research group at KAIST led by Professor Seokwoo Jeon and Professor Jihun Oh from the Department of Materials Science and Engineering designed a 3D hierarchically porous Au nanostructure with two different sizes of macropores and nanopores. The team used proximity-field nanopatterning (PnP) and electroplating techniques that are effective for fabricating the 3D well-ordered nanostructures.
The proposed nanostructure, comprised of interconnected macroporous channels 200 to 300 nanometers (nm) wide and 10 nm nanopores, induces efficient mass transport through the interconnected macroporous channels as well as high selectivity by producing highly active stable sites from numerous nanopores.
As a result, its electrodes show a high CO selectivity of 85.8% at a low overpotential of 0.264 V and efficient mass activity that is up to 3.96 times higher than that of de-alloyed nanoporous Au electrodes.
“These results are expected to solve the problem of mass transfer in the field of similar electrochemical reactions and can be applied to a wide range of green energy applications for the efficient utilization of electrocatalysts,” said the researchers.
This work was supported by the National Research Foundation (NRF) of Korea.
Image credit: Professor Seokwoo Jeon and Professor Jihun Oh, KAIST
Image usage restrictions: News organizations may use or redistribute this image, with proper attribution, as part of news coverage of this paper only.
Publication:
Hyun et al. (2020) Hierarchically porous Au nanostructures with interconnected channels for efficient mass transport in electrocatalytic CO2 reduction. Proceedings of the National Academy of Sciences of the USA (PNAS). Available online at https://doi.org/10.1073/pnas.1918837117
Profile:
Seokwoo Jeon, PhD
Professor
jeon39@kaist.ac.kr
http://fdml.kaist.ac.kr
Department of Materials Science and Engineering (MSE)
https://www.kaist.ac.kr
Korea Advanced Institute of Science and Technology (KAIST)Daejeon, Republic of Korea
Profile:
Jihun Oh, PhD
Associate Professor
jihun.oh@kaist.ac.kr
http://les.kaist.ac.kr
Department of Materials Science and Engineering (MSE)
Department of Energy, Environment, Water and Sustainability (EEWS)
KAIST
Profile:
Gayea Hyun
PhD Candidate
cldywkd93@kaist.ac.kr
http://fdml.kaist.ac.kr
Flexible Devices and Metamaterials Laboratory (FDML)
Department of Materials Science and Engineering (MSE)
KAIST
Profile:
Jun Tae Song, PhD
Assistant Professor
song.juntae@cstf.kyushu-u.ac.jp
http://www.cstf.kyushu-u.ac.jp/~ishihara-lab/
Department of Applied Chemistry
https://www.kyushu-u.ac.jp
Kyushu UniversityFukuoka, Japan
(END)
2020.03.13 View 19454 -
 A System Controlling Road Active Noise to Hit the Road
The research team led by Professor Youngjin Park of the Department of Mechanical Engineering has developed a road noise active noise control (RANC) system to be commercialized in partnership with Hyundai Motor Group.
On December 11, Hyundai Motor Group announced the successful development of the RANC system, which significantly reduces the road noise flowing into cars. The carmaker has completed the domestic and American patent applications for the location of sensors and the signal selection method, the core technology of RANC.
RANC is a technology for reducing road noise during driving. This system consists of an acceleration sensor, digital signal processor (the control computer to analyze sound signals), microphone, amplifier, and audio system. To make the system as simple as possible, the audio system utilizes the original audio system embedded in the car instead of a separate system.
The acceleration sensor first calculates the vibration from the road into the car. The location of the sensor is important for accurately identifying the vibration path. The research team was able to find the optimal sensor location through a number of tests.
The System Dynamics and Applied Control Laboratory of Professor Park researched ways to significantly reduce road noise with Hyundai Motor Group for four years from 1993 as a G7 national project and published the results in international journals. In 2002, the researchers published an article titled “Noise Quietens Driving” in Nature, where they announced the first success in reducing road noise in actual cars. The achievement did not lead to commercialization, however, due to the lack of auxiliary technologies at the time, digital amplifiers and DSP for cars for example, and pricing issues.
Since 2013, Professor Park’s research team has participated in one technology transfer and eight university-industry projects. Based on these efforts, the team was able to successfully develop the RANC system with domestic technology in partnership with Hyundai’s NVH Research Lab (Research Fellow, Dr. Gangdeok Lee; Ph.D. in aviation engineering, 1996), Optomech (Founder, Professor Gyeongsu Kim; Ph.D. in mechanical engineering, 1999), ARE (CEO Hyeonseok Kim; Ph.D. in mechanical engineering, 1998), WeAcom, and BurnYoung.
Professor Park’s team led the project by performing theory-based research during the commercialization stage in collaboration with Hyundai Motor Group.
For the commercialization of the RANC system, Hyundai Motor Group is planning to collaborate with the global car audio company Harman to increase the degree of completion and apply the RANC system to the GV 80, the first SUV model of the Genesis brand.
“I am very delighted as an engineer to see the research I worked on from my early days at KAIST be commercialized after 20 years,” noted Professor Park. “I am thrilled to make a contribution to such commercialization with my students in my lab.”
2019.12.27 View 13188
A System Controlling Road Active Noise to Hit the Road
The research team led by Professor Youngjin Park of the Department of Mechanical Engineering has developed a road noise active noise control (RANC) system to be commercialized in partnership with Hyundai Motor Group.
On December 11, Hyundai Motor Group announced the successful development of the RANC system, which significantly reduces the road noise flowing into cars. The carmaker has completed the domestic and American patent applications for the location of sensors and the signal selection method, the core technology of RANC.
RANC is a technology for reducing road noise during driving. This system consists of an acceleration sensor, digital signal processor (the control computer to analyze sound signals), microphone, amplifier, and audio system. To make the system as simple as possible, the audio system utilizes the original audio system embedded in the car instead of a separate system.
The acceleration sensor first calculates the vibration from the road into the car. The location of the sensor is important for accurately identifying the vibration path. The research team was able to find the optimal sensor location through a number of tests.
The System Dynamics and Applied Control Laboratory of Professor Park researched ways to significantly reduce road noise with Hyundai Motor Group for four years from 1993 as a G7 national project and published the results in international journals. In 2002, the researchers published an article titled “Noise Quietens Driving” in Nature, where they announced the first success in reducing road noise in actual cars. The achievement did not lead to commercialization, however, due to the lack of auxiliary technologies at the time, digital amplifiers and DSP for cars for example, and pricing issues.
Since 2013, Professor Park’s research team has participated in one technology transfer and eight university-industry projects. Based on these efforts, the team was able to successfully develop the RANC system with domestic technology in partnership with Hyundai’s NVH Research Lab (Research Fellow, Dr. Gangdeok Lee; Ph.D. in aviation engineering, 1996), Optomech (Founder, Professor Gyeongsu Kim; Ph.D. in mechanical engineering, 1999), ARE (CEO Hyeonseok Kim; Ph.D. in mechanical engineering, 1998), WeAcom, and BurnYoung.
Professor Park’s team led the project by performing theory-based research during the commercialization stage in collaboration with Hyundai Motor Group.
For the commercialization of the RANC system, Hyundai Motor Group is planning to collaborate with the global car audio company Harman to increase the degree of completion and apply the RANC system to the GV 80, the first SUV model of the Genesis brand.
“I am very delighted as an engineer to see the research I worked on from my early days at KAIST be commercialized after 20 years,” noted Professor Park. “I am thrilled to make a contribution to such commercialization with my students in my lab.”
2019.12.27 View 13188 -
 KAIST GSAI and SNUBH Join Hands for AI in Healthcare
< Dean Song Chong (left) and Director Chang Wan Oh (right)
at the KAIST GSAI - SNUBH MOU Signing Ceremony >
The Graduate School of AI (GSAI) at KAIST and the Seoul National University Bundang Hospital (SNUBH) signed a memorandum of understanding (MOU) to cooperate in AI education and research in the field of healthcare last month. The two institutions have agreed to collaborate on research and technology development through the implementation of academic and personnel exchange programs.
The GSAI, opened in August 2019 as Korea’s first AI graduate school, has been in the forefront of nurturing top-tier AI specialists in the era of Fourth Industrial Revolution. The school employs a two-track strategy that not only provides students with core AI-related courses on machine learning, data mining, computer vision, and natural language processing, but also a multidisciplinary curriculum incorporating the five key fields of healthcare, autonomous vehicles, manufacturing, security, and emerging technologies. Its faculty members are "the cream of the crop” in their early 40s, achieving world-class performance in their respective fields.
SNUBH opened the Healthcare Innovation Park in 2016, the first hospital-led convergence research complex among Korean medical institutions. It is leading future medical research in five specialized areas: medical devices, healthcare ICT, human genetics, nano-machines, and regenerative medicine.
The Dean of the GSAI, Song Chong, said, “We have set the stage for a cooperative platform for continuous and efficient joint education and research by the two institutions.” He expressed his excitement, saying, “Through this platform and our expertise in AI engineering and medicine, we will lead future AI-based medical technology.”
The Director of the SNUBH Research Division, Chang Wan Oh, stressed that “the mutual cooperation between the two institutions will become a crucial turning point in AI education and research, which is at the core of future healthcare.” He added, “Through a high level of cooperation, we will have the ability to bring about global competitiveness and innovation.”
(END)
2019.12.27 View 9768
KAIST GSAI and SNUBH Join Hands for AI in Healthcare
< Dean Song Chong (left) and Director Chang Wan Oh (right)
at the KAIST GSAI - SNUBH MOU Signing Ceremony >
The Graduate School of AI (GSAI) at KAIST and the Seoul National University Bundang Hospital (SNUBH) signed a memorandum of understanding (MOU) to cooperate in AI education and research in the field of healthcare last month. The two institutions have agreed to collaborate on research and technology development through the implementation of academic and personnel exchange programs.
The GSAI, opened in August 2019 as Korea’s first AI graduate school, has been in the forefront of nurturing top-tier AI specialists in the era of Fourth Industrial Revolution. The school employs a two-track strategy that not only provides students with core AI-related courses on machine learning, data mining, computer vision, and natural language processing, but also a multidisciplinary curriculum incorporating the five key fields of healthcare, autonomous vehicles, manufacturing, security, and emerging technologies. Its faculty members are "the cream of the crop” in their early 40s, achieving world-class performance in their respective fields.
SNUBH opened the Healthcare Innovation Park in 2016, the first hospital-led convergence research complex among Korean medical institutions. It is leading future medical research in five specialized areas: medical devices, healthcare ICT, human genetics, nano-machines, and regenerative medicine.
The Dean of the GSAI, Song Chong, said, “We have set the stage for a cooperative platform for continuous and efficient joint education and research by the two institutions.” He expressed his excitement, saying, “Through this platform and our expertise in AI engineering and medicine, we will lead future AI-based medical technology.”
The Director of the SNUBH Research Division, Chang Wan Oh, stressed that “the mutual cooperation between the two institutions will become a crucial turning point in AI education and research, which is at the core of future healthcare.” He added, “Through a high level of cooperation, we will have the ability to bring about global competitiveness and innovation.”
(END)
2019.12.27 View 9768 -
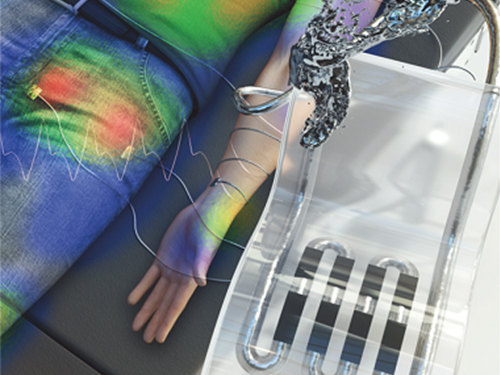 New Liquid Metal Wearable Pressure Sensor Created for Health Monitoring Applications
Soft pressure sensors have received significant research attention in a variety of fields, including soft robotics, electronic skin, and wearable electronics. Wearable soft pressure sensors have great potential for the real-time health monitoring and for the early diagnosis of diseases.
A KAIST research team led by Professor Inkyu Park from the Department of Mechanical Engineering developed a highly sensitive wearable pressure sensor for health monitoring applications. This work was reported in Advanced Healthcare Materials on November 21 as a front cover article.
This technology is capable of sensitive, precise, and continuous measurement of physiological and physical signals and shows great potential for health monitoring applications and the early diagnosis of diseases.
A soft pressure sensor is required to have high compliance, high sensitivity, low cost, long-term performance stability, and environmental stability in order to be employed for continuous health monitoring. Conventional solid-state soft pressure sensors using functional materials including carbon nanotubes and graphene have showed great sensing performance. However, these sensors suffer from limited stretchability, signal drifting, and long-term instability due to the distance between the stretchable substrate and the functional materials.
To overcome these issues, liquid-state electronics using liquid metal have been introduced for various wearable applications. Of these materials, Galinstan, a eutectic metal alloy of gallium, indium, and tin, has great mechanical and electrical properties that can be employed in wearable applications. But today’s liquid metal-based pressure sensors have low-pressure sensitivity, limiting their applicability for health monitoring devices.
The research team developed a 3D-printed rigid microbump array-integrated, liquid metal-based soft pressure sensor. With the help of 3D printing, the integration of a rigid microbump array and the master mold for a liquid metal microchannel could be achieved simultaneously, reducing the complexity of the manufacturing process. Through the integration of the rigid microbump and the microchannel, the new pressure sensor has an extremely low detection limit and enhanced pressure sensitivity compared to previously reported liquid metal-based pressure sensors. The proposed sensor also has a negligible signal drift over 10,000 cycles of pressure, bending, and stretching and exhibited excellent stability when subjected to various environmental conditions.
These performance outcomes make it an excellent sensor for various health monitoring devices. First, the research team demonstrated a wearable wristband device that can continuously monitor one’s pulse during exercise and be employed in a noninvasive cuffless BP monitoring system based on PTT calculations. Then, they introduced a wireless wearable heel pressure monitoring system that integrates three 3D-BLiPS with a wireless communication module.
Professor Park said, “It was possible to measure health indicators including pulse and blood pressure continuously as well as pressure of body parts using our proposed soft pressure sensor. We expect it to be used in health care applications, such as the prevention and the monitoring of the pressure-driven diseases such as pressure ulcers in the near future. There will be more opportunities for future research including a whole-body pressure monitoring system related to other physical parameters.”
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Ministry of Science and ICT.
< Figure 1. The front cover image of Advanced Healthcare Materials, Volume 8, Issue 22. >
< Figure 2. Highly sensitive liquid metal-based soft pressure sensor integrated with 3D-printed microbump array. >
< Figure 3. High pressure sensitivity and reliable sensing performances of the proposed sensor and wireless heel pressure monitoring application. >
-ProfileProfessor Inkyu ParkMicro/Nano Transducers Laboratoryhttp://mintlab1.kaist.ac.kr/
Department of Mechanical EngineeringKAIST
2019.12.20 View 16227
New Liquid Metal Wearable Pressure Sensor Created for Health Monitoring Applications
Soft pressure sensors have received significant research attention in a variety of fields, including soft robotics, electronic skin, and wearable electronics. Wearable soft pressure sensors have great potential for the real-time health monitoring and for the early diagnosis of diseases.
A KAIST research team led by Professor Inkyu Park from the Department of Mechanical Engineering developed a highly sensitive wearable pressure sensor for health monitoring applications. This work was reported in Advanced Healthcare Materials on November 21 as a front cover article.
This technology is capable of sensitive, precise, and continuous measurement of physiological and physical signals and shows great potential for health monitoring applications and the early diagnosis of diseases.
A soft pressure sensor is required to have high compliance, high sensitivity, low cost, long-term performance stability, and environmental stability in order to be employed for continuous health monitoring. Conventional solid-state soft pressure sensors using functional materials including carbon nanotubes and graphene have showed great sensing performance. However, these sensors suffer from limited stretchability, signal drifting, and long-term instability due to the distance between the stretchable substrate and the functional materials.
To overcome these issues, liquid-state electronics using liquid metal have been introduced for various wearable applications. Of these materials, Galinstan, a eutectic metal alloy of gallium, indium, and tin, has great mechanical and electrical properties that can be employed in wearable applications. But today’s liquid metal-based pressure sensors have low-pressure sensitivity, limiting their applicability for health monitoring devices.
The research team developed a 3D-printed rigid microbump array-integrated, liquid metal-based soft pressure sensor. With the help of 3D printing, the integration of a rigid microbump array and the master mold for a liquid metal microchannel could be achieved simultaneously, reducing the complexity of the manufacturing process. Through the integration of the rigid microbump and the microchannel, the new pressure sensor has an extremely low detection limit and enhanced pressure sensitivity compared to previously reported liquid metal-based pressure sensors. The proposed sensor also has a negligible signal drift over 10,000 cycles of pressure, bending, and stretching and exhibited excellent stability when subjected to various environmental conditions.
These performance outcomes make it an excellent sensor for various health monitoring devices. First, the research team demonstrated a wearable wristband device that can continuously monitor one’s pulse during exercise and be employed in a noninvasive cuffless BP monitoring system based on PTT calculations. Then, they introduced a wireless wearable heel pressure monitoring system that integrates three 3D-BLiPS with a wireless communication module.
Professor Park said, “It was possible to measure health indicators including pulse and blood pressure continuously as well as pressure of body parts using our proposed soft pressure sensor. We expect it to be used in health care applications, such as the prevention and the monitoring of the pressure-driven diseases such as pressure ulcers in the near future. There will be more opportunities for future research including a whole-body pressure monitoring system related to other physical parameters.”
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Ministry of Science and ICT.
< Figure 1. The front cover image of Advanced Healthcare Materials, Volume 8, Issue 22. >
< Figure 2. Highly sensitive liquid metal-based soft pressure sensor integrated with 3D-printed microbump array. >
< Figure 3. High pressure sensitivity and reliable sensing performances of the proposed sensor and wireless heel pressure monitoring application. >
-ProfileProfessor Inkyu ParkMicro/Nano Transducers Laboratoryhttp://mintlab1.kaist.ac.kr/
Department of Mechanical EngineeringKAIST
2019.12.20 View 16227 -
 AI to Determine When to Intervene with Your Driving
(Professor Uichin Lee (left) and PhD candidate Auk Kim)
Can your AI agent judge when to talk to you while you are driving? According to a KAIST research team, their in-vehicle conservation service technology will judge when it is appropriate to contact you to ensure your safety.
Professor Uichin Lee from the Department of Industrial and Systems Engineering at KAIST and his research team have developed AI technology that automatically detects safe moments for AI agents to provide conversation services to drivers.
Their research focuses on solving the potential problems of distraction created by in-vehicle conversation services. If an AI agent talks to a driver at an inopportune moment, such as while making a turn, a car accident will be more likely to occur.
In-vehicle conversation services need to be convenient as well as safe. However, the cognitive burden of multitasking negatively influences the quality of the service. Users tend to be more distracted during certain traffic conditions. To address this long-standing challenge of the in-vehicle conversation services, the team introduced a composite cognitive model that considers both safe driving and auditory-verbal service performance and used a machine-learning model for all collected data.
The combination of these individual measures is able to determine the appropriate moments for conversation and most appropriate types of conversational services. For instance, in the case of delivering simple-context information, such as a weather forecast, driver safety alone would be the most appropriate consideration. Meanwhile, when delivering information that requires a driver response, such as a “Yes” or “No,” the combination of driver safety and auditory-verbal performance should be considered.
The research team developed a prototype of an in-vehicle conversation service based on a navigation app that can be used in real driving environments. The app was also connected to the vehicle to collect in-vehicle OBD-II/CAN data, such as the steering wheel angle and brake pedal position, and mobility and environmental data such as the distance between successive cars and traffic flow.
Using pseudo-conversation services, the research team collected a real-world driving dataset consisting of 1,388 interactions and sensor data from 29 drivers who interacted with AI conversational agents. Machine learning analysis based on the dataset demonstrated that the opportune moments for driver interruption could be correctly inferred with 87% accuracy.
The safety enhancement technology developed by the team is expected to minimize driver distractions caused by in-vehicle conversation services. This technology can be directly applied to current in-vehicle systems that provide conversation services. It can also be extended and applied to the real-time detection of driver distraction problems caused by the use of a smartphone while driving.
Professor Lee said, “In the near future, cars will proactively deliver various in-vehicle conversation services. This technology will certainly help vehicles interact with their drivers safely as it can fairly accurately determine when to provide conversation services using only basic sensor data generated by cars.”
The researchers presented their findings at the ACM International Joint Conference on Pervasive and Ubiquitous Computing (Ubicomp’19) in London, UK. This research was supported in part by Hyundai NGV and by the Next-Generation Information Computing Development Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT.
(Figure: Visual description of safe enhancement technology for in-vehicle conversation services)
2019.11.13 View 19580
AI to Determine When to Intervene with Your Driving
(Professor Uichin Lee (left) and PhD candidate Auk Kim)
Can your AI agent judge when to talk to you while you are driving? According to a KAIST research team, their in-vehicle conservation service technology will judge when it is appropriate to contact you to ensure your safety.
Professor Uichin Lee from the Department of Industrial and Systems Engineering at KAIST and his research team have developed AI technology that automatically detects safe moments for AI agents to provide conversation services to drivers.
Their research focuses on solving the potential problems of distraction created by in-vehicle conversation services. If an AI agent talks to a driver at an inopportune moment, such as while making a turn, a car accident will be more likely to occur.
In-vehicle conversation services need to be convenient as well as safe. However, the cognitive burden of multitasking negatively influences the quality of the service. Users tend to be more distracted during certain traffic conditions. To address this long-standing challenge of the in-vehicle conversation services, the team introduced a composite cognitive model that considers both safe driving and auditory-verbal service performance and used a machine-learning model for all collected data.
The combination of these individual measures is able to determine the appropriate moments for conversation and most appropriate types of conversational services. For instance, in the case of delivering simple-context information, such as a weather forecast, driver safety alone would be the most appropriate consideration. Meanwhile, when delivering information that requires a driver response, such as a “Yes” or “No,” the combination of driver safety and auditory-verbal performance should be considered.
The research team developed a prototype of an in-vehicle conversation service based on a navigation app that can be used in real driving environments. The app was also connected to the vehicle to collect in-vehicle OBD-II/CAN data, such as the steering wheel angle and brake pedal position, and mobility and environmental data such as the distance between successive cars and traffic flow.
Using pseudo-conversation services, the research team collected a real-world driving dataset consisting of 1,388 interactions and sensor data from 29 drivers who interacted with AI conversational agents. Machine learning analysis based on the dataset demonstrated that the opportune moments for driver interruption could be correctly inferred with 87% accuracy.
The safety enhancement technology developed by the team is expected to minimize driver distractions caused by in-vehicle conversation services. This technology can be directly applied to current in-vehicle systems that provide conversation services. It can also be extended and applied to the real-time detection of driver distraction problems caused by the use of a smartphone while driving.
Professor Lee said, “In the near future, cars will proactively deliver various in-vehicle conversation services. This technology will certainly help vehicles interact with their drivers safely as it can fairly accurately determine when to provide conversation services using only basic sensor data generated by cars.”
The researchers presented their findings at the ACM International Joint Conference on Pervasive and Ubiquitous Computing (Ubicomp’19) in London, UK. This research was supported in part by Hyundai NGV and by the Next-Generation Information Computing Development Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT.
(Figure: Visual description of safe enhancement technology for in-vehicle conversation services)
2019.11.13 View 19580 -
 Ultrafast Quantum Motion in a Nanoscale Trap Detected
< Professor Heung-Sun Sim (left) and Co-author Dr. Sungguen Ryu (right) >
KAIST researchers have reported the detection of a picosecond electron motion in a silicon transistor. This study has presented a new protocol for measuring ultrafast electronic dynamics in an effective time-resolved fashion of picosecond resolution. The detection was made in collaboration with Nippon Telegraph and Telephone Corp. (NTT) in Japan and National Physical Laboratory (NPL) in the UK and is the first report to the best of our knowledge.
When an electron is captured in a nanoscale trap in solids, its quantum mechanical wave function can exhibit spatial oscillation at sub-terahertz frequencies. Time-resolved detection of such picosecond dynamics of quantum waves is important, as the detection provides a way of understanding the quantum behavior of electrons in nano-electronics. It also applies to quantum information technologies such as the ultrafast quantum-bit operation of quantum computing and high-sensitivity electromagnetic-field sensing. However, detecting picosecond dynamics has been a challenge since the sub-terahertz scale is far beyond the latest bandwidth measurement tools.
A KAIST team led by Professor Heung-Sun Sim developed a theory of ultrafast electron dynamics in a nanoscale trap, and proposed a scheme for detecting the dynamics, which utilizes a quantum-mechanical resonant state formed beside the trap. The coupling between the electron dynamics and the resonant state is switched on and off at a picosecond so that information on the dynamics is read out on the electric current being generated when the coupling is switched on.
NTT realized, together with NPL, the detection scheme and applied it to electron motions in a nanoscale trap formed in a silicon transistor. A single electron was captured in the trap by controlling electrostatic gates, and a resonant state was formed in the potential barrier of the trap.
The switching on and off of the coupling between the electron and the resonant state was achieved by aligning the resonance energy with the energy of the electron within a picosecond. An electric current from the trap through the resonant state to an electrode was measured at only a few Kelvin degrees, unveiling the spatial quantum-coherent oscillation of the electron with 250 GHz frequency inside the trap.
Professor Sim said, “This work suggests a scheme of detecting picosecond electron motions in submicron scales by utilizing quantum resonance. It will be useful in dynamical control of quantum mechanical electron waves for various purposes in nano-electronics, quantum sensing, and quantum information”.
This work was published online at Nature Nanotechnology on November 4. It was partly supported by the Korea National Research Foundation through the SRC Center for Quantum Coherence in Condensed Matter. For more on the NTT news release this article, please visit https://www.ntt.co.jp/news2019/1911e/191105a.html
-ProfileProfessor Heung-Sun Sim
Department of PhysicsDirector, SRC Center for Quantum Coherence in Condensed Matterhttps://qet.kaist.ac.kr KAIST
-Publication:Gento Yamahata, Sungguen Ryu, Nathan Johnson, H.-S. Sim, Akira Fujiwara, and Masaya Kataoka. 2019. Picosecond coherent electron motion in a silicon single-electron source. Nature Nanotechnology (Online Publication). 6 pages. https://doi.org/10.1038/s41565-019-0563-2
2019.11.05 View 19706
Ultrafast Quantum Motion in a Nanoscale Trap Detected
< Professor Heung-Sun Sim (left) and Co-author Dr. Sungguen Ryu (right) >
KAIST researchers have reported the detection of a picosecond electron motion in a silicon transistor. This study has presented a new protocol for measuring ultrafast electronic dynamics in an effective time-resolved fashion of picosecond resolution. The detection was made in collaboration with Nippon Telegraph and Telephone Corp. (NTT) in Japan and National Physical Laboratory (NPL) in the UK and is the first report to the best of our knowledge.
When an electron is captured in a nanoscale trap in solids, its quantum mechanical wave function can exhibit spatial oscillation at sub-terahertz frequencies. Time-resolved detection of such picosecond dynamics of quantum waves is important, as the detection provides a way of understanding the quantum behavior of electrons in nano-electronics. It also applies to quantum information technologies such as the ultrafast quantum-bit operation of quantum computing and high-sensitivity electromagnetic-field sensing. However, detecting picosecond dynamics has been a challenge since the sub-terahertz scale is far beyond the latest bandwidth measurement tools.
A KAIST team led by Professor Heung-Sun Sim developed a theory of ultrafast electron dynamics in a nanoscale trap, and proposed a scheme for detecting the dynamics, which utilizes a quantum-mechanical resonant state formed beside the trap. The coupling between the electron dynamics and the resonant state is switched on and off at a picosecond so that information on the dynamics is read out on the electric current being generated when the coupling is switched on.
NTT realized, together with NPL, the detection scheme and applied it to electron motions in a nanoscale trap formed in a silicon transistor. A single electron was captured in the trap by controlling electrostatic gates, and a resonant state was formed in the potential barrier of the trap.
The switching on and off of the coupling between the electron and the resonant state was achieved by aligning the resonance energy with the energy of the electron within a picosecond. An electric current from the trap through the resonant state to an electrode was measured at only a few Kelvin degrees, unveiling the spatial quantum-coherent oscillation of the electron with 250 GHz frequency inside the trap.
Professor Sim said, “This work suggests a scheme of detecting picosecond electron motions in submicron scales by utilizing quantum resonance. It will be useful in dynamical control of quantum mechanical electron waves for various purposes in nano-electronics, quantum sensing, and quantum information”.
This work was published online at Nature Nanotechnology on November 4. It was partly supported by the Korea National Research Foundation through the SRC Center for Quantum Coherence in Condensed Matter. For more on the NTT news release this article, please visit https://www.ntt.co.jp/news2019/1911e/191105a.html
-ProfileProfessor Heung-Sun Sim
Department of PhysicsDirector, SRC Center for Quantum Coherence in Condensed Matterhttps://qet.kaist.ac.kr KAIST
-Publication:Gento Yamahata, Sungguen Ryu, Nathan Johnson, H.-S. Sim, Akira Fujiwara, and Masaya Kataoka. 2019. Picosecond coherent electron motion in a silicon single-electron source. Nature Nanotechnology (Online Publication). 6 pages. https://doi.org/10.1038/s41565-019-0563-2
2019.11.05 View 19706 -
 A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29599
A Mathematical Model Reveals Long-Distance Cell Communication Mechanism
How can tens of thousands of people in a large football stadium all clap together with the same beat even though they can only hear the people near them clapping?
A combination of a partial differential equation and a synthetic circuit in microbes answers this question. An interdisciplinary collaborative team of Professor Jae Kyoung Kim at KAIST, Professor Krešimir Josić at the University of Houston, and Professor Matt Bennett at Rice University has identified how a large community can communicate with each other almost simultaneously even with very short distance signaling. The research was reported at Nature Chemical Biology.
Cells often communicate using signaling molecules, which can travel only a short distance. Nevertheless, the cells can also communicate over large distances to spur collective action. The team revealed a cell communication mechanism that quickly forms a network of local interactions to spur collective action, even in large communities.
The research team used an engineered transcriptional circuit of combined positive and negative feedback loops in E. coli, which can periodically release two types of signaling molecules: activator and repressor. As the signaling molecules travel over a short distance, cells can only talk to their nearest neighbors. However, cell communities synchronize oscillatory gene expression in spatially extended systems as long as the transcriptional circuit contains a positive feedback loop for the activator.
Professor Kim said that analyzing and understanding such high-dimensional dynamics was extremely difficult. He explained, “That’s why we used high-dimensional partial differential equation to describe the system based on the interactions among various types of molecules.” Surprisingly, the mathematical model accurately simulates the synthesis of the signaling molecules in the cell and their spatial diffusion throughout the chamber and their effect on neighboring cells.
The team simplified the high-dimensional system into a one-dimensional orbit, noting that the system repeats periodically. This allowed them to discover that cells can make one voice when they lowered their own voice and listened to the others. “It turns out the positive feedback loop reduces the distance between moving points and finally makes them move all together. That’s why you clap louder when you hear applause from nearby neighbors and everyone eventually claps together at almost the same time,” said Professor Kim.
Professor Kim added, “Math is a powerful as it simplifies complex thing so that we can find an essential underlying property. This finding would not have been possible without the simplification of complex systems using mathematics."
The National Institutes of Health, the National Science Foundation, the Robert A. Welch Foundation, the Hamill Foundation, the National Research Foundation of Korea, and the T.J. Park Science Fellowship of POSCO supported the research.
(Figure: Complex molecular interactions among microbial consortia is simplified as interactions among points on a limit cycle (right).)
2019.10.15 View 29599 -
 Professor Byong-Guk Park Named Scientist of October
< Professor Byong-Guk Park >
Professor Byong-Guk Park from the Department of Materials Science and Engineering was selected as the ‘Scientist of the Month’ for October 2019 by the Ministry of Science and ICT and the National Research Foundation of Korea. Professor Park was recognized for his contributions to the advancement of spin-orbit torque (SOT)-based magnetic random access memory (MRAM) technology. He received 10 million KRW in prize money.
A next-generation, non-volatile memory device MRAM consists of thin magnetic films. It can be applied in “logic-in-memory” devices, in which logic and memory functionalities coexist, thus drastically improving the performance of complementary metal-oxide semiconductor (CMOS) processors. Conventional MRAM technology is limited in its ability to increase the operation speed of a memory device while maintaining a high density.
Professor Park tackled this challenge by introducing a new material, antiferromagnet (IrMn), that generates a sizable amount of SOT as well as an exchange-bias field, which makes successful data writing possible without an external magnetic field. This research outcome paved the way for the development of MRAM, which has a simple device structure but features high speeds and density.
Professor Park said, “I feel rewarded to have forwarded the feasibility and applicability of MRAM. I will continue devoting myself to studying further on the development of new materials that can help enhance the performance of memory devices."
(END)
2019.10.10 View 11576
Professor Byong-Guk Park Named Scientist of October
< Professor Byong-Guk Park >
Professor Byong-Guk Park from the Department of Materials Science and Engineering was selected as the ‘Scientist of the Month’ for October 2019 by the Ministry of Science and ICT and the National Research Foundation of Korea. Professor Park was recognized for his contributions to the advancement of spin-orbit torque (SOT)-based magnetic random access memory (MRAM) technology. He received 10 million KRW in prize money.
A next-generation, non-volatile memory device MRAM consists of thin magnetic films. It can be applied in “logic-in-memory” devices, in which logic and memory functionalities coexist, thus drastically improving the performance of complementary metal-oxide semiconductor (CMOS) processors. Conventional MRAM technology is limited in its ability to increase the operation speed of a memory device while maintaining a high density.
Professor Park tackled this challenge by introducing a new material, antiferromagnet (IrMn), that generates a sizable amount of SOT as well as an exchange-bias field, which makes successful data writing possible without an external magnetic field. This research outcome paved the way for the development of MRAM, which has a simple device structure but features high speeds and density.
Professor Park said, “I feel rewarded to have forwarded the feasibility and applicability of MRAM. I will continue devoting myself to studying further on the development of new materials that can help enhance the performance of memory devices."
(END)
2019.10.10 View 11576 -
 Professor Ki-Jun Yoon selected as the 2019 SUHF Young Investigator
< Professor Ki-Jun Yoon >
Professor Ki-Jun Yoon from the Department of Biological Sciences was named one of four recipients of the 2019 Suh Kyung-Bae Science Foundation (SUHF) Young Investigator Awards.
The SUHF is a non-profit organization established in 2016 and funded by a personal donation of 300 billion KRW in shares from Chairman and CEO Kyung-Bae Suh of the Amorepacific Group. The primary purpose of the foundation is to serve as a platform to nurture and provide comprehensive long-term support for creative and passionate young Korean scientists committed to pursuing research in the field of life sciences. The SUHF selects three to five scientists through an open recruiting process every year, and grants each scientist a maximum of 2.5 billion KRW over a period of up to five years.
Since January this year, the foundation received 83 research proposals from scientists across the nation, especially from those who had less than five years of experience as professors, and selected the four recipients, including Professor Yoon.
Professor Yoon was recognized for his contributions to the advancement of research on how post-transcriptional mechanisms may modulate stem cell properties. His research project involves deciphering the molecular mechanisms controlling RNA metabolism in neural stem cells during normal development, and how alterations in RNA regulatory programs lead to human brain disorders.
< (From left) Professor Joo-Hong Park, Professor Yuree Lee, Chairman and CEO Kyung-Bae Suh, Professor Eunjung Lee, Professor Ki-Jun Yoon, ⓒ Amorepacific Group >
The other awards were given to Professor Joo-Hong Park and Professor Yuree Lee of Seoul National University, and Professor Eunjung Lee of Boston Children's Hospital and Harvard Medical School.
The awards ceremony was held on September 18 at the Amorepacific Headquarters in Seoul.
With these four new awardees, a total of 14 scientists have been named as SUHF Young Investigators to date.
(END)
2019.09.23 View 10239
Professor Ki-Jun Yoon selected as the 2019 SUHF Young Investigator
< Professor Ki-Jun Yoon >
Professor Ki-Jun Yoon from the Department of Biological Sciences was named one of four recipients of the 2019 Suh Kyung-Bae Science Foundation (SUHF) Young Investigator Awards.
The SUHF is a non-profit organization established in 2016 and funded by a personal donation of 300 billion KRW in shares from Chairman and CEO Kyung-Bae Suh of the Amorepacific Group. The primary purpose of the foundation is to serve as a platform to nurture and provide comprehensive long-term support for creative and passionate young Korean scientists committed to pursuing research in the field of life sciences. The SUHF selects three to five scientists through an open recruiting process every year, and grants each scientist a maximum of 2.5 billion KRW over a period of up to five years.
Since January this year, the foundation received 83 research proposals from scientists across the nation, especially from those who had less than five years of experience as professors, and selected the four recipients, including Professor Yoon.
Professor Yoon was recognized for his contributions to the advancement of research on how post-transcriptional mechanisms may modulate stem cell properties. His research project involves deciphering the molecular mechanisms controlling RNA metabolism in neural stem cells during normal development, and how alterations in RNA regulatory programs lead to human brain disorders.
< (From left) Professor Joo-Hong Park, Professor Yuree Lee, Chairman and CEO Kyung-Bae Suh, Professor Eunjung Lee, Professor Ki-Jun Yoon, ⓒ Amorepacific Group >
The other awards were given to Professor Joo-Hong Park and Professor Yuree Lee of Seoul National University, and Professor Eunjung Lee of Boston Children's Hospital and Harvard Medical School.
The awards ceremony was held on September 18 at the Amorepacific Headquarters in Seoul.
With these four new awardees, a total of 14 scientists have been named as SUHF Young Investigators to date.
(END)
2019.09.23 View 10239