BIO
-
 A Korean research team develops a new clinical candidate for fatty liver disease
A team of Korean researchers have succeeded in developing a new drug candidate for the treatment of non-alcoholic fatty liver disease (NAFLD) acting on peripheral tissues. To date, there has not been an optimal treatment for non-alcoholic steatohepatitis (NASH), and this discovery is expected to set the grounds for the development of new drugs that can safely suppress both liver fat accumulation and liver fibrosis at the same time.
A joint research team led by Professor Jin Hee Ahn from Gwangju Institute of Science and Technology (GIST) and Professor Hail Kim from the KAIST Graduate School of Medical Science and Engineering developed a new chemical that can suppress disease-specific protein (HTR2A) through years of basic research. The team also revealed to have verified its efficacy and safety through preclinical tests (animal tests) at JD Bioscience Inc., a start-up company founded by Professor Ahn.
Although NAFLD has a prevalence rate as high as 20-30%, and about 5% of the global adult population suffers from NASH, there are no commercial drugs targeting them to date. NAFLD is a chronic disease that starts from the fatty liver and progresses into steatohepatitis, fibrosis, cirrhosis, and liver cancer. The mortality rate of patients increases with accompanied cardiovascular diseases and liver-related complications, and appropriate treatment in the early stage is hence necessary.
< Figure 1. Strategy and history of 5HT2A antagonists. Library and rational design for the development of compound 11c as a potent 5HT2A antagonist. Previous research efforts were discontinued due to limited oral absorption and safety. A therapeutic candidate to overcome this problem was identified and phase 1 clinical trials are currently in progress. >
The new synthetic chemical developed by the joint GIST-KAIST research is an innovative drug candidate that shows therapeutic effects on NASH based on a dual action mechanism that inhibits the accumulation of fat in the liver and liver fibrosis by suppressing the serotonin receptor protein 5HT2A.
The research team confirmed its therapeutic effects in animal models for NAFLD and NASH, in which hepatic steatosis and liver fibrosis* caused by fat accumulation in the liver were suppressed simultaneously by 50-70%.
*fibrosis: stiffening of parts of the liver, also used as a major indicator to track the prognosis of steatosis
The research team explained that the material was designed with optimal polarity and lipid affinity to minimize its permeability across the blood-brain barrier. It therefore does not affect the brain, and causes little side effects in the central nervous system (CNS) such as depression and suicidal ideations, while demonstrating excellent inhibition on its target protein present in tissues outside brain (IC50* = 14 nM). The team also demonstrated its superior efficacy in improving liver fibrosis when compared to similar drugs in the phase 3 clinical trial.
*IC50 (half maximal inhibitory concentration): the concentration at which a chemical suppresses 50% of a particular biological function
< Figure 2. GM-60106 (11c)'s effect on obesity: When GM-60106 was administered to an obese animal model (mice) for 2 months, body weight, body fat mass, and blood sugar were significantly reduced (a-d). In addition, the steatohepatitis level (NAFLD Activity Score) and the expression of genes of the treated mice involved in adipogenesis along with blood/liver fat decreased (e-h) >
Based on the pharmacological data obtained through preclinical trials, the team evaluated the effects of the drug on 88 healthy adults as part of their phase 1 clinical trial, where the side effects and the safe dosage of a drug are tested against healthy adults. Results showed no serious side effects and a good level of drug safety.
In addition, a preliminary efficacy evaluation on eight adults with steatohepatitis is currently underway.
Professor Jin Hee Ahn said, “The aim of this research is to develop a treatment for NASH with little side effects and guaranteed safety by developing a new target. The developed chemical is currently going through phase 1 of the global clinical trial in Australia through JD Bioscience Inc., a bio venture company for innovative drug development.” he added, “The candidate material the research team is currently developing shows not only a high level of safety and preventative effects by suppressing fat accumulation in the liver, but also a direct therapeutic effect on liver fibrosis. This is a strength that distinguishes our material from other competing drugs.”
< Figure 3. Efficacy of GM-60106 (11c) on liver fibrosis: When GM-60106 was administered to a steatohepatitis model (mice) for 3 months, the expression of genes associated with tissue fibrosis was significantly reduced (b-c). As a result of a detailed analysis of the tissues of the animal model, it was confirmed that the rate of tissue fibrosis was reduced and the expression rate of genes related to tissue fibrosis and inflammation was also significantly reduced (e-h). >
Professor Hail Kim from KAIST said, “Until now, this disease did not have a method of treatment other than weight control, and there has been no attempt to develop a drug that can be used for non-obese patients.” He added, “Through this research, we look forward to the development of various treatment techniques targeting a range of metabolic diseases including NASH that do not affect the weight of the patient.”
This study, conducted together by the research teams led by Professor Ahn from GIST and Professor Kim from KAIST, as well as the research team from JD Bioscience Inc., was supported by the Ministry of Science and ICT, and the National New Drug Development Project. The results of this research were published by Nature Communications on January 20.
The team also presented the results of their clinical study on the candidate material coded GM-60106 targeting metabolic abnormality-related MASH* at NASH-TAG Conference 2024, which was held in Utah for three days starting on January 4, which was selected as an excellent abstract.
*MASH (Metabolic Dysfunction-Associated Steatohepatitis): new replacement term for NASH
2024.02.21 View 9590
A Korean research team develops a new clinical candidate for fatty liver disease
A team of Korean researchers have succeeded in developing a new drug candidate for the treatment of non-alcoholic fatty liver disease (NAFLD) acting on peripheral tissues. To date, there has not been an optimal treatment for non-alcoholic steatohepatitis (NASH), and this discovery is expected to set the grounds for the development of new drugs that can safely suppress both liver fat accumulation and liver fibrosis at the same time.
A joint research team led by Professor Jin Hee Ahn from Gwangju Institute of Science and Technology (GIST) and Professor Hail Kim from the KAIST Graduate School of Medical Science and Engineering developed a new chemical that can suppress disease-specific protein (HTR2A) through years of basic research. The team also revealed to have verified its efficacy and safety through preclinical tests (animal tests) at JD Bioscience Inc., a start-up company founded by Professor Ahn.
Although NAFLD has a prevalence rate as high as 20-30%, and about 5% of the global adult population suffers from NASH, there are no commercial drugs targeting them to date. NAFLD is a chronic disease that starts from the fatty liver and progresses into steatohepatitis, fibrosis, cirrhosis, and liver cancer. The mortality rate of patients increases with accompanied cardiovascular diseases and liver-related complications, and appropriate treatment in the early stage is hence necessary.
< Figure 1. Strategy and history of 5HT2A antagonists. Library and rational design for the development of compound 11c as a potent 5HT2A antagonist. Previous research efforts were discontinued due to limited oral absorption and safety. A therapeutic candidate to overcome this problem was identified and phase 1 clinical trials are currently in progress. >
The new synthetic chemical developed by the joint GIST-KAIST research is an innovative drug candidate that shows therapeutic effects on NASH based on a dual action mechanism that inhibits the accumulation of fat in the liver and liver fibrosis by suppressing the serotonin receptor protein 5HT2A.
The research team confirmed its therapeutic effects in animal models for NAFLD and NASH, in which hepatic steatosis and liver fibrosis* caused by fat accumulation in the liver were suppressed simultaneously by 50-70%.
*fibrosis: stiffening of parts of the liver, also used as a major indicator to track the prognosis of steatosis
The research team explained that the material was designed with optimal polarity and lipid affinity to minimize its permeability across the blood-brain barrier. It therefore does not affect the brain, and causes little side effects in the central nervous system (CNS) such as depression and suicidal ideations, while demonstrating excellent inhibition on its target protein present in tissues outside brain (IC50* = 14 nM). The team also demonstrated its superior efficacy in improving liver fibrosis when compared to similar drugs in the phase 3 clinical trial.
*IC50 (half maximal inhibitory concentration): the concentration at which a chemical suppresses 50% of a particular biological function
< Figure 2. GM-60106 (11c)'s effect on obesity: When GM-60106 was administered to an obese animal model (mice) for 2 months, body weight, body fat mass, and blood sugar were significantly reduced (a-d). In addition, the steatohepatitis level (NAFLD Activity Score) and the expression of genes of the treated mice involved in adipogenesis along with blood/liver fat decreased (e-h) >
Based on the pharmacological data obtained through preclinical trials, the team evaluated the effects of the drug on 88 healthy adults as part of their phase 1 clinical trial, where the side effects and the safe dosage of a drug are tested against healthy adults. Results showed no serious side effects and a good level of drug safety.
In addition, a preliminary efficacy evaluation on eight adults with steatohepatitis is currently underway.
Professor Jin Hee Ahn said, “The aim of this research is to develop a treatment for NASH with little side effects and guaranteed safety by developing a new target. The developed chemical is currently going through phase 1 of the global clinical trial in Australia through JD Bioscience Inc., a bio venture company for innovative drug development.” he added, “The candidate material the research team is currently developing shows not only a high level of safety and preventative effects by suppressing fat accumulation in the liver, but also a direct therapeutic effect on liver fibrosis. This is a strength that distinguishes our material from other competing drugs.”
< Figure 3. Efficacy of GM-60106 (11c) on liver fibrosis: When GM-60106 was administered to a steatohepatitis model (mice) for 3 months, the expression of genes associated with tissue fibrosis was significantly reduced (b-c). As a result of a detailed analysis of the tissues of the animal model, it was confirmed that the rate of tissue fibrosis was reduced and the expression rate of genes related to tissue fibrosis and inflammation was also significantly reduced (e-h). >
Professor Hail Kim from KAIST said, “Until now, this disease did not have a method of treatment other than weight control, and there has been no attempt to develop a drug that can be used for non-obese patients.” He added, “Through this research, we look forward to the development of various treatment techniques targeting a range of metabolic diseases including NASH that do not affect the weight of the patient.”
This study, conducted together by the research teams led by Professor Ahn from GIST and Professor Kim from KAIST, as well as the research team from JD Bioscience Inc., was supported by the Ministry of Science and ICT, and the National New Drug Development Project. The results of this research were published by Nature Communications on January 20.
The team also presented the results of their clinical study on the candidate material coded GM-60106 targeting metabolic abnormality-related MASH* at NASH-TAG Conference 2024, which was held in Utah for three days starting on January 4, which was selected as an excellent abstract.
*MASH (Metabolic Dysfunction-Associated Steatohepatitis): new replacement term for NASH
2024.02.21 View 9590 -
 KAIST presents strategies for environmentally friendly and sustainable polyamides production
- Provides current research trends in bio-based polyamide production
- Research on bio-based polyamides production gains importance for achieving a carbon-neutral society
Global industries focused on carbon neutrality, under the slogan "Net-Zero," are gaining increasing attention. In particular, research on microbial production of polymers, replacing traditional chemical methods with biological approaches, is actively progressing.
Polyamides, represented by nylon, are linear polymers widely used in various industries such as automotive, electronics, textiles, and medical fields. They possess beneficial properties such as high tensile strength, electrical insulation, heat resistance, wear resistance, and biocompatibility. Since the commercialization of nylon in 1938, approximately 7 million tons of polyamides are produced worldwide annually. Considering their broad applications and significance, producing polyamides through bio-based methods holds considerable environmental and industrial importance.
KAIST (President Kwang-Hyung Lee) announced that a research team led by Distinguished Professor Sang Yup Lee, including Dr. Jong An Lee and doctoral candidate Ji Yeon Kim from the Department of Chemical and Biomolecular Engineering, published a paper titled "Current Advancements in Bio-Based Production of Polyamides”. The paper was featured on the cover of the monthly issue of "Trends in Chemistry” by Cell Press.
As part of climate change response technologies, bio-refineries involve using biotechnological and chemical methods to produce industrially important chemicals and biofuels from renewable biomass without relying on fossil resources. Notably, systems metabolic engineering, pioneered by KAIST's Distinguished Professor Sang Yup Lee, is a research field that effectively manipulates microbial metabolic pathways to produce valuable chemicals, forming the core technology for bio-refineries. The research team has successfully developed high-performance strains producing a variety of compounds, including succinic acid, biodegradable plastics, biofuels, and natural products, using systems metabolic engineering tools and strategies.
The research team predicted that if bio-based polyamide production technology, which is widely used in the production of clothing and textiles, becomes widespread, it will attract attention as a future technology that can respond to the climate crisis due to its environment-friendly production technology. In this study, the research team comprehensively reviewed the bio-based polyamide production strategies. They provided insights into the advancements in polyamide monomer production using metabolically engineered microorganisms and highlighted the recent trends in bio-based polyamide advancements utilizing these monomers. Additionally, they reviewed the strategies for synthesizing bio-based polyamides through chemical conversion of natural oils and discussed the biodegradability and recycling of the polyamides. Furthermore, the paper presented the future direction in which metabolic engineering can be applied for the bio-based polyamide production, contributing to environmentally friendly and sustainable society.
Ji Yeon Kim, the co-first author of this paper from KAIST, stated "The importance of utilizing systems metabolic engineering tools and strategies for bio-based polyamides production is becoming increasingly prominent in achieving carbon neutrality."
Professor Sang Yup Lee emphasized, "Amid growing concerns about climate change, the significance of environmentally friendly and sustainable industrial development is greater than ever. Systems metabolic engineering is expected to have a significant impact not only on the chemical industry but also in various fields."
< [Figure 1] A schematic overview of the overall process for polyamides production >
This paper by Dr. Jong An Lee, PhD student Ji Yeon Kim, Dr. Jung Ho Ahn, and Master Yeah-Ji Ahn from the Department of Chemical and Biomolecular Engineering at KAIST was published in the December issue of 'Trends in Chemistry', an authoritative review journal in the field of chemistry published by Cell. It was published on December 7 as the cover paper and featured review.
※ Paper title: Current advancements in the bio-based production of polyamides
※ Author information: Jong An Lee, Ji Yeon Kim, Jung Ho Ahn, Yeah-Ji Ahn, and Sang Yup Lee
This research was conducted with the support from the development of platform technologies of microbial cell factories for the next-generation biorefineries project and C1 gas refinery program by Korean Ministry of Science and ICT.
< [Figure 2] Cover paper of the December issue of Trends in Chemistry >
2023.12.21 View 7381
KAIST presents strategies for environmentally friendly and sustainable polyamides production
- Provides current research trends in bio-based polyamide production
- Research on bio-based polyamides production gains importance for achieving a carbon-neutral society
Global industries focused on carbon neutrality, under the slogan "Net-Zero," are gaining increasing attention. In particular, research on microbial production of polymers, replacing traditional chemical methods with biological approaches, is actively progressing.
Polyamides, represented by nylon, are linear polymers widely used in various industries such as automotive, electronics, textiles, and medical fields. They possess beneficial properties such as high tensile strength, electrical insulation, heat resistance, wear resistance, and biocompatibility. Since the commercialization of nylon in 1938, approximately 7 million tons of polyamides are produced worldwide annually. Considering their broad applications and significance, producing polyamides through bio-based methods holds considerable environmental and industrial importance.
KAIST (President Kwang-Hyung Lee) announced that a research team led by Distinguished Professor Sang Yup Lee, including Dr. Jong An Lee and doctoral candidate Ji Yeon Kim from the Department of Chemical and Biomolecular Engineering, published a paper titled "Current Advancements in Bio-Based Production of Polyamides”. The paper was featured on the cover of the monthly issue of "Trends in Chemistry” by Cell Press.
As part of climate change response technologies, bio-refineries involve using biotechnological and chemical methods to produce industrially important chemicals and biofuels from renewable biomass without relying on fossil resources. Notably, systems metabolic engineering, pioneered by KAIST's Distinguished Professor Sang Yup Lee, is a research field that effectively manipulates microbial metabolic pathways to produce valuable chemicals, forming the core technology for bio-refineries. The research team has successfully developed high-performance strains producing a variety of compounds, including succinic acid, biodegradable plastics, biofuels, and natural products, using systems metabolic engineering tools and strategies.
The research team predicted that if bio-based polyamide production technology, which is widely used in the production of clothing and textiles, becomes widespread, it will attract attention as a future technology that can respond to the climate crisis due to its environment-friendly production technology. In this study, the research team comprehensively reviewed the bio-based polyamide production strategies. They provided insights into the advancements in polyamide monomer production using metabolically engineered microorganisms and highlighted the recent trends in bio-based polyamide advancements utilizing these monomers. Additionally, they reviewed the strategies for synthesizing bio-based polyamides through chemical conversion of natural oils and discussed the biodegradability and recycling of the polyamides. Furthermore, the paper presented the future direction in which metabolic engineering can be applied for the bio-based polyamide production, contributing to environmentally friendly and sustainable society.
Ji Yeon Kim, the co-first author of this paper from KAIST, stated "The importance of utilizing systems metabolic engineering tools and strategies for bio-based polyamides production is becoming increasingly prominent in achieving carbon neutrality."
Professor Sang Yup Lee emphasized, "Amid growing concerns about climate change, the significance of environmentally friendly and sustainable industrial development is greater than ever. Systems metabolic engineering is expected to have a significant impact not only on the chemical industry but also in various fields."
< [Figure 1] A schematic overview of the overall process for polyamides production >
This paper by Dr. Jong An Lee, PhD student Ji Yeon Kim, Dr. Jung Ho Ahn, and Master Yeah-Ji Ahn from the Department of Chemical and Biomolecular Engineering at KAIST was published in the December issue of 'Trends in Chemistry', an authoritative review journal in the field of chemistry published by Cell. It was published on December 7 as the cover paper and featured review.
※ Paper title: Current advancements in the bio-based production of polyamides
※ Author information: Jong An Lee, Ji Yeon Kim, Jung Ho Ahn, Yeah-Ji Ahn, and Sang Yup Lee
This research was conducted with the support from the development of platform technologies of microbial cell factories for the next-generation biorefineries project and C1 gas refinery program by Korean Ministry of Science and ICT.
< [Figure 2] Cover paper of the December issue of Trends in Chemistry >
2023.12.21 View 7381 -
 KAIST introduces eco-friendly technologies for plastic production and biodegradation
- A research team under Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering published a paper in Nature Microbiology on the overview and trends of plastic production and degradation technology using microorganisms.
- Eco-friendly and sustainable plastic production and degradation technology using microorganisms as a core technology to achieve a plastic circular economy was presented.
Plastic is one of the important materials in modern society, with approximately 460 million tons produced annually and with expected production reaching approximately 1.23 billion tons in 2060. However, since 1950, plastic waste totaling more than 6.3 billion tons has been generated, and it is believed that more than 140 million tons of plastic waste has accumulated in the aquatic environment. Recently, the seriousness of microplastic pollution has emerged, not only posing a risk to the marine ecosystem and human health, but also worsening global warming by inhibiting the activity of marine plankton, which play an important role in lowering the Earth's carbon dioxide concentration.
KAIST President Kwang-Hyung Lee announced on December 11 that a research team under Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering had published a paper titled 'Sustainable production and degradation of plastics using microbes', which covers the latest technologies for producing plastics and processing waste plastics in an eco-friendly manner using microorganisms.
As the international community moves to solve this plastic problem, various efforts are being made, including 175 countries participating to conclude a legally binding agreement with the goal of ending plastic pollution by 2024. Various technologies are being developed for sustainable plastic production and processing, and among them, biotechnology using microorganisms is attracting attention.
Microorganisms have the ability to naturally produce or decompose certain compounds, and this ability is maximized through biotechnologies such as metabolic engineering and enzyme engineering to produce plastics from renewable biomass resources instead of fossil raw materials and to decompose waste plastics.
Accordingly, the research team comprehensively analyzed the latest microorganism-based technologies for the sustainable production and decomposition of plastics and presented how they actually contribute to solving the plastic problem. Based on this, they presented limitations, prospects, and research directions of the technologies for achieving a circular economy for plastics.
Microorganism-based technologies for various plastics range from widely used synthetic plastics such as polyethylene (PE) to promising bioplastics such as natural polymers derived from microorganisms (polyhydroxyalkanoate (PHA)) that are completely biodegradable in the natural environment and do not pose a risk of microplastic generation. Commercialization statuses and latest technologies were also discussed. In addition, the technology to decompose these plastics using microorganisms and their enzymes and the upcycling technology to convert them into other useful compounds after decomposition were introduced, highlighting the competitiveness and potential of technology using microorganisms.
First author So Young Choi, a research assistant professor in the Department of Chemical and Biomolecular Engineering at KAIST, said, “In the future, we will be able to easily find eco-friendly plastics made using microorganisms all around us,” and corresponding author Distinguished Professor Sang Yup Lee said, “Plastic can be made more sustainable. It is important to use plastics responsibly to protect the environment and simultaneously achieve economic and social development through the new plastics industry, and we look forward to the improved performance of microbial metabolic engineering technology.”
This paper was published on November 30th in the online edition of Nature Microbiology.
※ Paper Title : Sustainable production and degradation of plastics using microbes
Authors: So Young Choi, Youngjoon Lee, Hye Eun Yu, In Jin Cho, Minju Kang & Sang Yup Lee
2023.12.11 View 7325
KAIST introduces eco-friendly technologies for plastic production and biodegradation
- A research team under Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering published a paper in Nature Microbiology on the overview and trends of plastic production and degradation technology using microorganisms.
- Eco-friendly and sustainable plastic production and degradation technology using microorganisms as a core technology to achieve a plastic circular economy was presented.
Plastic is one of the important materials in modern society, with approximately 460 million tons produced annually and with expected production reaching approximately 1.23 billion tons in 2060. However, since 1950, plastic waste totaling more than 6.3 billion tons has been generated, and it is believed that more than 140 million tons of plastic waste has accumulated in the aquatic environment. Recently, the seriousness of microplastic pollution has emerged, not only posing a risk to the marine ecosystem and human health, but also worsening global warming by inhibiting the activity of marine plankton, which play an important role in lowering the Earth's carbon dioxide concentration.
KAIST President Kwang-Hyung Lee announced on December 11 that a research team under Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering had published a paper titled 'Sustainable production and degradation of plastics using microbes', which covers the latest technologies for producing plastics and processing waste plastics in an eco-friendly manner using microorganisms.
As the international community moves to solve this plastic problem, various efforts are being made, including 175 countries participating to conclude a legally binding agreement with the goal of ending plastic pollution by 2024. Various technologies are being developed for sustainable plastic production and processing, and among them, biotechnology using microorganisms is attracting attention.
Microorganisms have the ability to naturally produce or decompose certain compounds, and this ability is maximized through biotechnologies such as metabolic engineering and enzyme engineering to produce plastics from renewable biomass resources instead of fossil raw materials and to decompose waste plastics.
Accordingly, the research team comprehensively analyzed the latest microorganism-based technologies for the sustainable production and decomposition of plastics and presented how they actually contribute to solving the plastic problem. Based on this, they presented limitations, prospects, and research directions of the technologies for achieving a circular economy for plastics.
Microorganism-based technologies for various plastics range from widely used synthetic plastics such as polyethylene (PE) to promising bioplastics such as natural polymers derived from microorganisms (polyhydroxyalkanoate (PHA)) that are completely biodegradable in the natural environment and do not pose a risk of microplastic generation. Commercialization statuses and latest technologies were also discussed. In addition, the technology to decompose these plastics using microorganisms and their enzymes and the upcycling technology to convert them into other useful compounds after decomposition were introduced, highlighting the competitiveness and potential of technology using microorganisms.
First author So Young Choi, a research assistant professor in the Department of Chemical and Biomolecular Engineering at KAIST, said, “In the future, we will be able to easily find eco-friendly plastics made using microorganisms all around us,” and corresponding author Distinguished Professor Sang Yup Lee said, “Plastic can be made more sustainable. It is important to use plastics responsibly to protect the environment and simultaneously achieve economic and social development through the new plastics industry, and we look forward to the improved performance of microbial metabolic engineering technology.”
This paper was published on November 30th in the online edition of Nature Microbiology.
※ Paper Title : Sustainable production and degradation of plastics using microbes
Authors: So Young Choi, Youngjoon Lee, Hye Eun Yu, In Jin Cho, Minju Kang & Sang Yup Lee
2023.12.11 View 7325 -
 KAIST holds its first ‘KAIST Tech Fair’ in New York, USA
< Photo 1. 2023 KAIST Tech Fair in New York >
KAIST (President Kwang-Hyung Lee) announced on the 11th that it will hold the ‘2023 KAIST Tech Fair in New York’ at the Kimmel Center at New York University in Manhattan, USA, on the 22nd of this month. It is an event designed to be the starting point for KAIST to expand its startup ecosystem into the global stage, and it is to attract investments and secure global customers in New York by demonstrating the technological value of KAIST startup companies directly at location.
< Photo 2. President Kwang Hyung Lee at the 2023 KAIST Tech Fair in New York >
KAIST has been holding briefing sessions for technology transfer in Korea every year since 2018, and this year is the first time to hold a tech fair overseas for global companies.
KAIST Institute of Technology Value Creation (Director Sung-Yool Choi) has prepared for this event over the past six months with the Korea International Trade Association (hereinafter KITA, CEO Christopher Koo) to survey customer base and investment companies to conduct market analysis.
Among the companies founded with the technologies developed by the faculty and students of KAIST and their partners, 7 companies were selected to be matched with companies overseas that expressed interests in these technologies. Global multinational companies in the fields of IT, artificial intelligence, environment, logistics, distribution, and retail are participating as demand agencies and are testing the marketability of the start-up's technology as of September.
Daim Research, founded by Professor Young Jae Jang of the Department of Industrial and Systems Engineering, is a company specializing in smart factory automation solutions and is knocking on the door of the global market with a platform technology optimized for automated logistics systems.
< Photo 3. Presentation by Professor Young Jae Jang for DAIM Research >
It is a ‘collaborative intelligence’ solution that maximizes work productivity by having a number of robots used in industrial settings collaborate with one another. The strength of their solution is that logistics robots equipped with AI reinforced learning technology can respond to processes and environmental changes on their own, minimizing maintenance costs and the system can achieve excellent performance even with a small amount of data when it is combined with the digital twin technology the company has developed on its own.
A student startup, ‘Aniai’, is entering the US market, the home of hamburgers, with hamburger patty automation equipments and solutions. This is a robot kitchen startup founded by its CEO Gunpil Hwang, a graduate of KAIST’s School of Electrical Engineering which gathered together the experts in the fields of robot control, design, and artificial intelligence and cognitive technology to develop technology to automatically cook hamburger patties.
At the touch of a button, both sides of the patty are cooked simultaneously for consistent taste and quality according to the set condition. Since it can cook about 200 dishes in an hour, it is attracting attention as a technology that can not only solve manpower shortages but also accelerate the digital transformation of the restaurant industry.
Also, at the tech fair to be held at the Kimmel Center of New York University on the 22nd, the following startups who are currently under market verification in the U.S. will be participating: ▴'TheWaveTalk', which developed a water quality management system that can measure external substances and metal ions by transferring original technology from KAIST; ▴‘VIRNECT’, which helps workers improve their skills by remotely managing industrial sites using XR*; ▴‘Datumo’, a solution that helps process and analyze artificial intelligence big data, ▴‘VESSL AI’, the provider of a solution to eliminate the overhead** of machine learning systems; and ▴ ‘DolbomDream’, which developed an inflatable vest that helps the psychological stability of people with developmental disabilities.
* XR (eXtended Reality): Ultra-realistic technology that enhances immersion by utilizing augmented reality, virtual reality, and mixed reality technologies
** Overhead: Additional time required for stable processing of the program
In addition, two companies (Plasmapp and NotaAI) that are participating in the D-Unicorn program with the support of the Daejeon City and two companies (Enget and ILIAS Biologics) that are receiving support from the Scale Up Tips of the Ministry of SMEs and Startups, three companies (WiPowerOne, IDK Lab, and Artificial Photosynthesis Lab) that are continuing to realize the sustainable development goals for a total of 14 KAIST startups, will hold a corporate information session with about 100 invited guests from global companies and venture capital.
< Photo 4. Presentation for AP Lab >
Prior to this event, participating startups will be visiting the New York Economic Development Corporation and large law firms to receive advice on U.S. government support programs and on their attemps to enter the U.S. market. In addition, the participating companies plan to visit a startup support investment institution pursuing sustainable development goals and the Leslie eLab, New York University's one-stop startup support space, to lay the foundation for KAIST's leap forward in global technology commercialization.
< Photo 5. Sung-Yool Choi, the Director of KAIST Institute of Technology Value Creation (left) at the 2023 KAIST Tech Fair in New York with the key participants >
Sung-Yool Choi, the Director of KAIST Institute of Technology Value Creation, said, “KAIST prepared this event to realize its vision of being a leading university in creating global value.” He added, “We hope that our startups founded with KAIST technology would successfully completed market verification to be successful in securing global demands and in attracting investments for their endeavors.”
2023.09.11 View 18577
KAIST holds its first ‘KAIST Tech Fair’ in New York, USA
< Photo 1. 2023 KAIST Tech Fair in New York >
KAIST (President Kwang-Hyung Lee) announced on the 11th that it will hold the ‘2023 KAIST Tech Fair in New York’ at the Kimmel Center at New York University in Manhattan, USA, on the 22nd of this month. It is an event designed to be the starting point for KAIST to expand its startup ecosystem into the global stage, and it is to attract investments and secure global customers in New York by demonstrating the technological value of KAIST startup companies directly at location.
< Photo 2. President Kwang Hyung Lee at the 2023 KAIST Tech Fair in New York >
KAIST has been holding briefing sessions for technology transfer in Korea every year since 2018, and this year is the first time to hold a tech fair overseas for global companies.
KAIST Institute of Technology Value Creation (Director Sung-Yool Choi) has prepared for this event over the past six months with the Korea International Trade Association (hereinafter KITA, CEO Christopher Koo) to survey customer base and investment companies to conduct market analysis.
Among the companies founded with the technologies developed by the faculty and students of KAIST and their partners, 7 companies were selected to be matched with companies overseas that expressed interests in these technologies. Global multinational companies in the fields of IT, artificial intelligence, environment, logistics, distribution, and retail are participating as demand agencies and are testing the marketability of the start-up's technology as of September.
Daim Research, founded by Professor Young Jae Jang of the Department of Industrial and Systems Engineering, is a company specializing in smart factory automation solutions and is knocking on the door of the global market with a platform technology optimized for automated logistics systems.
< Photo 3. Presentation by Professor Young Jae Jang for DAIM Research >
It is a ‘collaborative intelligence’ solution that maximizes work productivity by having a number of robots used in industrial settings collaborate with one another. The strength of their solution is that logistics robots equipped with AI reinforced learning technology can respond to processes and environmental changes on their own, minimizing maintenance costs and the system can achieve excellent performance even with a small amount of data when it is combined with the digital twin technology the company has developed on its own.
A student startup, ‘Aniai’, is entering the US market, the home of hamburgers, with hamburger patty automation equipments and solutions. This is a robot kitchen startup founded by its CEO Gunpil Hwang, a graduate of KAIST’s School of Electrical Engineering which gathered together the experts in the fields of robot control, design, and artificial intelligence and cognitive technology to develop technology to automatically cook hamburger patties.
At the touch of a button, both sides of the patty are cooked simultaneously for consistent taste and quality according to the set condition. Since it can cook about 200 dishes in an hour, it is attracting attention as a technology that can not only solve manpower shortages but also accelerate the digital transformation of the restaurant industry.
Also, at the tech fair to be held at the Kimmel Center of New York University on the 22nd, the following startups who are currently under market verification in the U.S. will be participating: ▴'TheWaveTalk', which developed a water quality management system that can measure external substances and metal ions by transferring original technology from KAIST; ▴‘VIRNECT’, which helps workers improve their skills by remotely managing industrial sites using XR*; ▴‘Datumo’, a solution that helps process and analyze artificial intelligence big data, ▴‘VESSL AI’, the provider of a solution to eliminate the overhead** of machine learning systems; and ▴ ‘DolbomDream’, which developed an inflatable vest that helps the psychological stability of people with developmental disabilities.
* XR (eXtended Reality): Ultra-realistic technology that enhances immersion by utilizing augmented reality, virtual reality, and mixed reality technologies
** Overhead: Additional time required for stable processing of the program
In addition, two companies (Plasmapp and NotaAI) that are participating in the D-Unicorn program with the support of the Daejeon City and two companies (Enget and ILIAS Biologics) that are receiving support from the Scale Up Tips of the Ministry of SMEs and Startups, three companies (WiPowerOne, IDK Lab, and Artificial Photosynthesis Lab) that are continuing to realize the sustainable development goals for a total of 14 KAIST startups, will hold a corporate information session with about 100 invited guests from global companies and venture capital.
< Photo 4. Presentation for AP Lab >
Prior to this event, participating startups will be visiting the New York Economic Development Corporation and large law firms to receive advice on U.S. government support programs and on their attemps to enter the U.S. market. In addition, the participating companies plan to visit a startup support investment institution pursuing sustainable development goals and the Leslie eLab, New York University's one-stop startup support space, to lay the foundation for KAIST's leap forward in global technology commercialization.
< Photo 5. Sung-Yool Choi, the Director of KAIST Institute of Technology Value Creation (left) at the 2023 KAIST Tech Fair in New York with the key participants >
Sung-Yool Choi, the Director of KAIST Institute of Technology Value Creation, said, “KAIST prepared this event to realize its vision of being a leading university in creating global value.” He added, “We hope that our startups founded with KAIST technology would successfully completed market verification to be successful in securing global demands and in attracting investments for their endeavors.”
2023.09.11 View 18577 -
 KAIST presents a microbial cell factory as a source of eco-friendly food and cosmetic coloring
Despite decades of global population growth, global food crisis seems to be at hand yet again because the food productivity is cut severely due to prolonged presence of abnormal weather from intensifying climate change and global food supply chain is deteriorated due to international conflicts such as wars exacerbating food shortages and nutritional inequality around the globe. At the same time, however, as awareness of the environment and sustainability rises, an increase in demand for more eco-friendly and high-quality food and beauty products is being observed not without a sense of irony. At a time like this, microorganisms are attracting attention as a key that can handle this couple of seemingly distant problems.
KAIST (President Kwang-Hyung Lee) announced on the 26th that Kyeong Rok Choi, a research professor of the Bioprocess Research Center and Sang Yup Lee, a Distinguished Professor of the Department of Chemical and Biomolecular Engineering, published a paper titled “Metabolic Engineering of Microorganisms for Food and Cosmetics Production” upon invitation by “Nature Reviews Bioengineering” to be published online published by Nature after peer review.
※ Paper title: Systems metabolic engineering of microorganisms for food and cosmetics production
※ Author information: Kyeong Rok Choi (first author) and Sang Yup Lee (corresponding author)
Systems metabolic engineering is a research field founded by Distinguished Professor Sang Yup Lee of KAIST to more effectively develop microbial cell factories, the core factor of the next-generation bio industry to replace the existing chemical industry that relies heavily on petroleum. By applying a systemic metabolic engineering strategy, the researchers have developed a number of high-performance microbial cell factories that produce a variety of food and cosmetic compounds including natural substances like heme and zinc protoporphyrin IX compounds which can improve the flavor and color of synthetic meat, lycopene and β-carotene which are functional natural pigments that can be widely used in food and cosmetics, and methyl anthranilate, a grape-derived compound widely used to impart grape flavor in food and beverage manufacturing.
In this paper written upon invitation by Nature, the research team covered remarkable cases of microbial cell factory that can produce amino acids, proteins, fats and fatty acids, vitamins, flavors, pigments, alcohols, functional compounds and other food additives used in various foods and cosmetics and the companies that have successfully commercialized these microbial-derived materials Furthermore, the paper organized and presents systems metabolic engineering strategies that can spur the development of industrial microbial cell factories that can produce more diverse food and cosmetic compounds in an eco-friendly way with economic feasibility.
< Figure 1. Examples of production of food and cosmetic compounds using microbial cell factories >
For example, by producing proteins or amino acids with high nutritional value through non-edible biomass used as animal feed or fertilizer through the microbial fermentation process, it will contribute to the increase in production and stable supply of food around the world. Furthermore, by contributing to developing more viable alternative meat, further reducing dependence on animal protein, it can also contribute to reducing greenhouse gases and environmental pollution generated through livestock breeding or fish farming.
In addition, vanillin or methyl anthranilate, which give off vanilla or grape flavor, are widely added to various foods, but natural products isolated and refined from plants are low in production and high in production cost, so in most cases, petrochemicals substances derived from vanillin and methylanthranilic acid are added to food. These materials can also be produced through an eco-friendly and human-friendly method by borrowing the power of microorganisms.
Ethical and resource problems that arise in producing compounds like Calmin (cochineal pigment), a coloring added to various cosmetics and foods such as red lipstick and strawberry-flavored milk, which must be extracted from cochineal insects that live only in certain cacti. and Hyaluronic acid, which is widely consumed as a health supplement, but is only present in omega-3 fatty acids extracted from shark or fish livers, can also be resolved when they can be produced in an eco-friendly way using microorganisms.
KAIST Research Professor Kyeong Rok Choi, the first author of this paper, said, “In addition to traditional fermented foods such as kimchi and yogurt, foods produced with the help of microorganisms like cocoa butter, a base ingredient for chocolate that can only be obtained from fermented cacao beans, and monosodium glutamate, a seasoning produced through microbial fermentation are already familiar to us”. “In the future, we will be able to acquire a wider variety of foods and cosmetics even more easily produced in an eco-friendly and sustainable way in our daily lives through microbial cell factories.” he added.
< Figure 2. Systems metabolic engineering strategy to improve metabolic flow in microbial cell factories >
Distinguished Professor Sang Yup Lee said, “It is engineers’ mission to make the world a better place utilizing science and technology.” and added, “Continuous advancement and active use of systems metabolic engineering will contribute greatly to easing and resolving the problems arising from both the food crisis and the climate change."
This research was carried out as a part of the “Development of Protein Production Technology from Inorganic Substances through Control of Microbial Metabolism System Project” (Project Leader: Kyeong Rok Choi, KAIST Research Professor) of the the Center for Agricultural Microorganism and Enzyme (Director Pahn-Shick Chang) supported by the Rural Development Administration and the “Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project” (Project Leader: Sang Yup Lee, KAIST Distinguished Professor) of the Petroleum-Substitute Eco-friendly Chemical Technology Development Program supported by the Ministry of Science and ICT.
2023.07.28 View 9557
KAIST presents a microbial cell factory as a source of eco-friendly food and cosmetic coloring
Despite decades of global population growth, global food crisis seems to be at hand yet again because the food productivity is cut severely due to prolonged presence of abnormal weather from intensifying climate change and global food supply chain is deteriorated due to international conflicts such as wars exacerbating food shortages and nutritional inequality around the globe. At the same time, however, as awareness of the environment and sustainability rises, an increase in demand for more eco-friendly and high-quality food and beauty products is being observed not without a sense of irony. At a time like this, microorganisms are attracting attention as a key that can handle this couple of seemingly distant problems.
KAIST (President Kwang-Hyung Lee) announced on the 26th that Kyeong Rok Choi, a research professor of the Bioprocess Research Center and Sang Yup Lee, a Distinguished Professor of the Department of Chemical and Biomolecular Engineering, published a paper titled “Metabolic Engineering of Microorganisms for Food and Cosmetics Production” upon invitation by “Nature Reviews Bioengineering” to be published online published by Nature after peer review.
※ Paper title: Systems metabolic engineering of microorganisms for food and cosmetics production
※ Author information: Kyeong Rok Choi (first author) and Sang Yup Lee (corresponding author)
Systems metabolic engineering is a research field founded by Distinguished Professor Sang Yup Lee of KAIST to more effectively develop microbial cell factories, the core factor of the next-generation bio industry to replace the existing chemical industry that relies heavily on petroleum. By applying a systemic metabolic engineering strategy, the researchers have developed a number of high-performance microbial cell factories that produce a variety of food and cosmetic compounds including natural substances like heme and zinc protoporphyrin IX compounds which can improve the flavor and color of synthetic meat, lycopene and β-carotene which are functional natural pigments that can be widely used in food and cosmetics, and methyl anthranilate, a grape-derived compound widely used to impart grape flavor in food and beverage manufacturing.
In this paper written upon invitation by Nature, the research team covered remarkable cases of microbial cell factory that can produce amino acids, proteins, fats and fatty acids, vitamins, flavors, pigments, alcohols, functional compounds and other food additives used in various foods and cosmetics and the companies that have successfully commercialized these microbial-derived materials Furthermore, the paper organized and presents systems metabolic engineering strategies that can spur the development of industrial microbial cell factories that can produce more diverse food and cosmetic compounds in an eco-friendly way with economic feasibility.
< Figure 1. Examples of production of food and cosmetic compounds using microbial cell factories >
For example, by producing proteins or amino acids with high nutritional value through non-edible biomass used as animal feed or fertilizer through the microbial fermentation process, it will contribute to the increase in production and stable supply of food around the world. Furthermore, by contributing to developing more viable alternative meat, further reducing dependence on animal protein, it can also contribute to reducing greenhouse gases and environmental pollution generated through livestock breeding or fish farming.
In addition, vanillin or methyl anthranilate, which give off vanilla or grape flavor, are widely added to various foods, but natural products isolated and refined from plants are low in production and high in production cost, so in most cases, petrochemicals substances derived from vanillin and methylanthranilic acid are added to food. These materials can also be produced through an eco-friendly and human-friendly method by borrowing the power of microorganisms.
Ethical and resource problems that arise in producing compounds like Calmin (cochineal pigment), a coloring added to various cosmetics and foods such as red lipstick and strawberry-flavored milk, which must be extracted from cochineal insects that live only in certain cacti. and Hyaluronic acid, which is widely consumed as a health supplement, but is only present in omega-3 fatty acids extracted from shark or fish livers, can also be resolved when they can be produced in an eco-friendly way using microorganisms.
KAIST Research Professor Kyeong Rok Choi, the first author of this paper, said, “In addition to traditional fermented foods such as kimchi and yogurt, foods produced with the help of microorganisms like cocoa butter, a base ingredient for chocolate that can only be obtained from fermented cacao beans, and monosodium glutamate, a seasoning produced through microbial fermentation are already familiar to us”. “In the future, we will be able to acquire a wider variety of foods and cosmetics even more easily produced in an eco-friendly and sustainable way in our daily lives through microbial cell factories.” he added.
< Figure 2. Systems metabolic engineering strategy to improve metabolic flow in microbial cell factories >
Distinguished Professor Sang Yup Lee said, “It is engineers’ mission to make the world a better place utilizing science and technology.” and added, “Continuous advancement and active use of systems metabolic engineering will contribute greatly to easing and resolving the problems arising from both the food crisis and the climate change."
This research was carried out as a part of the “Development of Protein Production Technology from Inorganic Substances through Control of Microbial Metabolism System Project” (Project Leader: Kyeong Rok Choi, KAIST Research Professor) of the the Center for Agricultural Microorganism and Enzyme (Director Pahn-Shick Chang) supported by the Rural Development Administration and the “Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project” (Project Leader: Sang Yup Lee, KAIST Distinguished Professor) of the Petroleum-Substitute Eco-friendly Chemical Technology Development Program supported by the Ministry of Science and ICT.
2023.07.28 View 9557 -
 A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 11964
A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 11964 -
 Synthetic sRNAs to knockdown genes in medical and industrial bacteria
Bacteria are intimately involved in our daily lives. These microorganisms have been used in human history for food such as cheese, yogurt, and wine, In more recent years, through metabolic engineering, microorganisms been used extensively as microbial cell factories to manufacture plastics, feed for livestock, dietary supplements, and drugs. However, in addition to these bacteria that are beneficial to human lives, pathogens such as Pneumonia, Salmonella, and Staphylococcus that cause various infectious diseases are also ubiquitously present. It is important to be able to metabolically control these beneficial industrial bacteria for high value-added chemicals production and to manipulate harmful pathogens to suppress its pathogenic traits.
KAIST (President Kwang Hyung Lee) announced on the 10th that a research team led by Distinguished Professor Sang Yup Lee of the Department of Biochemical Engineering has developed a new sRNA tool that can effectively inhibit target genes in various bacteria, including both Gram-negative and Gram-positive bacteria. The research results were published online on April 24 in Nature Communications.
※ Thesis title: Targeted and high-throughput gene knockdown in diverse bacteria using synthetic sRNAs
※ Author information : Jae Sung Cho (co-1st), Dongsoo Yang (co-1st), Cindy Pricilia Surya Prabowo (co-author), Mohammad Rifqi Ghiffary (co-author), Taehee Han (co-author), Kyeong Rok Choi (co-author), Cheon Woo Moon (co-author), Hengrui Zhou (co-author), Jae Yong Ryu (co-author), Hyun Uk Kim (co-author) and Sang Yup Lee (corresponding author).
sRNA is an effective tool for synthesizing and regulating target genes in E. coli, but it has been difficult to apply to industrially useful Gram-positive bacteria such as Bacillus subtilis and Corynebacterium in addition to Gram-negative bacteria such as E. coli.
To address this issue, a research team led by Distinguished Professor Lee Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a new sRNA platform that can effectively suppress target genes in various bacteria, including both Gram-negative and positive bacteria. The research team surveyed thousands of microbial-derived sRNA systems in the microbial database, and eventually designated the sRNA system derived from 'Bacillus subtilis' that showed the highest gene knockdown efficiency, and designated it as “Broad-Host-Range sRNA”, or BHR-sRNA.
A similar well-known system is the CRISPR interference (CRISPRi) system, which is a modified CRISPR system that knocks down gene expression by suppressing the gene transcription process. However, the Cas9 protein in the CRISPRi system has a very high molecular weight, and there have been reports growth inhibition in bacteria. The BHR-sRNA system developed in this study did not affect bacterial growth while showing similar gene knockdown efficiencies to CRISPRi.
< Figure 1. a) Schematic illustration demonstrating the mechanism of syntetic sRNA b) Phylogenetic tree of the 16 Gram-negative and Gram-positive bacterial species tested for gene knockdown by the BHR-sRNA system. >
To validate the versatility of the BHR-sRNA system, 16 different gram-negative and gram-positive bacteria were selected and tested, where the BHR-sRNA system worked successfully in 15 of them. In addition, it was demonstrated that the gene knockdown capability was more effective than that of the existing E. coli-based sRNA system in 10 bacteria. The BHR-sRNA system proved to be a universal tool capable of effectively inhibiting gene expression in various bacteria.
In order to address the problem of antibiotic-resistant pathogens that have recently become more serious, the BHR-sRNA was demonstrated to suppress the pathogenicity by suppressing the gene producing the virulence factor. By using BHR-sRNA, biofilm formation, one of the factors resulting in antibiotic resistance, was inhibited by 73% in Staphylococcus epidermidis a pathogen that can cause hospital-acquired infections. Antibiotic resistance was also weakened by 58% in the pneumonia causing bacteria Klebsiella pneumoniae. In addition, BHR-sRNA was applied to industrial bacteria to develop microbial cell factories to produce high value-added chemicals with better production performance. Notably, superior industrial strains were constructed with the aid of BHR-sRNA to produce the following chemicals: valerolactam, a raw material for polyamide polymers, methyl-anthranilate, a grape-flavor food additive, and indigoidine, a blue-toned natural dye.
The BHR-sRNA developed through this study will help expedite the commercialization of bioprocesses to produce high value-added compounds and materials such as artificial meat, jet fuel, health supplements, pharmaceuticals, and plastics. It is also anticipated that to help eradicating antibiotic-resistant pathogens in preparation for another upcoming pandemic. “In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria” said Distinguished Professor Sang Yup Lee.
This work was supported by the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project from NRF supported by the Korean MSIT.
2023.05.10 View 8498
Synthetic sRNAs to knockdown genes in medical and industrial bacteria
Bacteria are intimately involved in our daily lives. These microorganisms have been used in human history for food such as cheese, yogurt, and wine, In more recent years, through metabolic engineering, microorganisms been used extensively as microbial cell factories to manufacture plastics, feed for livestock, dietary supplements, and drugs. However, in addition to these bacteria that are beneficial to human lives, pathogens such as Pneumonia, Salmonella, and Staphylococcus that cause various infectious diseases are also ubiquitously present. It is important to be able to metabolically control these beneficial industrial bacteria for high value-added chemicals production and to manipulate harmful pathogens to suppress its pathogenic traits.
KAIST (President Kwang Hyung Lee) announced on the 10th that a research team led by Distinguished Professor Sang Yup Lee of the Department of Biochemical Engineering has developed a new sRNA tool that can effectively inhibit target genes in various bacteria, including both Gram-negative and Gram-positive bacteria. The research results were published online on April 24 in Nature Communications.
※ Thesis title: Targeted and high-throughput gene knockdown in diverse bacteria using synthetic sRNAs
※ Author information : Jae Sung Cho (co-1st), Dongsoo Yang (co-1st), Cindy Pricilia Surya Prabowo (co-author), Mohammad Rifqi Ghiffary (co-author), Taehee Han (co-author), Kyeong Rok Choi (co-author), Cheon Woo Moon (co-author), Hengrui Zhou (co-author), Jae Yong Ryu (co-author), Hyun Uk Kim (co-author) and Sang Yup Lee (corresponding author).
sRNA is an effective tool for synthesizing and regulating target genes in E. coli, but it has been difficult to apply to industrially useful Gram-positive bacteria such as Bacillus subtilis and Corynebacterium in addition to Gram-negative bacteria such as E. coli.
To address this issue, a research team led by Distinguished Professor Lee Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a new sRNA platform that can effectively suppress target genes in various bacteria, including both Gram-negative and positive bacteria. The research team surveyed thousands of microbial-derived sRNA systems in the microbial database, and eventually designated the sRNA system derived from 'Bacillus subtilis' that showed the highest gene knockdown efficiency, and designated it as “Broad-Host-Range sRNA”, or BHR-sRNA.
A similar well-known system is the CRISPR interference (CRISPRi) system, which is a modified CRISPR system that knocks down gene expression by suppressing the gene transcription process. However, the Cas9 protein in the CRISPRi system has a very high molecular weight, and there have been reports growth inhibition in bacteria. The BHR-sRNA system developed in this study did not affect bacterial growth while showing similar gene knockdown efficiencies to CRISPRi.
< Figure 1. a) Schematic illustration demonstrating the mechanism of syntetic sRNA b) Phylogenetic tree of the 16 Gram-negative and Gram-positive bacterial species tested for gene knockdown by the BHR-sRNA system. >
To validate the versatility of the BHR-sRNA system, 16 different gram-negative and gram-positive bacteria were selected and tested, where the BHR-sRNA system worked successfully in 15 of them. In addition, it was demonstrated that the gene knockdown capability was more effective than that of the existing E. coli-based sRNA system in 10 bacteria. The BHR-sRNA system proved to be a universal tool capable of effectively inhibiting gene expression in various bacteria.
In order to address the problem of antibiotic-resistant pathogens that have recently become more serious, the BHR-sRNA was demonstrated to suppress the pathogenicity by suppressing the gene producing the virulence factor. By using BHR-sRNA, biofilm formation, one of the factors resulting in antibiotic resistance, was inhibited by 73% in Staphylococcus epidermidis a pathogen that can cause hospital-acquired infections. Antibiotic resistance was also weakened by 58% in the pneumonia causing bacteria Klebsiella pneumoniae. In addition, BHR-sRNA was applied to industrial bacteria to develop microbial cell factories to produce high value-added chemicals with better production performance. Notably, superior industrial strains were constructed with the aid of BHR-sRNA to produce the following chemicals: valerolactam, a raw material for polyamide polymers, methyl-anthranilate, a grape-flavor food additive, and indigoidine, a blue-toned natural dye.
The BHR-sRNA developed through this study will help expedite the commercialization of bioprocesses to produce high value-added compounds and materials such as artificial meat, jet fuel, health supplements, pharmaceuticals, and plastics. It is also anticipated that to help eradicating antibiotic-resistant pathogens in preparation for another upcoming pandemic. “In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria” said Distinguished Professor Sang Yup Lee.
This work was supported by the Development of Next-generation Biorefinery Platform Technologies for Leading Bio-based Chemicals Industry Project and the Development of Platform Technologies of Microbial Cell Factories for the Next-generation Biorefineries Project from NRF supported by the Korean MSIT.
2023.05.10 View 8498 -
 KAIST gearing up to train physician-scientists and BT Professionals joining hands with Boston-based organizations
KAIST (President Kwang Hyung Lee) announced on the 29th that it has signed MOUs with Massachusetts General Hospital, a founding member of the Mass General Brigham health care system and a world-class research-oriented hospital, and Moderna, a biotechnology company that developed a COVID-19 vaccine at the Langham Hotel in Boston, MA, USA on the morning of April 28th (local time). The signing ceremony was attended by officials from each institution joined by others headed by Minister LEE Young of the Korean Ministry of SMEs and Startups (MSS), and Commissioner LEE Insil of the Korean Intellectual Property Office.
< Photo 1. Photo from the Signing of MOU between KAIST-Harvard University Massachusetts General Hospital and KAIST-Moderna >
Mass General is the first and largest teaching hospital of Harvard Medical School in Boston, USA, and it is one of the most innovative hospitals in the world being the alma mater of more than 13 Nobel Prize winners and the home of the Mass General Research Institute, the world’s largest hospital-based research program that utilizes an annual research budget of more than $1.3 billion.
KAIST signed a general agreement to explore research and academic exchange with Mass General in September of last year and this MOU is a part of its follow-ups.
Mass General works with Harvard and the Massachusetts Institute of Technology (MIT), as well as local hospitals, to support students learn the theories of medicine and engineering, and gain rich clinical research experience.
Through this MOU, KAIST will explore cooperation with an innovative ecosystem created through the convergence of medicine and engineering. In particular, KAIST’s goal is to develop a Korean-style training program and implement a differentiated educational program when establishing the science and technology-oriented medical school in the future by further strengthening the science and engineering part of the training including a curriculum on artificial intelligence (AI) and the likes there of.
Also, in order to foster innovative physician-scientists, KAIST plans to pursue cooperation to develop programs for exchange of academic and human resources including programs for student and research exchanges and a program for students of the science and technology-oriented medical school at KAIST to have a chance to take part in practical training at Mass General.
David F.M. Brown, MD, Mass General President, said, “The collaboration with KAIST has a wide range of potentials, including advice on training of physician-scientists, academic and human resource exchanges, and vitalization of joint research by faculty from both institutions. Through this agreement, we will be able to actively contribute to global cooperation and achieve mutual goals.”
Meanwhile, an MOU between KAIST and Moderna was also held on the same day. Its main focus is to foster medical experts in cooperation with KAIST Graduate School of Medical Science and Engineering (GSMSE), and plans to cooperate in various ways in the future, including collaborating for development of vaccine and new drugs, virus research, joint mRNA research, and facilitation of technology commercialization.
In over 10 years since its inception, Moderna has transformed from a research-stage company advancing programs in the field of messenger RNA (mRNA) to an enterprise with a diverse clinical portfolio of vaccines and therapeutics across seven modalities. The Company has 48 programs in development across 45 development candidates, of which 38 are currently in active clinical trials.
“We are grateful to have laid a foundation for collaboration to foster industry experts with the Korea Advanced Institute of Science and Technology, a leader of science and technology innovation in Korea,” said Arpa Garay, Chief Commercial Officer, Moderna. “Based on our leadership and expertise in developing innovative mRNA vaccines and therapeutics, we hope to contribute to educating and collaborating with professionals in the bio-health field of Korea.“
President Kwang Hyung Lee of KAIST, said, “We deem this occasion to be of grave significance to be able to work closely with Massachusetts General Hospital, one of the world's best research-oriented hospitals, and Moderna, one of the most influential biomedical companies.”
President Lee continued, "On the basis of the collaboration with the two institutions, we will be able to bring up qualified physician-scientists and global leaders of the biomedical business who will solve problems of human health and their progress will in turn, accelerate the national R&D efforts in general and diversify the industry."
2023.04.29 View 15630
KAIST gearing up to train physician-scientists and BT Professionals joining hands with Boston-based organizations
KAIST (President Kwang Hyung Lee) announced on the 29th that it has signed MOUs with Massachusetts General Hospital, a founding member of the Mass General Brigham health care system and a world-class research-oriented hospital, and Moderna, a biotechnology company that developed a COVID-19 vaccine at the Langham Hotel in Boston, MA, USA on the morning of April 28th (local time). The signing ceremony was attended by officials from each institution joined by others headed by Minister LEE Young of the Korean Ministry of SMEs and Startups (MSS), and Commissioner LEE Insil of the Korean Intellectual Property Office.
< Photo 1. Photo from the Signing of MOU between KAIST-Harvard University Massachusetts General Hospital and KAIST-Moderna >
Mass General is the first and largest teaching hospital of Harvard Medical School in Boston, USA, and it is one of the most innovative hospitals in the world being the alma mater of more than 13 Nobel Prize winners and the home of the Mass General Research Institute, the world’s largest hospital-based research program that utilizes an annual research budget of more than $1.3 billion.
KAIST signed a general agreement to explore research and academic exchange with Mass General in September of last year and this MOU is a part of its follow-ups.
Mass General works with Harvard and the Massachusetts Institute of Technology (MIT), as well as local hospitals, to support students learn the theories of medicine and engineering, and gain rich clinical research experience.
Through this MOU, KAIST will explore cooperation with an innovative ecosystem created through the convergence of medicine and engineering. In particular, KAIST’s goal is to develop a Korean-style training program and implement a differentiated educational program when establishing the science and technology-oriented medical school in the future by further strengthening the science and engineering part of the training including a curriculum on artificial intelligence (AI) and the likes there of.
Also, in order to foster innovative physician-scientists, KAIST plans to pursue cooperation to develop programs for exchange of academic and human resources including programs for student and research exchanges and a program for students of the science and technology-oriented medical school at KAIST to have a chance to take part in practical training at Mass General.
David F.M. Brown, MD, Mass General President, said, “The collaboration with KAIST has a wide range of potentials, including advice on training of physician-scientists, academic and human resource exchanges, and vitalization of joint research by faculty from both institutions. Through this agreement, we will be able to actively contribute to global cooperation and achieve mutual goals.”
Meanwhile, an MOU between KAIST and Moderna was also held on the same day. Its main focus is to foster medical experts in cooperation with KAIST Graduate School of Medical Science and Engineering (GSMSE), and plans to cooperate in various ways in the future, including collaborating for development of vaccine and new drugs, virus research, joint mRNA research, and facilitation of technology commercialization.
In over 10 years since its inception, Moderna has transformed from a research-stage company advancing programs in the field of messenger RNA (mRNA) to an enterprise with a diverse clinical portfolio of vaccines and therapeutics across seven modalities. The Company has 48 programs in development across 45 development candidates, of which 38 are currently in active clinical trials.
“We are grateful to have laid a foundation for collaboration to foster industry experts with the Korea Advanced Institute of Science and Technology, a leader of science and technology innovation in Korea,” said Arpa Garay, Chief Commercial Officer, Moderna. “Based on our leadership and expertise in developing innovative mRNA vaccines and therapeutics, we hope to contribute to educating and collaborating with professionals in the bio-health field of Korea.“
President Kwang Hyung Lee of KAIST, said, “We deem this occasion to be of grave significance to be able to work closely with Massachusetts General Hospital, one of the world's best research-oriented hospitals, and Moderna, one of the most influential biomedical companies.”
President Lee continued, "On the basis of the collaboration with the two institutions, we will be able to bring up qualified physician-scientists and global leaders of the biomedical business who will solve problems of human health and their progress will in turn, accelerate the national R&D efforts in general and diversify the industry."
2023.04.29 View 15630 -
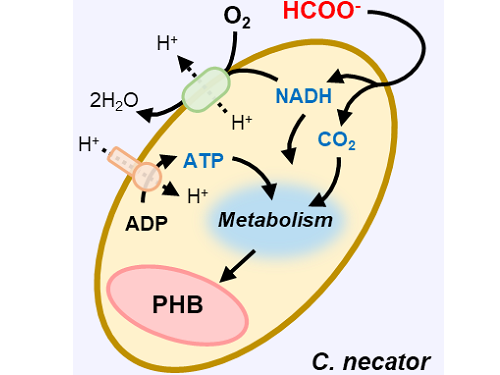 A biohybrid system to extract 20 times more bioplastic from CO2 developed by KAIST researchers
As the issues surrounding global climate change intensify, more attention and determined efforts are required to re-grasp the issue as a state of “crisis” and respond to it properly. Among the various methods of recycling CO2, the electrochemical CO2 conversion technology is a technology that can convert CO2 into useful chemical substances using electrical energy. Since it is easy to operate facilities and can use the electricity from renewable sources like the solar cells or the wind power, it has received a lot of attention as an eco-friendly technology can contribute to reducing greenhouse gases and achieve carbon neutrality.
KAIST (President Kwang Hyung Lee) announced on the 30th that the joint research team led by Professor Hyunjoo Lee and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering succeeded in developing a technology that produces bioplastics from CO2 with high efficiency by developing a hybrid system that interlinked the electrochemical CO2 conversion and microbial bio conversion methods together. The results of the research, which showed the world's highest productivity by more than 20 times compared to similar systems, were published online on March 27th in the "Proceedings of the National Academy of Sciences (PNAS)".
※ Paper title: Biohybrid CO2 electrolysis for the direct synthesis of polyesters from CO2
※ Author information: Jinkyu Lim (currently at Stanford Linear Accelerator Center, co-first author), So Young Choi (KAIST, co-first author), Jae Won Lee (KAIST, co-first author), Hyunjoo Lee (KAIST, corresponding author), Sang Yup Lee (KAIST, corresponding author)
For the efficient conversion of CO2, high-efficiency electrode catalysts and systems are actively being developed. As conversion products, only compounds containing one or up to three carbon atoms are produced on a limited basis. Compounds of one carbon, such as CO, formic acid, and ethylene, are produced with relatively high efficiency. Liquid compounds of several carbons, such as ethanol, acetic acid, and propanol, can also be produced by these systems, but due to the nature of the chemical reaction that requires more electrons, there are limitations involving the conversion efficiency and the product selection.
Accordingly, a joint research team led by Professor Hyunjoo Lee and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a technology to produce bioplastics from CO2 by linking electrochemical conversion technology with bioconversion method that uses microorganisms.
This electrochemical-bio hybrid system is in the form of having an electrolyzer, in which electrochemical conversion reactions occur, connected to a fermenter, in which microorganisms are cultured. When CO2 is converted to formic acid in the electrolyzer, and it is fed into the fermenter in which the microbes like the Cupriavidus necator, in this case, consumes the carbon source to produce polyhydroxyalkanoate (PHA), a microbial-derived bioplastic.
According to the research results of the existing hybrid concepts, there was a disadvantage of having low productivity or stopping at a non-continuous process due to problems of low efficiency of the electrolysis and irregular results arising from the culturing conditions of the microbes.
In order to overcome these problems, the joint research team made formic acid with a gas diffusion electrode using gaseous CO2. In addition, the team developed a 'physiologically compatible catholyte' that can be used as a culture medium for microorganisms as well as an electrolyte that allows the electrolysis to occur sufficiently without inhibiting the growth of microorganisms, without having to have a additional separation and purification process, which allowed the acide to be supplied directly to microorganisms.
Through this, the electrolyte solution containing formic acid made from CO2 enters the fermentation tank, is used for microbial culture, and enters the electrolyzer to be circulated, maximizing the utilization of the electrolyte solution and remaining formic acid. In addition, a filter was installed to ensure that only the electrolyte solution with any and all microorganisms that can affect the electrosis filtered out is supplied back to the electrolyzer, and that the microorganisms exist only in the fermenter, designing the two system to work well together with utmost efficiency.
Through the developed hybrid system, the produced bioplastic, poly-3-hydroxybutyrate (PHB), of up to 83% of the cell dry weight was produced from CO2, which produced 1.38g of PHB from a 4 cm2 electrode, which is the world's first gram(g) level production and is more than 20 times more productive than previous research. In addition, the hybrid system is expected to be applied to various industrial processes in the future as it shows promises of the continuous culture system.
The corresponding authors, Professor Hyunjoo Lee and Distinguished Professor Sang Yup Lee noted that “The results of this research are technologies that can be applied to the production of various chemical substances as well as bioplastics, and are expected to be used as key parts needed in achieving carbon neutrality in the future.”
This research was received and performed with the supports from the CO2 Reduction Catalyst and Energy Device Technology Development Project, the Heterogeneous Atomic Catalyst Control Project, and the Next-generation Biorefinery Source Technology Development Project to lead the Biochemical Industry of the Oil-replacement Eco-friendly Chemical Technology Development Program by the Ministry of Science and ICT.
Figure 1. Schematic diagram and photo of the biohybrid CO2 electrolysis system.
(A) A conceptual scheme and (B) a photograph of the biohybrid CO2 electrolysis system. (C) A detailed scheme of reaction inside the system. Gaseous CO2 was converted to formate in the electrolyzer, and the formate was converted to PHB by the cells in the fermenter. The catholyte was developed so that it is compatible with both CO2 electrolysis and fermentation and was continuously circulated.
2023.03.30 View 11876
A biohybrid system to extract 20 times more bioplastic from CO2 developed by KAIST researchers
As the issues surrounding global climate change intensify, more attention and determined efforts are required to re-grasp the issue as a state of “crisis” and respond to it properly. Among the various methods of recycling CO2, the electrochemical CO2 conversion technology is a technology that can convert CO2 into useful chemical substances using electrical energy. Since it is easy to operate facilities and can use the electricity from renewable sources like the solar cells or the wind power, it has received a lot of attention as an eco-friendly technology can contribute to reducing greenhouse gases and achieve carbon neutrality.
KAIST (President Kwang Hyung Lee) announced on the 30th that the joint research team led by Professor Hyunjoo Lee and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering succeeded in developing a technology that produces bioplastics from CO2 with high efficiency by developing a hybrid system that interlinked the electrochemical CO2 conversion and microbial bio conversion methods together. The results of the research, which showed the world's highest productivity by more than 20 times compared to similar systems, were published online on March 27th in the "Proceedings of the National Academy of Sciences (PNAS)".
※ Paper title: Biohybrid CO2 electrolysis for the direct synthesis of polyesters from CO2
※ Author information: Jinkyu Lim (currently at Stanford Linear Accelerator Center, co-first author), So Young Choi (KAIST, co-first author), Jae Won Lee (KAIST, co-first author), Hyunjoo Lee (KAIST, corresponding author), Sang Yup Lee (KAIST, corresponding author)
For the efficient conversion of CO2, high-efficiency electrode catalysts and systems are actively being developed. As conversion products, only compounds containing one or up to three carbon atoms are produced on a limited basis. Compounds of one carbon, such as CO, formic acid, and ethylene, are produced with relatively high efficiency. Liquid compounds of several carbons, such as ethanol, acetic acid, and propanol, can also be produced by these systems, but due to the nature of the chemical reaction that requires more electrons, there are limitations involving the conversion efficiency and the product selection.
Accordingly, a joint research team led by Professor Hyunjoo Lee and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST developed a technology to produce bioplastics from CO2 by linking electrochemical conversion technology with bioconversion method that uses microorganisms.
This electrochemical-bio hybrid system is in the form of having an electrolyzer, in which electrochemical conversion reactions occur, connected to a fermenter, in which microorganisms are cultured. When CO2 is converted to formic acid in the electrolyzer, and it is fed into the fermenter in which the microbes like the Cupriavidus necator, in this case, consumes the carbon source to produce polyhydroxyalkanoate (PHA), a microbial-derived bioplastic.
According to the research results of the existing hybrid concepts, there was a disadvantage of having low productivity or stopping at a non-continuous process due to problems of low efficiency of the electrolysis and irregular results arising from the culturing conditions of the microbes.
In order to overcome these problems, the joint research team made formic acid with a gas diffusion electrode using gaseous CO2. In addition, the team developed a 'physiologically compatible catholyte' that can be used as a culture medium for microorganisms as well as an electrolyte that allows the electrolysis to occur sufficiently without inhibiting the growth of microorganisms, without having to have a additional separation and purification process, which allowed the acide to be supplied directly to microorganisms.
Through this, the electrolyte solution containing formic acid made from CO2 enters the fermentation tank, is used for microbial culture, and enters the electrolyzer to be circulated, maximizing the utilization of the electrolyte solution and remaining formic acid. In addition, a filter was installed to ensure that only the electrolyte solution with any and all microorganisms that can affect the electrosis filtered out is supplied back to the electrolyzer, and that the microorganisms exist only in the fermenter, designing the two system to work well together with utmost efficiency.
Through the developed hybrid system, the produced bioplastic, poly-3-hydroxybutyrate (PHB), of up to 83% of the cell dry weight was produced from CO2, which produced 1.38g of PHB from a 4 cm2 electrode, which is the world's first gram(g) level production and is more than 20 times more productive than previous research. In addition, the hybrid system is expected to be applied to various industrial processes in the future as it shows promises of the continuous culture system.
The corresponding authors, Professor Hyunjoo Lee and Distinguished Professor Sang Yup Lee noted that “The results of this research are technologies that can be applied to the production of various chemical substances as well as bioplastics, and are expected to be used as key parts needed in achieving carbon neutrality in the future.”
This research was received and performed with the supports from the CO2 Reduction Catalyst and Energy Device Technology Development Project, the Heterogeneous Atomic Catalyst Control Project, and the Next-generation Biorefinery Source Technology Development Project to lead the Biochemical Industry of the Oil-replacement Eco-friendly Chemical Technology Development Program by the Ministry of Science and ICT.
Figure 1. Schematic diagram and photo of the biohybrid CO2 electrolysis system.
(A) A conceptual scheme and (B) a photograph of the biohybrid CO2 electrolysis system. (C) A detailed scheme of reaction inside the system. Gaseous CO2 was converted to formate in the electrolyzer, and the formate was converted to PHB by the cells in the fermenter. The catholyte was developed so that it is compatible with both CO2 electrolysis and fermentation and was continuously circulated.
2023.03.30 View 11876 -
 KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 12635
KAIST develops biocompatible adhesive applicable to hair transplants
Aside from being used as a new medical adhesive, the new material can be applied to developing a new method of hair transplants, which cannot be repeated multiple times using current method of implanting the wholly intact follicles into the skin.
Medical adhesives are materials that can be applied to various uses such as wound healing, hemostasis, vascular anastomosis, and tissue engineering, and is expected to contribute greatly to the development of minimally invasive surgery and organ transplants. However, adhesives with high adhesion, low toxicity, and capable of decomposing in the body are rare. Adhesives based on natural proteins, such as fibrin and collagen, have high biocompatibility but insufficient adhesive strength. Synthetic polymer adhesives based on urethane or acrylic have greater adhesion but do not decompose well and may cause an inflammatory reaction in the body.
A joint research team led by Professor Myungeun Seo and Professor Haeshin Lee from the KAIST Department of Chemistry developed a bio-friendly adhesive from biocompatible polymers using tannic acid, the source of astringency in wine.
The research team focused on tannic acid, a natural polyphenolic product. Tannic acid is a polyphenol present in large amounts in fruit peels, nuts, and cacao. It has a high affinity and coating ability on other substances, and we sense the astringent taste in wine when tannic acid sticks to the surface of our tongue. When tannic acid is mixed with hydrophilic polymers, they form coacervates, or small droplets of jelly-like fluids that sink. If the polymers used are biocompatible, the mixture can be applied as a medical adhesive with low toxicity. However, coacervates are fundamentally fluid-like and cannot withstand large forces, which limits their adhesive capabilities. Thus, while research to utilize it as an adhesive has been actively discussed, a biodegradable material exhibiting strong adhesion due to its high shear strength has not yet been developed.
The research team figured out a way to enhance adhesion by mixing two biocompatible FDA-approved polymers, polyethylene glycol (PEG) and polylactic acid (PLA). While PEG, which is used widely in eyedrops and cream, is hydrophilic, PLA, a well-known bioplastic derived from lactic acid, is insoluble in water. The team combined the two into a block copolymer, which forms hydrophilic PLA aggregates in water with PEG blocks surrounding them. A coacervate created by mixing the micelles and tannic acid would behave like a solid due to the hard PLA components, and show an elastic modulus improved by a thousand times compared to PEG, enabling it to withstand much greater force as an adhesive.
Figure 1. (Above) Principle of biodegradable adhesive made by mixing poly(ethylene glycol)-poly(lactic acid) diblock copolymer and tannic acid in water. Yellow coacervate is precipitated through hydrogen bonding between the block copolymer micelles and tannic acid, and exhibits adhesion. After heat treatment, hydrogen bonds are rearranged to further improve adhesion. (Bottom) Adhesion comparison. Compared to using poly(ethylene glycol) polymer (d), it can support 10 times more weight when using block copolymer (e) and 60 times more weight after heat treatment (f). The indicated G' values represent the elastic modulus of the material.
Furthermore, the research team observed that the material’s mechanical properties can be improved by over a hundred times through a heating and cooling process that is used to heat-treat metals. They also discovered that this is due to the enforced interactions between micelle and tannic acid arrays.
The research team used the fact that the material shows minimal irritation to the skin and decomposes well in the body to demonstrate its possible application as an adhesive for hair transplantation through an animal experiment. Professor Haeshin Lee, who has pioneered various application fields including medical adhesives, hemostatic agents, and browning shampoo, focused on the adhesive capacities and low toxicity of polyphenols like tannic acid, and now looks forward to it improving the limitations of current hair transplant methods, which still involve follicle transfer and are difficult to be repeated multiple times.
Figure 2. (a) Overview of a hair transplantation method using a biodegradable adhesive (right) compared to a conventional hair transplantation method (left) that transplants hair containing hair follicles. After applying an adhesive to the tip of the hair, it is fixed to the skin by implanting it through a subcutaneous injection, and repeated treatment is possible. (b) Initial animal test results. One day after 15 hair transplantation, 12 strands of hair remain. If you pull the 3 strands of hair, you can see that the whole body is pulled up, indicating that it is firmly implanted into the skin. All strands of hair applied without the new adhesive material fell off, and in the case of adhesive without heat treatment, the efficiency was 1/7.
This research was conducted by first co-authors Dr. Jongmin Park (currently a senior researcher at the Korea Research Institute of Chemical Technology) from Professor Myeongeun Seo’s team and Dr. Eunsook Park from Professor Haeshin Lee’s team in the KAIST Department of Chemistry, and through joint research with the teams led by Professor Hyungjun Kim from the KAIST Department of Chemistry and Professor Siyoung Choi from the Department of Chemical and Biomolecular Engineering. The research was published online on August 22 in the international journal Au (JACS Au) under the title Biodegradable Block Copolymer-Tannic Acid Glue.
This study was funded by the Support Research Under Protection Project of the National Research Foundation (NRF), Leading Research Center Support Project (Research Center for Multiscale Chiral Structure), Biodegradable Plastics Commercialization and Demonstration Project by the Ministry of Trade and Industry, and institutional funding from the Korea Research Institute of Chemical Technology.
2022.10.07 View 12635 -
 Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 15178
Phage resistant Escherichia coli strains developed to reduce fermentation failure
A genome engineering-based systematic strategy for developing phage resistant Escherichia coli strains has been successfully developed through the collaborative efforts of a team led by Professor Sang Yup Lee, Professor Shi Chen, and Professor Lianrong Wang. This study by Xuan Zou et al. was published in Nature Communications in August 2022 and featured in Nature Communications Editors’ Highlights. The collaboration by the School of Pharmaceutical Sciences at Wuhan University, the First Affiliated Hospital of Shenzhen University, and the KAIST Department of Chemical and Biomolecular Engineering has made an important advance in the metabolic engineering and fermentation industry as it solves a big problem of phage infection causing fermentation failure.
Systems metabolic engineering is a highly interdisciplinary field that has made the development of microbial cell factories to produce various bioproducts including chemicals, fuels, and materials possible in a sustainable and environmentally friendly way, mitigating the impact of worldwide resource depletion and climate change. Escherichia coli is one of the most important chassis microbial strains, given its wide applications in the bio-based production of a diverse range of chemicals and materials. With the development of tools and strategies for systems metabolic engineering using E. coli, a highly optimized and well-characterized cell factory will play a crucial role in converting cheap and readily available raw materials into products of great economic and industrial value.
However, the consistent problem of phage contamination in fermentation imposes a devastating impact on host cells and threatens the productivity of bacterial bioprocesses in biotechnology facilities, which can lead to widespread fermentation failure and immeasurable economic loss. Host-controlled defense systems can be developed into effective genetic engineering solutions to address bacteriophage contamination in industrial-scale fermentation; however, most of the resistance mechanisms only narrowly restrict phages and their effect on phage contamination will be limited.
There have been attempts to develop diverse abilities/systems for environmental adaptation or antiviral defense. The team’s collaborative efforts developed a new type II single-stranded DNA phosphorothioation (Ssp) defense system derived from E. coli 3234/A, which can be used in multiple industrial E. coli strains (e.g., E. coli K-12, B and W) to provide broad protection against various types of dsDNA coliphages. Furthermore, they developed a systematic genome engineering strategy involving the simultaneous genomic integration of the Ssp defense module and mutations in components that are essential to the phage life cycle. This strategy can be used to transform E. coli hosts that are highly susceptible to phage attack into strains with powerful restriction effects on the tested bacteriophages. This endows hosts with strong resistance against a wide spectrum of phage infections without affecting bacterial growth and normal physiological function. More importantly, the resulting engineered phage-resistant strains maintained the capabilities of producing the desired chemicals and recombinant proteins even under high levels of phage cocktail challenge, which provides crucial protection against phage attacks.
This is a major step forward, as it provides a systematic solution for engineering phage-resistant bacterial strains, especially industrial bioproduction strains, to protect cells from a wide range of bacteriophages. Considering the functionality of this engineering strategy with diverse E. coli strains, the strategy reported in this study can be widely extended to other bacterial species and industrial applications, which will be of great interest to researchers in academia and industry alike.
Fig. A schematic model of the systematic strategy for engineering phage-sensitive industrial E. coli strains into strains with broad antiphage activities. Through the simultaneous genomic integration of a DNA phosphorothioation-based Ssp defense module and mutations of components essential for the phage life cycle, the engineered E. coli strains show strong resistance against diverse phages tested and maintain the capabilities of producing example recombinant proteins, even under high levels of phage cocktail challenge.
2022.08.23 View 15178 -
 Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 16472
Interactive Map of Metabolical Synthesis of Chemicals
An interactive map that compiled the chemicals produced by biological, chemical and combined reactions has been distributed on the web
- A team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering, organized and distributed an all-inclusive listing of chemical substances that can be synthesized using microorganisms
- It is expected to be used by researchers around the world as it enables easy assessment of the synthetic pathway through the web.
A research team comprised of Woo Dae Jang, Gi Bae Kim, and Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported an interactive metabolic map of bio-based chemicals. Their research paper “An interactive metabolic map of bio-based chemicals” was published online in Trends in Biotechnology on August 10, 2022.
As a response to rapid climate change and environmental pollution, research on the production of petrochemical products using microorganisms is receiving attention as a sustainable alternative to existing methods of productions. In order to synthesize various chemical substances, materials, and fuel using microorganisms, it is necessary to first construct the biosynthetic pathway toward desired product by exploration and discovery and introduce them into microorganisms. In addition, in order to efficiently synthesize various chemical substances, it is sometimes necessary to employ chemical methods along with bioengineering methods using microorganisms at the same time. For the production of non-native chemicals, novel pathways are designed by recruiting enzymes from heterologous sources or employing enzymes designed though rational engineering, directed evolution, or ab initio design.
The research team had completed a map of chemicals which compiled all available pathways of biological and/or chemical reactions that lead to the production of various bio-based chemicals back in 2019 and published the map in Nature Catalysis. The map was distributed in the form of a poster to industries and academia so that the synthesis paths of bio-based chemicals could be checked at a glance.
The research team has expanded the bio-based chemicals map this time in the form of an interactive map on the web so that anyone with internet access can quickly explore efficient paths to synthesize desired products. The web-based map provides interactive visual tools to allow interactive visualization, exploration, and analysis of complex networks of biological and/or chemical reactions toward the desired products. In addition, the reported paper also discusses the production of natural compounds that are used for diverse purposes such as food and medicine, which will help designing novel pathways through similar approaches or by exploiting the promiscuity of enzymes described in the map. The published bio-based chemicals map is also available at http://systemsbiotech.co.kr.
The co-first authors, Dr. Woo Dae Jang and Ph.D. student Gi Bae Kim, said, “We conducted this study to address the demand for updating the previously distributed chemicals map and enhancing its versatility.” “The map is expected to be utilized in a variety of research and in efforts to set strategies and prospects for chemical production incorporating bio and chemical methods that are detailed in the map.”
Distinguished Professor Sang Yup Lee said, “The interactive bio-based chemicals map is expected to help design and optimization of the metabolic pathways for the biosynthesis of target chemicals together with the strategies of chemical conversions, serving as a blueprint for developing further ideas on the production of desired chemicals through biological and/or chemical reactions.”
The interactive metabolic map of bio-based chemicals.
2022.08.11 View 16472