Sang+Yup+Lee
-
 Development of a Highly-Accurate Computational Model of Human Metabolism
A research team from KAIST developed a computational framework that enables the reconstruction of a comprehensive computational model of human metabolism, which allows for an accurate prediction of personal metabolic features (or phenotypes).
Understanding personal metabolic phenotypes allows us to design effective therapeutic strategies for various chronic and infectious diseases. A human computational model called the genome-scale metabolic model (GEM) contains information on thousands of metabolic genes and their corresponding reactions and metabolites, and has played an important role in predicting metabolic phenotypes. Although several versions of human GEMs have been released, they had room for further development, especially as to incorporating biological information coming from a human genetics mechanism called “alternative splicing.” Alternative splicing is a genetic mechanism that allows a gene to give rise to multiple reactions, and is strongly associated with pathology.
To tackle this problem, Jae Yong Ryu (a Ph.D. student), Dr. Hyun Uk Kim (Research Fellow), and Distinguished Professor Sang Yup Lee, all from the Department of Chemical and Biomolecular Engineering at KAIST, developed a computational framework that systematically generates metabolic reactions, and adds them to the human GEM. The resulting human GEM was demonstrated to accurately predict metabolic phenotypes under varied environmental conditions. The research results were published online in Proceedings of the National Academy of Sciences (PNAS) on October 24, 2017, under the title “Framework and resource for more than 11,000 gene-transcript-protein-reaction associations in human metabolism.”
The research team first updated the biological contents of a previous version of the human GEM. The updated biological contents include metabolic genes and their corresponding metabolites and reactions. In particular, metabolic reactions catalyzed by already-known protein isoforms were additionally incorporated into the human GEM; protein isoforms are multiple variants of proteins generated from individual genes through the alternative splicing process. Each protein isoform is often responsible for the operation of a metabolic reaction. Although multiple protein isoforms generated from one gene can play different functions by having different sets of protein domains and/or subcellular localizations, such information was not properly considered in previous versions of human GEMs.
Upon the initial update of the human GEM, named Recon 2M.1, the research team subsequently implemented a computational framework that systematically generates information on Gene-Transcript-Protein-Reaction Associations (GeTPRA) in order to identify protein isoforms that were previously not identified. This framework was developed in this study. As a result of the implementation of the framework for GeTPRA, more than 11,000 GeTPRA were automatically predicted, and thoroughly validated. Additional metabolic reactions were then added to Recon 2M.1 based on the predicted GeTPRA for the previously uncharacterized protein isoforms; Recon 2M.1 was renamed Recon 2M.2 from this upgrade.
Finally, Recon 2M.2 was integrated with 446 sets of personal biological data (RNA-Seq data) in order to build patient-specific cancer models. These patient-specific cancer models were used to predict cancer metabolism activities and anticancer targets.
The development of a new version of human GEMs along with the computational framework for GeTPRA is expected to boost studies in fundamental human genetics and medicine. Model files of the human GEMs Recon 2M.1 and 2M.2, a full list of the GeTPRA and the source code for the computational framework to predict the GeTPRA are all available as part of the publication of this study.
Distinguished Professor Lee said, “The predicted GeTPRA from the computational framework is expected to serve as a guideline for future experiments on human genetics and biochemistry, whereas the resulting Recon 2M.2 can be used to predict drug targets for various human diseases.”
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556 and NRF-2012M1A2A2026557) from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
(Figure 1:A scheme of Recon 2M.1 development and its use in reconstructing personal genome-scale metabolic models (GEMs). (A) A concept of alternative splicing of human genes and its use in Gene-Transcript-Protein-Reaction Associations (GeTPRA) of Recon 2M.1. (B) A procedure of systematic refinement of the Recon 2Q. Recon 2Q is one of the previously released human GEMs. Biochemically inconsistent reactions include unbalanced, artificial, blocked, and/or redundant reactions. Iterative manual curation was conducted while validating the Recon 2M.1. (C) Reconstruction of cancer patient-specific GEMs using Recon 2M.1 for further simulation studies. In this study, personal biological data (RNA-Seq data) were obtained from The Cancer Genome Atlas (TCGA; https://cancergenome.nih.gov/ ) across the ten cancer types.
(Figure 2: Computational framework for the systematic generation of Gene-Transcript-Protein-Reaction Associations (GeTPRA; red box in the flowchart). Peptide sequences of metabolic genes defined in Recon 2M.1 were retrieved from a database called Ensembl. EC numbers and subcellular localizations of all the protein isoforms of metabolic genes in Recon 2M.1 were predicted using software programs EFICAz2.5 and Wolf PSort, respectively. Information on the newly predicted GeTPRA was systematically incorporated into the Recon 2M.1, thereby resulting in Recon 2M.2.)
2017.10.25 View 10884
Development of a Highly-Accurate Computational Model of Human Metabolism
A research team from KAIST developed a computational framework that enables the reconstruction of a comprehensive computational model of human metabolism, which allows for an accurate prediction of personal metabolic features (or phenotypes).
Understanding personal metabolic phenotypes allows us to design effective therapeutic strategies for various chronic and infectious diseases. A human computational model called the genome-scale metabolic model (GEM) contains information on thousands of metabolic genes and their corresponding reactions and metabolites, and has played an important role in predicting metabolic phenotypes. Although several versions of human GEMs have been released, they had room for further development, especially as to incorporating biological information coming from a human genetics mechanism called “alternative splicing.” Alternative splicing is a genetic mechanism that allows a gene to give rise to multiple reactions, and is strongly associated with pathology.
To tackle this problem, Jae Yong Ryu (a Ph.D. student), Dr. Hyun Uk Kim (Research Fellow), and Distinguished Professor Sang Yup Lee, all from the Department of Chemical and Biomolecular Engineering at KAIST, developed a computational framework that systematically generates metabolic reactions, and adds them to the human GEM. The resulting human GEM was demonstrated to accurately predict metabolic phenotypes under varied environmental conditions. The research results were published online in Proceedings of the National Academy of Sciences (PNAS) on October 24, 2017, under the title “Framework and resource for more than 11,000 gene-transcript-protein-reaction associations in human metabolism.”
The research team first updated the biological contents of a previous version of the human GEM. The updated biological contents include metabolic genes and their corresponding metabolites and reactions. In particular, metabolic reactions catalyzed by already-known protein isoforms were additionally incorporated into the human GEM; protein isoforms are multiple variants of proteins generated from individual genes through the alternative splicing process. Each protein isoform is often responsible for the operation of a metabolic reaction. Although multiple protein isoforms generated from one gene can play different functions by having different sets of protein domains and/or subcellular localizations, such information was not properly considered in previous versions of human GEMs.
Upon the initial update of the human GEM, named Recon 2M.1, the research team subsequently implemented a computational framework that systematically generates information on Gene-Transcript-Protein-Reaction Associations (GeTPRA) in order to identify protein isoforms that were previously not identified. This framework was developed in this study. As a result of the implementation of the framework for GeTPRA, more than 11,000 GeTPRA were automatically predicted, and thoroughly validated. Additional metabolic reactions were then added to Recon 2M.1 based on the predicted GeTPRA for the previously uncharacterized protein isoforms; Recon 2M.1 was renamed Recon 2M.2 from this upgrade.
Finally, Recon 2M.2 was integrated with 446 sets of personal biological data (RNA-Seq data) in order to build patient-specific cancer models. These patient-specific cancer models were used to predict cancer metabolism activities and anticancer targets.
The development of a new version of human GEMs along with the computational framework for GeTPRA is expected to boost studies in fundamental human genetics and medicine. Model files of the human GEMs Recon 2M.1 and 2M.2, a full list of the GeTPRA and the source code for the computational framework to predict the GeTPRA are all available as part of the publication of this study.
Distinguished Professor Lee said, “The predicted GeTPRA from the computational framework is expected to serve as a guideline for future experiments on human genetics and biochemistry, whereas the resulting Recon 2M.2 can be used to predict drug targets for various human diseases.”
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556 and NRF-2012M1A2A2026557) from the Ministry of Science and ICT through the National Research Foundation (NRF) of Korea.
(Figure 1:A scheme of Recon 2M.1 development and its use in reconstructing personal genome-scale metabolic models (GEMs). (A) A concept of alternative splicing of human genes and its use in Gene-Transcript-Protein-Reaction Associations (GeTPRA) of Recon 2M.1. (B) A procedure of systematic refinement of the Recon 2Q. Recon 2Q is one of the previously released human GEMs. Biochemically inconsistent reactions include unbalanced, artificial, blocked, and/or redundant reactions. Iterative manual curation was conducted while validating the Recon 2M.1. (C) Reconstruction of cancer patient-specific GEMs using Recon 2M.1 for further simulation studies. In this study, personal biological data (RNA-Seq data) were obtained from The Cancer Genome Atlas (TCGA; https://cancergenome.nih.gov/ ) across the ten cancer types.
(Figure 2: Computational framework for the systematic generation of Gene-Transcript-Protein-Reaction Associations (GeTPRA; red box in the flowchart). Peptide sequences of metabolic genes defined in Recon 2M.1 were retrieved from a database called Ensembl. EC numbers and subcellular localizations of all the protein isoforms of metabolic genes in Recon 2M.1 were predicted using software programs EFICAz2.5 and Wolf PSort, respectively. Information on the newly predicted GeTPRA was systematically incorporated into the Recon 2M.1, thereby resulting in Recon 2M.2.)
2017.10.25 View 10884 -
 Bio-based p-Xylene Oxidation into Terephthalic Acid by Engineered E.coli
KAIST researchers have established an efficient biocatalytic system to produce terephthalic acid (TPA) from p-xylene (pX). It will allow this industrially important bulk chemical to be made available in a more environmentally-friendly manner.
The research team developed metabolically engineered Escherichia coli (E.coli) to biologically transform pX into TPA, a chemical necessary in the manufacturing of polyethylene terephthalate (PET). This biocatalysis system represents a greener and more efficient alternative to the traditional chemical methods for TPA production. This research, headed by Distinguished Professor Sang Yup Lee, was published in Nature Communications on May 31.
The research team utilized a metabolic engineering and synthetic biology approach to develop a recombinant microorganism that can oxidize pX into TPA using microbial fermentation. TPA is a globally important chemical commodity for manufacturing PET. It can be applied to manufacture plastic bottles, clothing fibers, films, and many other products. Currently, TPA is produced from pX oxidation through an industrially well-known chemical process (with a typical TPA yield of over 95 mol%), which shows, however, such drawbacks as intensive energy requirements at high temperatures and pressure, usage of heavy metal catalysts, and the unavoidable byproduct formation of 4-carboxybenzaldehyde.
The research team designed and constructed a synthetic metabolic pathway by incorporating the upper xylene degradation pathway of Pseudomonas putida F1 and the lower p-toluene sulfonate pathway of Comamonas testosteroni T-2, which successfully produced TPA from pX in small-scale cultures, with the formation of p-toluate (pTA) as the major byproduct. The team further optimized the pathway gene expression levels by using a synthetic biology toolkit, which gave the final engineered E. coli strain showing increased TPA production and the complete elimination of the byproduct.
Using this best-performing strain, the team designed an elegant two-phase (aqueous/organic) fermentation system for TPA production on a larger scale, where pX was supplied in the organic phase. Through a number of optimization steps, the team ultimately achieved production of 13.3 g TPA from 8.8 g pX, which represented an extraordinary yield of 97 mol%.
The team has developed a microbial biotechnology application which is reportedly the first successful example of the bio-based production of TPA from pX by the microbial fermentation of engineered E. coli. This bio-based TPA technology presents several advantages such as ambient reaction temperature and pressure, no use of heavy metals or other toxic chemicals, the removable of byproduct formation, and it is 100% environmentally compatible.
Professor Lee said, “We presented promising biotechnology for producing large amounts of the commodity chemical TPA, which is used for PET manufacturing, through metabolically engineered gut bacterium. Our research is meaningful in that it demonstrates the feasibility of the biotechnological production of bulk chemicals, and if reproducible when up-scaled, it will represent a breakthrough in hydrocarbon bioconversions.”
Ph.D. candidate Zi Wei Luo is the first author of this research (DOI:10.1038/ncomms15689).The research was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT & Future Planning through the National Research Foundation of Korea.
Figure: Biotransformation of pX into TPA by engineered E. coli.
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced TPA from pX. The engineered E. coli was developed through reconstituting a synthetic metabolic pathway for pX conversion to TPA and optimized for increased TPA yield and byproduct elimination. Two-phase partitioning fermentation system was developed for demonstrating the feasibility of large-scale production of TPA from pX using the engineered E. coli strains, where pX was supplied in the organic phase and TPA was produced in the aqueous phase.
2017.06.05 View 12759
Bio-based p-Xylene Oxidation into Terephthalic Acid by Engineered E.coli
KAIST researchers have established an efficient biocatalytic system to produce terephthalic acid (TPA) from p-xylene (pX). It will allow this industrially important bulk chemical to be made available in a more environmentally-friendly manner.
The research team developed metabolically engineered Escherichia coli (E.coli) to biologically transform pX into TPA, a chemical necessary in the manufacturing of polyethylene terephthalate (PET). This biocatalysis system represents a greener and more efficient alternative to the traditional chemical methods for TPA production. This research, headed by Distinguished Professor Sang Yup Lee, was published in Nature Communications on May 31.
The research team utilized a metabolic engineering and synthetic biology approach to develop a recombinant microorganism that can oxidize pX into TPA using microbial fermentation. TPA is a globally important chemical commodity for manufacturing PET. It can be applied to manufacture plastic bottles, clothing fibers, films, and many other products. Currently, TPA is produced from pX oxidation through an industrially well-known chemical process (with a typical TPA yield of over 95 mol%), which shows, however, such drawbacks as intensive energy requirements at high temperatures and pressure, usage of heavy metal catalysts, and the unavoidable byproduct formation of 4-carboxybenzaldehyde.
The research team designed and constructed a synthetic metabolic pathway by incorporating the upper xylene degradation pathway of Pseudomonas putida F1 and the lower p-toluene sulfonate pathway of Comamonas testosteroni T-2, which successfully produced TPA from pX in small-scale cultures, with the formation of p-toluate (pTA) as the major byproduct. The team further optimized the pathway gene expression levels by using a synthetic biology toolkit, which gave the final engineered E. coli strain showing increased TPA production and the complete elimination of the byproduct.
Using this best-performing strain, the team designed an elegant two-phase (aqueous/organic) fermentation system for TPA production on a larger scale, where pX was supplied in the organic phase. Through a number of optimization steps, the team ultimately achieved production of 13.3 g TPA from 8.8 g pX, which represented an extraordinary yield of 97 mol%.
The team has developed a microbial biotechnology application which is reportedly the first successful example of the bio-based production of TPA from pX by the microbial fermentation of engineered E. coli. This bio-based TPA technology presents several advantages such as ambient reaction temperature and pressure, no use of heavy metals or other toxic chemicals, the removable of byproduct formation, and it is 100% environmentally compatible.
Professor Lee said, “We presented promising biotechnology for producing large amounts of the commodity chemical TPA, which is used for PET manufacturing, through metabolically engineered gut bacterium. Our research is meaningful in that it demonstrates the feasibility of the biotechnological production of bulk chemicals, and if reproducible when up-scaled, it will represent a breakthrough in hydrocarbon bioconversions.”
Ph.D. candidate Zi Wei Luo is the first author of this research (DOI:10.1038/ncomms15689).The research was supported by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) of the Ministry of Science, ICT & Future Planning through the National Research Foundation of Korea.
Figure: Biotransformation of pX into TPA by engineered E. coli.
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced TPA from pX. The engineered E. coli was developed through reconstituting a synthetic metabolic pathway for pX conversion to TPA and optimized for increased TPA yield and byproduct elimination. Two-phase partitioning fermentation system was developed for demonstrating the feasibility of large-scale production of TPA from pX using the engineered E. coli strains, where pX was supplied in the organic phase and TPA was produced in the aqueous phase.
2017.06.05 View 12759 -
 Mystery of Biological Plastic Synthesis Machinery Unveiled
Plastics and other polymers are used every day. These polymers are mostly made from fossil resources by refining petrochemicals. On the other hand, many microorganisms naturally synthesize polyesters known as polyhydroxyalkanoates (PHAs) as distinct granules inside cells.
PHAs are a family of microbial polyesters that have attracted much attention as biodegradable and biocompatible plastics and elastomers that can substitute petrochemical counterparts. There have been numerous papers and patents on gene cloning and metabolic engineering of PHA biosynthetic machineries, biochemical studies, and production of PHAs; simple Google search with “polyhydroxyalkanoates” yielded returns of 223,000 document pages. PHAs have always been considered amazing examples of biological polymer synthesis. It is astounding to see PHAs of 500 kDa to sometimes as high as 10,000 kDa can be synthesized in vivo by PHA synthase, the key polymerizing enzyme in PHA biosynthesis. They have attracted great interest in determining the crystal structure of PHA synthase over the last 30 years, but unfortunately without success. Thus, the characteristics and molecular mechanisms of PHA synthase were under a dark veil.
In two papers published back-to-back in Biotechnology Journal online on November 30, 2016, a Korean research team led by Professor Kyung-Jin Kim at Kyungpook National University and Distinguished Professor Sang Yup Lee at the Korea Advanced Institute of Science and Technology (KAIST) described the crystal structure of PHA synthase from Ralstonia eutropha, the best studied bacterium for PHA production, and reported the structural basis for the detailed molecular mechanisms of PHA biosynthesis. The crystal structure has been deposited to Protein Data Bank in February 2016. After deciphering the crystal structure of the catalytic domain of PHA synthase, in addition to other structural studies on whole enzyme and related proteins, the research team also performed experiments to elucidate the mechanisms of the enzyme reaction, validating detailed structures, enzyme engineering, and also N-terminal domain studies among others.
Through several biochemical studies based on crystal structure, the authors show that PHA synthase exists as a dimer and is divided into two distinct domains, the N-terminal domain (RePhaC1ND) and the C-terminal domain (RePhaC1CD). The RePhaC1CD catalyzes the polymerization reaction via a non-processive ping-pong mechanism using a Cys-His-Asp catalytic triad. The two catalytic sites of the RePhaC1CD dimer are positioned 33.4 Å apart, suggesting that the polymerization reaction occurs independently at each site. This study also presents the structure-based mechanisms for substrate specificities of various PHA synthases from different classes.
Professor Sang Yup Lee, who has worked on this topic for more than 20 years, said,
“The results and information presented in these two papers have long been awaited not only in the PHA community, but also metabolic engineering, bacteriology/microbiology, and in general biological sciences communities. The structural information on PHA synthase together with the recently deciphered reaction mechanisms will be valuable for understanding the detailed mechanisms of biosynthesizing this important energy/redox storage material, and also for the rational engineering of PHA synthases to produce designer bioplastics from various monomers more efficiently.”
Indeed, these two papers published in Biotechnology Journal finally reveal the 30-year mystery of machinery of biological polyester synthesis, and will serve as the essential compass in creating designer and more efficient bioplastic machineries.
References:
Jieun Kim, Yeo-Jin Kim, So Young Choi, Sang Yup Lee and Kyung-Jin Kim. “Crystal structure of Ralstonia eutropha polyhydroxyalkanoate synthase C-terminal domain and reaction mechanisms” Biotechnology Journal DOI: 10.1002/biot.201600648
http://onlinelibrary.wiley.com/doi/10.1002/biot.201600648/abstract
Yeo-Jin Kim, So Young Choi, Jieun Kim, Kyeong Sik Jin, Sang Yup Lee and Kyung-Jin Kim. “Structure and function of the N-terminal domain of Ralstonia eutropha polyhydroxyalkanoate synthase, and the proposed structure and mechanisms of the whole enzyme” Biotechnology Journal DOI: 10.1002/biot.201600649
http://onlinelibrary.wiley.com/doi/10.1002/biot.201600649/abstract
2016.12.02 View 11312
Mystery of Biological Plastic Synthesis Machinery Unveiled
Plastics and other polymers are used every day. These polymers are mostly made from fossil resources by refining petrochemicals. On the other hand, many microorganisms naturally synthesize polyesters known as polyhydroxyalkanoates (PHAs) as distinct granules inside cells.
PHAs are a family of microbial polyesters that have attracted much attention as biodegradable and biocompatible plastics and elastomers that can substitute petrochemical counterparts. There have been numerous papers and patents on gene cloning and metabolic engineering of PHA biosynthetic machineries, biochemical studies, and production of PHAs; simple Google search with “polyhydroxyalkanoates” yielded returns of 223,000 document pages. PHAs have always been considered amazing examples of biological polymer synthesis. It is astounding to see PHAs of 500 kDa to sometimes as high as 10,000 kDa can be synthesized in vivo by PHA synthase, the key polymerizing enzyme in PHA biosynthesis. They have attracted great interest in determining the crystal structure of PHA synthase over the last 30 years, but unfortunately without success. Thus, the characteristics and molecular mechanisms of PHA synthase were under a dark veil.
In two papers published back-to-back in Biotechnology Journal online on November 30, 2016, a Korean research team led by Professor Kyung-Jin Kim at Kyungpook National University and Distinguished Professor Sang Yup Lee at the Korea Advanced Institute of Science and Technology (KAIST) described the crystal structure of PHA synthase from Ralstonia eutropha, the best studied bacterium for PHA production, and reported the structural basis for the detailed molecular mechanisms of PHA biosynthesis. The crystal structure has been deposited to Protein Data Bank in February 2016. After deciphering the crystal structure of the catalytic domain of PHA synthase, in addition to other structural studies on whole enzyme and related proteins, the research team also performed experiments to elucidate the mechanisms of the enzyme reaction, validating detailed structures, enzyme engineering, and also N-terminal domain studies among others.
Through several biochemical studies based on crystal structure, the authors show that PHA synthase exists as a dimer and is divided into two distinct domains, the N-terminal domain (RePhaC1ND) and the C-terminal domain (RePhaC1CD). The RePhaC1CD catalyzes the polymerization reaction via a non-processive ping-pong mechanism using a Cys-His-Asp catalytic triad. The two catalytic sites of the RePhaC1CD dimer are positioned 33.4 Å apart, suggesting that the polymerization reaction occurs independently at each site. This study also presents the structure-based mechanisms for substrate specificities of various PHA synthases from different classes.
Professor Sang Yup Lee, who has worked on this topic for more than 20 years, said,
“The results and information presented in these two papers have long been awaited not only in the PHA community, but also metabolic engineering, bacteriology/microbiology, and in general biological sciences communities. The structural information on PHA synthase together with the recently deciphered reaction mechanisms will be valuable for understanding the detailed mechanisms of biosynthesizing this important energy/redox storage material, and also for the rational engineering of PHA synthases to produce designer bioplastics from various monomers more efficiently.”
Indeed, these two papers published in Biotechnology Journal finally reveal the 30-year mystery of machinery of biological polyester synthesis, and will serve as the essential compass in creating designer and more efficient bioplastic machineries.
References:
Jieun Kim, Yeo-Jin Kim, So Young Choi, Sang Yup Lee and Kyung-Jin Kim. “Crystal structure of Ralstonia eutropha polyhydroxyalkanoate synthase C-terminal domain and reaction mechanisms” Biotechnology Journal DOI: 10.1002/biot.201600648
http://onlinelibrary.wiley.com/doi/10.1002/biot.201600648/abstract
Yeo-Jin Kim, So Young Choi, Jieun Kim, Kyeong Sik Jin, Sang Yup Lee and Kyung-Jin Kim. “Structure and function of the N-terminal domain of Ralstonia eutropha polyhydroxyalkanoate synthase, and the proposed structure and mechanisms of the whole enzyme” Biotechnology Journal DOI: 10.1002/biot.201600649
http://onlinelibrary.wiley.com/doi/10.1002/biot.201600649/abstract
2016.12.02 View 11312 -
 Unveiling the Distinctive Features of Industrial Microorganism
KAIST researchers have sequenced the whole genome of Clostridium tyrobutyricum, which has a higher tolerance to toxic chemicals, such as 1-butanol, compared to other clostridial bacterial strains.
Clostridium tyrobutyricum, a Gram-positive, anaerobic spore-forming bacterium, is considered a promising industrial host strain for the production of various chemicals including butyric acid which has many applications in different industries such as a precursor to biofuels. Despite such potential, C. tyrobutyricum has received little attention, mainly due to a limited understanding of its genotypic and metabolic characteristics at the genome level.
A Korean research team headed by Distinguished Professor Sang Yup Lee of the Chemical and Biomolecular Engineering Department at the Korea Advanced Institute of Science and Technology (KAIST) deciphered the genome sequence of C. tyrobutyricum and its proteome profiles during the course of batch fermentation. As a result, the research team learned that the bacterium is not only capable of producing a large amount of butyric acid but also can tolerate toxic compounds such as 1-butanol. The research results were published in mBio on June 14, 2016.
The team adopted a genoproteomic approach, combining genomics and proteomics, to investigate the metabolic features of C. tyrobutyricum. Unlike Clostridium acetobutylicum, the most widely used organism for 1-butanol production, C. tyrobutyricum has a novel butyrate-producing pathway and various mechanisms for energy conservation under anaerobic conditions. The expression of various metabolic genes, including those involved in butyrate formation, was analyzed using the “shotgun” proteome approach.
To date, the bio-based production of 1-butanol, a next-generation biofuel, has relied on several clostridial hosts including C. acetobutylicum and C. beijerinckii. However, these organisms have a low tolerance against 1-butanol even though they are naturally capable of producing it. C. tyrobutyricum cannot produce 1-butanol itself, but has a higher 1-butanol-tolerance and rapid uptake of monosaccharides, compared to those two species.
The team identified most of the genes involved in the central metabolism of C. tyrobutyricum from the whole-genome and shotgun proteome data, and this study will accelerate the bacterium’s engineering to produce useful chemicals including butyric acid and 1-butanol, replacing traditional bacterial hosts.
Professor Lee said,
“The unique metabolic features and energy conservation mechanisms of C. tyrobutyricum can be employed in the various microbial hosts we have previously developed to further improve their productivity and yield. Moreover, findings on C. tyrobutyricum revealed by this study will be the first step to directly engineer this bacterium.”
Director Jin-Woo Kim at the Platform Technology Division of the Ministry of Science, ICT and Future Planning of Korea, who oversees the Technology Development Program to Solve Climate Change, said,
“Over the years, Professor Lee’s team has researched the development of a bio-refinery system to produce natural and non-natural chemicals with the systems metabolic engineering of microorganisms. They were able to design strategies for the development of diverse industrial microbial strains to produce useful chemicals from inedible biomass-based carbon dioxide fixation. We believe the efficient production of butyric acid using a metabolic engineering approach will play an important role in the establishment of a bioprocess for chemical production.”
The title of the research paper is “Deciphering Clostridium tyrobutyricum Metabolism Based on the Who-Genome Sequence and Proteome Analyses.” (DOI: 10.1128/mBio.00743-16)
The lead authors are Joungmin Lee, a post-doctoral fellow in the BioProcess Research Center at KAIST, currently working in CJ CheilJedang Research Institute; Yu-Sin Jang, a research fellow in the BioProcess Research Center at KAIST, currently working at Gyeongsang National University as an assistant professor; and Mee-Jung Han, an assistant professor in the Environmental Engineering and Energy Department at Dongyang University. Jin Young Kim, a senior researcher at the Korea Basic Science Institute, also participated in the research.
This research was supported by the Technology Development Program to Solve Climate Change’s research project entitled “Systems Metabolic Engineering for Biorefineries” from the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea (NRF-2012M1A2A2026556).
Schematic Diagram of C. tyrobutyricum’s Genome Sequence and Its Proteome Profiles
The picture below shows the complete genome sequence, global protein expression profiles, and the genome-based metabolic characteristics during batch fermentation of C. tyrobutyricum.
2016.06.20 View 11594
Unveiling the Distinctive Features of Industrial Microorganism
KAIST researchers have sequenced the whole genome of Clostridium tyrobutyricum, which has a higher tolerance to toxic chemicals, such as 1-butanol, compared to other clostridial bacterial strains.
Clostridium tyrobutyricum, a Gram-positive, anaerobic spore-forming bacterium, is considered a promising industrial host strain for the production of various chemicals including butyric acid which has many applications in different industries such as a precursor to biofuels. Despite such potential, C. tyrobutyricum has received little attention, mainly due to a limited understanding of its genotypic and metabolic characteristics at the genome level.
A Korean research team headed by Distinguished Professor Sang Yup Lee of the Chemical and Biomolecular Engineering Department at the Korea Advanced Institute of Science and Technology (KAIST) deciphered the genome sequence of C. tyrobutyricum and its proteome profiles during the course of batch fermentation. As a result, the research team learned that the bacterium is not only capable of producing a large amount of butyric acid but also can tolerate toxic compounds such as 1-butanol. The research results were published in mBio on June 14, 2016.
The team adopted a genoproteomic approach, combining genomics and proteomics, to investigate the metabolic features of C. tyrobutyricum. Unlike Clostridium acetobutylicum, the most widely used organism for 1-butanol production, C. tyrobutyricum has a novel butyrate-producing pathway and various mechanisms for energy conservation under anaerobic conditions. The expression of various metabolic genes, including those involved in butyrate formation, was analyzed using the “shotgun” proteome approach.
To date, the bio-based production of 1-butanol, a next-generation biofuel, has relied on several clostridial hosts including C. acetobutylicum and C. beijerinckii. However, these organisms have a low tolerance against 1-butanol even though they are naturally capable of producing it. C. tyrobutyricum cannot produce 1-butanol itself, but has a higher 1-butanol-tolerance and rapid uptake of monosaccharides, compared to those two species.
The team identified most of the genes involved in the central metabolism of C. tyrobutyricum from the whole-genome and shotgun proteome data, and this study will accelerate the bacterium’s engineering to produce useful chemicals including butyric acid and 1-butanol, replacing traditional bacterial hosts.
Professor Lee said,
“The unique metabolic features and energy conservation mechanisms of C. tyrobutyricum can be employed in the various microbial hosts we have previously developed to further improve their productivity and yield. Moreover, findings on C. tyrobutyricum revealed by this study will be the first step to directly engineer this bacterium.”
Director Jin-Woo Kim at the Platform Technology Division of the Ministry of Science, ICT and Future Planning of Korea, who oversees the Technology Development Program to Solve Climate Change, said,
“Over the years, Professor Lee’s team has researched the development of a bio-refinery system to produce natural and non-natural chemicals with the systems metabolic engineering of microorganisms. They were able to design strategies for the development of diverse industrial microbial strains to produce useful chemicals from inedible biomass-based carbon dioxide fixation. We believe the efficient production of butyric acid using a metabolic engineering approach will play an important role in the establishment of a bioprocess for chemical production.”
The title of the research paper is “Deciphering Clostridium tyrobutyricum Metabolism Based on the Who-Genome Sequence and Proteome Analyses.” (DOI: 10.1128/mBio.00743-16)
The lead authors are Joungmin Lee, a post-doctoral fellow in the BioProcess Research Center at KAIST, currently working in CJ CheilJedang Research Institute; Yu-Sin Jang, a research fellow in the BioProcess Research Center at KAIST, currently working at Gyeongsang National University as an assistant professor; and Mee-Jung Han, an assistant professor in the Environmental Engineering and Energy Department at Dongyang University. Jin Young Kim, a senior researcher at the Korea Basic Science Institute, also participated in the research.
This research was supported by the Technology Development Program to Solve Climate Change’s research project entitled “Systems Metabolic Engineering for Biorefineries” from the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea (NRF-2012M1A2A2026556).
Schematic Diagram of C. tyrobutyricum’s Genome Sequence and Its Proteome Profiles
The picture below shows the complete genome sequence, global protein expression profiles, and the genome-based metabolic characteristics during batch fermentation of C. tyrobutyricum.
2016.06.20 View 11594 -
 Non-Natural Biomedical Polymers Produced from Microorganisms
KAIST researchers have developed metabolically engineered Escherichia coli strains to synthesize non-natural, biomedically important polymers including poly(lactate-co-glycolate) (PLGA), previously considered impossible to obtain from biobased materials.
Renewable non-food biomass could potentially replace petrochemical raw materials to produce energy sources, useful chemicals, or a vast array of petroleum-based end products such as plastics, lubricants, paints, fertilizers, and vitamin capsules. In recent years, biorefineries which transform non-edible biomass into fuel, heat, power, chemicals, and materials have received a great deal of attention as a sustainable alternative to decreasing the reliance on fossil fuels.
A research team headed by Distinguished Professor Sang Yup Lee of the Chemical and Biomolecular Engineering Department at KAIST has established a biorefinery system to create non-natural polymers from natural sources, allowing various plastics to be made in an environmentally-friendly and sustainable manner. The research results were published online in Nature Biotechnology on March 7, 2016. The print version will be issued in April 2016.
The research team adopted a systems metabolic engineering approach to develop a microorganism that can produce diverse non-natural, biomedically important polymers and succeeded in synthesizing poly(lactate-co-glycolate) (PLGA), a copolymer of two different polymer monomers, lactic and glycolic acid. PLGA is biodegradable, biocompatible, and non-toxic, and has been widely used in biomedical and therapeutic applications such as surgical sutures, prosthetic devices, drug delivery, and tissue engineering.
Inspired by the biosynthesis process for polyhydroxyalkanoates (PHAs), biologically-derived polyesters produced in nature by the bacterial fermentation of sugar or lipids, the research team designed a metabolic pathway for the biosynthesis of PLGA through microbial fermentation directly from carbohydrates in Escherichia coli (E. coli) strains.
The team had previously reported a recombinant E. coli producing PLGA by using the glyoxylate shunt pathway for the generation of glycolate from glucose, which was disclosed in their patents KR10-1575585-0000 (filing date of March 11, 2011), US08883463 and JP5820363. However, they discovered that the polymer content and glycolate fraction of PLGA could not be significantly enhanced via further engineering techniques. Thus, in this research, the team introduced a heterologous pathway to produce glycolate from xylose and succeeded in developing the recombinant E. coli producing PLGA and various novel copolymers much more efficiently.
In order to produce PLGA by microbial fermentation directly from carbohydrates, the team incorporated external and engineered enzymes as catalysts to co-polymerize PLGA while establishing a few additional metabolic pathways for the biosynthesis to produce a range of different non-natural polymers, some for the first time. This bio-based synthetic process for PLGA and other polymers could substitute for the existing complicated chemical production that involves the preparation and purification of precursors, chemical polymerization processes, and the elimination of metal catalysts.
Professor Lee and his team performed in silico genome-scale metabolic simulations of the E. coli cell to predict and analyze changes in the metabolic fluxes of cells which were caused by the introduction of external metabolic pathways. Based on these results, genes are manipulated to optimize metabolic fluxes by eliminating the genes responsible for byproducts formation and enhancing the expression levels of certain genes, thereby achieving the effective production of target polymers as well as stimulating cell growth.
The team utilized the structural basis of broad substrate specificity of the key synthesizing enzyme, PHA synthase, to incorporate various co-monomers with main and side chains of different lengths. These monomers were produced inside the cell by metabolic engineering, and then copolymerized to improve the material properties of PLGA. As a result, a variety of PLGA copolymers with different monomer compositions such as the US Food and Drug Administration (FDA)-approved monomers, 3-hydroxyburate, 4-hydroxyburate, and 6-hydroxyhexanoate, were produced. Newly applied bioplastics such as 5-hydroxyvalerate and 2-hydroxyisovalerate were also made.
The team employed a systems metabolic engineering application which, according to the researchers, is the first successful example of biological production of PGLA and several novel copolymers from renewable biomass by one-step direct fermentation of metabolically engineered E.coli.
Professor Lee said, “We presented important findings that non-natural polymers, such as PLGA which is commonly used for drug delivery or biomedical devices, were produced by a metabolically engineered gut bacterium. Our research is meaningful in that it proposes a platform strategy in metabolic engineering, which can be further utilized in the development of numerous non-natural, useful polymers.”
Director Ilsub Baek at the Platform Technology Division of the Ministry of Science, ICT and Future Planning of Korea, who oversees the Technology Development Program to Solve Climate Change, said, “Professor Lee has led one of our research projects, the Systems Metabolic Engineering for Biorefineries, which began as part of the Ministry’s Technology Development Program to Solve Climate Change. He and his team have continuously achieved promising results and been attracting greater interest from the global scientific community. As climate change technology grows more important, this research on the biological production of non-natural, high value polymers will have a great impact on science and industry.”
The title of the research paper is “One-step Fermentative Production of Poly(lactate-co-glycolate) from Carbohydrates in Escherichia coli (DOI: 10.1038/nbt.3485).” The lead authors are So Young Choi, a Ph.D. candidate in the Department of Chemical and Biomolecular Engineering at KAIST, and Si Jae Park, Assistant Professor of the Environmental Engineering and Energy Department at Myongji University. Won Jun Kim and Jung Eun Yang, both doctoral students in the Department of Chemical and Biomolecular Engineering at KAIST, also participated in the research.
This research was supported by the Technology Development Program to Solve Climate Change’s research project titled “Systems Metabolic Engineering for Biorefineries” from the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea (NRF-2012M1A2A2026556).
Figure: Production of PLGA and Other Non-Natural Copolymers
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced a variety of PLGAs with different monomer compositions, proposing the chemosynthetic process of non-natural polymers from biomass. The non-natural polymer PLGA and its other copolymers, which were produced by engineered bacteria developed by taking a systems metabolic engineering approach, accumulate in granule forms within a cell.
2016.03.08 View 12799
Non-Natural Biomedical Polymers Produced from Microorganisms
KAIST researchers have developed metabolically engineered Escherichia coli strains to synthesize non-natural, biomedically important polymers including poly(lactate-co-glycolate) (PLGA), previously considered impossible to obtain from biobased materials.
Renewable non-food biomass could potentially replace petrochemical raw materials to produce energy sources, useful chemicals, or a vast array of petroleum-based end products such as plastics, lubricants, paints, fertilizers, and vitamin capsules. In recent years, biorefineries which transform non-edible biomass into fuel, heat, power, chemicals, and materials have received a great deal of attention as a sustainable alternative to decreasing the reliance on fossil fuels.
A research team headed by Distinguished Professor Sang Yup Lee of the Chemical and Biomolecular Engineering Department at KAIST has established a biorefinery system to create non-natural polymers from natural sources, allowing various plastics to be made in an environmentally-friendly and sustainable manner. The research results were published online in Nature Biotechnology on March 7, 2016. The print version will be issued in April 2016.
The research team adopted a systems metabolic engineering approach to develop a microorganism that can produce diverse non-natural, biomedically important polymers and succeeded in synthesizing poly(lactate-co-glycolate) (PLGA), a copolymer of two different polymer monomers, lactic and glycolic acid. PLGA is biodegradable, biocompatible, and non-toxic, and has been widely used in biomedical and therapeutic applications such as surgical sutures, prosthetic devices, drug delivery, and tissue engineering.
Inspired by the biosynthesis process for polyhydroxyalkanoates (PHAs), biologically-derived polyesters produced in nature by the bacterial fermentation of sugar or lipids, the research team designed a metabolic pathway for the biosynthesis of PLGA through microbial fermentation directly from carbohydrates in Escherichia coli (E. coli) strains.
The team had previously reported a recombinant E. coli producing PLGA by using the glyoxylate shunt pathway for the generation of glycolate from glucose, which was disclosed in their patents KR10-1575585-0000 (filing date of March 11, 2011), US08883463 and JP5820363. However, they discovered that the polymer content and glycolate fraction of PLGA could not be significantly enhanced via further engineering techniques. Thus, in this research, the team introduced a heterologous pathway to produce glycolate from xylose and succeeded in developing the recombinant E. coli producing PLGA and various novel copolymers much more efficiently.
In order to produce PLGA by microbial fermentation directly from carbohydrates, the team incorporated external and engineered enzymes as catalysts to co-polymerize PLGA while establishing a few additional metabolic pathways for the biosynthesis to produce a range of different non-natural polymers, some for the first time. This bio-based synthetic process for PLGA and other polymers could substitute for the existing complicated chemical production that involves the preparation and purification of precursors, chemical polymerization processes, and the elimination of metal catalysts.
Professor Lee and his team performed in silico genome-scale metabolic simulations of the E. coli cell to predict and analyze changes in the metabolic fluxes of cells which were caused by the introduction of external metabolic pathways. Based on these results, genes are manipulated to optimize metabolic fluxes by eliminating the genes responsible for byproducts formation and enhancing the expression levels of certain genes, thereby achieving the effective production of target polymers as well as stimulating cell growth.
The team utilized the structural basis of broad substrate specificity of the key synthesizing enzyme, PHA synthase, to incorporate various co-monomers with main and side chains of different lengths. These monomers were produced inside the cell by metabolic engineering, and then copolymerized to improve the material properties of PLGA. As a result, a variety of PLGA copolymers with different monomer compositions such as the US Food and Drug Administration (FDA)-approved monomers, 3-hydroxyburate, 4-hydroxyburate, and 6-hydroxyhexanoate, were produced. Newly applied bioplastics such as 5-hydroxyvalerate and 2-hydroxyisovalerate were also made.
The team employed a systems metabolic engineering application which, according to the researchers, is the first successful example of biological production of PGLA and several novel copolymers from renewable biomass by one-step direct fermentation of metabolically engineered E.coli.
Professor Lee said, “We presented important findings that non-natural polymers, such as PLGA which is commonly used for drug delivery or biomedical devices, were produced by a metabolically engineered gut bacterium. Our research is meaningful in that it proposes a platform strategy in metabolic engineering, which can be further utilized in the development of numerous non-natural, useful polymers.”
Director Ilsub Baek at the Platform Technology Division of the Ministry of Science, ICT and Future Planning of Korea, who oversees the Technology Development Program to Solve Climate Change, said, “Professor Lee has led one of our research projects, the Systems Metabolic Engineering for Biorefineries, which began as part of the Ministry’s Technology Development Program to Solve Climate Change. He and his team have continuously achieved promising results and been attracting greater interest from the global scientific community. As climate change technology grows more important, this research on the biological production of non-natural, high value polymers will have a great impact on science and industry.”
The title of the research paper is “One-step Fermentative Production of Poly(lactate-co-glycolate) from Carbohydrates in Escherichia coli (DOI: 10.1038/nbt.3485).” The lead authors are So Young Choi, a Ph.D. candidate in the Department of Chemical and Biomolecular Engineering at KAIST, and Si Jae Park, Assistant Professor of the Environmental Engineering and Energy Department at Myongji University. Won Jun Kim and Jung Eun Yang, both doctoral students in the Department of Chemical and Biomolecular Engineering at KAIST, also participated in the research.
This research was supported by the Technology Development Program to Solve Climate Change’s research project titled “Systems Metabolic Engineering for Biorefineries” from the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea (NRF-2012M1A2A2026556).
Figure: Production of PLGA and Other Non-Natural Copolymers
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced a variety of PLGAs with different monomer compositions, proposing the chemosynthetic process of non-natural polymers from biomass. The non-natural polymer PLGA and its other copolymers, which were produced by engineered bacteria developed by taking a systems metabolic engineering approach, accumulate in granule forms within a cell.
2016.03.08 View 12799 -
 Asia Pacific Biotech News' Special Coverage of Korean Biotechnology
The Asia Pacific Biotech News covered five major biotechnology research projects sponsored by the Korean government in the areas of biofuels, biomedicine, bio-nano healthcare, and biorefinery.
The Asia Pacific Biotech News (APBN), a monthly magazine based in Singapore, which offers comprehensive reports on the fields of pharmaceuticals, healthcare, and biotechnology, recently published a special feature on Korea’s biotechnology research and development (R&D) programs.
The magazine feature selected five research programs sponsored by the Korean government, which are either part of the Global Frontier or the Climate Change Technology Development Projects.
The programs are:
Systems Metabolic Engineering Research: Distinguished Professor Sang Yup Lee
of the Chemical and Biomolecular Engineering Department at the Korea Advanced
Institute of Science and Technology (KAIST) has been leading a research group to
develop biorefining technology using renewable non-food biomass to produce
chemicals, fuels, and materials that were largely drawn from fossil resources
through petrochemical refinery processes. Applying a systems metabolic
engineering approach, the group succeeded in modifying the metabolic pathways of
microorganisms. As a result, they produced, for the first time in the world,
engineered plastic raw materials and gasoline. The team also developed a technique
to produce butanol and succinic acid with a higher titer and yield using metabolically
engineered microorganisms.
Next-generation Biomass Research: Under the leadership of Professor Yong-
Keun Chang of the Chemical and Biomolecular Engineering Department at KAIST,
the research project, which belongs to the Global Frontier Project, develops biofuels
and bioproducts utilizing microalgae typically found in water and other marine
systems.
Convergence Research for Biomedicine: Professor Sung-Hoon Kim of Seoul
National University leads this project that develops targeted new drugs based on
convergence research strategies.
Bionano Healthcare Chip Research: Director Bong-Hyun Chung of the Korea
Research Institute of Bioscience and Biotechnology has integrated information and
communications technology, nanotechnology, and biotechnology to develop a
diagnostic kit that can screen toxic germs, virus, and toxic materials in a prompt
and accurate manner.
Biosynergy Research: Led by Professor Do-Hun Lee of the Bio and Brain
Engineering Department at KAIST, this research project develops new treatments
with a multi-target, multi-component approach in the context of systems biology
through an analysis of synergistic reactions between multi-compounds in traditional
East Asian medicine and human metabolites. In East Asian medicine, treatment and
caring of the human body are considered analogous to the politics of governing a
nation. Based on such system, the research focuses on designing a foundation for
the integration of traditional medicine with modern drug discovery and development.
Director Ilsub Baek at the Platform Technology Division of the Ministry of Science, ICT and Future Planning, Republic of Korea, who is responsible for the Global Frontier Program and the Technology to Solve Climate Change, said, “It is great to see that Asia Pacific Biotech News published an extensive coverage of Korea’s several key research programs on biotechnology as its first issue of this year. I am sure that these programs will lead to great outcomes to solve many worldwide pending issues including climate change and healthcare in the aging society.”
Professor Sang Yup Lee, who served as an editor of the feature, said, “At the request of the magazine, we have already published lead articles on our biotechnology research three times in the past in 2002, 2006, and 2011. I am pleased to see continued coverage of Korean biotechnology by the magazine because it recognizes the excellence of our research. Biotechnology has emerged as one of the strong fields that addresses important global issues such as climate change and sustainability.”
2016.02.04 View 14230
Asia Pacific Biotech News' Special Coverage of Korean Biotechnology
The Asia Pacific Biotech News covered five major biotechnology research projects sponsored by the Korean government in the areas of biofuels, biomedicine, bio-nano healthcare, and biorefinery.
The Asia Pacific Biotech News (APBN), a monthly magazine based in Singapore, which offers comprehensive reports on the fields of pharmaceuticals, healthcare, and biotechnology, recently published a special feature on Korea’s biotechnology research and development (R&D) programs.
The magazine feature selected five research programs sponsored by the Korean government, which are either part of the Global Frontier or the Climate Change Technology Development Projects.
The programs are:
Systems Metabolic Engineering Research: Distinguished Professor Sang Yup Lee
of the Chemical and Biomolecular Engineering Department at the Korea Advanced
Institute of Science and Technology (KAIST) has been leading a research group to
develop biorefining technology using renewable non-food biomass to produce
chemicals, fuels, and materials that were largely drawn from fossil resources
through petrochemical refinery processes. Applying a systems metabolic
engineering approach, the group succeeded in modifying the metabolic pathways of
microorganisms. As a result, they produced, for the first time in the world,
engineered plastic raw materials and gasoline. The team also developed a technique
to produce butanol and succinic acid with a higher titer and yield using metabolically
engineered microorganisms.
Next-generation Biomass Research: Under the leadership of Professor Yong-
Keun Chang of the Chemical and Biomolecular Engineering Department at KAIST,
the research project, which belongs to the Global Frontier Project, develops biofuels
and bioproducts utilizing microalgae typically found in water and other marine
systems.
Convergence Research for Biomedicine: Professor Sung-Hoon Kim of Seoul
National University leads this project that develops targeted new drugs based on
convergence research strategies.
Bionano Healthcare Chip Research: Director Bong-Hyun Chung of the Korea
Research Institute of Bioscience and Biotechnology has integrated information and
communications technology, nanotechnology, and biotechnology to develop a
diagnostic kit that can screen toxic germs, virus, and toxic materials in a prompt
and accurate manner.
Biosynergy Research: Led by Professor Do-Hun Lee of the Bio and Brain
Engineering Department at KAIST, this research project develops new treatments
with a multi-target, multi-component approach in the context of systems biology
through an analysis of synergistic reactions between multi-compounds in traditional
East Asian medicine and human metabolites. In East Asian medicine, treatment and
caring of the human body are considered analogous to the politics of governing a
nation. Based on such system, the research focuses on designing a foundation for
the integration of traditional medicine with modern drug discovery and development.
Director Ilsub Baek at the Platform Technology Division of the Ministry of Science, ICT and Future Planning, Republic of Korea, who is responsible for the Global Frontier Program and the Technology to Solve Climate Change, said, “It is great to see that Asia Pacific Biotech News published an extensive coverage of Korea’s several key research programs on biotechnology as its first issue of this year. I am sure that these programs will lead to great outcomes to solve many worldwide pending issues including climate change and healthcare in the aging society.”
Professor Sang Yup Lee, who served as an editor of the feature, said, “At the request of the magazine, we have already published lead articles on our biotechnology research three times in the past in 2002, 2006, and 2011. I am pleased to see continued coverage of Korean biotechnology by the magazine because it recognizes the excellence of our research. Biotechnology has emerged as one of the strong fields that addresses important global issues such as climate change and sustainability.”
2016.02.04 View 14230 -
 KAIST and Hanwha Chemical Agree on Research Collaboration
KAIST signed a memorandum of understanding (MOU) with Hanwha Chemical Co., Ltd., a Korean chemical and auto manufacturer, on November 2, 2015 to establish a research center on campus.
The research center, which will be named “KAIST-Hanwha Chemical Future Technology Research Center,” will implement joint research projects for five years beginning from 2016 to develop innovative, green technologies that will help the Korean chemical industry boost its global competitiveness and to nurture top researchers and engineers in chemical engineering.
The research center will lead the development of next-generation petrochemical materials and manufacturing technology and the establishment of pure high-refining processes which are more energy-efficient and environmentally friendly. KAIST and Hanwha will strive to secure new technologies that have the greatest commercialization potential in the global market. They will also establish a scholarship fund for 15 KAIST doctoral students in the Department of Chemical and Biomolecular Engineering.
Many professors from the Chemical and Biomolecular Engineering Department including Distinguished Professor Sang Yup Lee, who was listed in the Top 20 Translational Researchers of 2014 by Nature Biotechnology this year, and Professor Hyunjoo Lee who received the Woman Scholar award at the 2015 World Chemistry Conference, will work at the research center.
Professor Lee, the head of the research center, said, “Collaborating with Hanwha will give us a strong basis for our efforts to carry out original research and train the best researchers in the field.”
Chang-Bum Kim, the Chief Executive Officer (CEO) of Hanwha Chemical, said,
“We hope our collaborations with KAIST will go beyond the typical industry and university cooperation. The two organizations will indeed jointly operate the research center, and this will become a new model for industry and university cooperation. We expect that the research center will play a crucial role in the development of new products and technologies to grow the Korean chemical industry.”
In the photo, President Steve Kang of KAIST (fourth from left) and CEO Chang-Bum Kim of Hanwha Chemical (fifth from left) hold the MOU together.
2015.11.01 View 13202
KAIST and Hanwha Chemical Agree on Research Collaboration
KAIST signed a memorandum of understanding (MOU) with Hanwha Chemical Co., Ltd., a Korean chemical and auto manufacturer, on November 2, 2015 to establish a research center on campus.
The research center, which will be named “KAIST-Hanwha Chemical Future Technology Research Center,” will implement joint research projects for five years beginning from 2016 to develop innovative, green technologies that will help the Korean chemical industry boost its global competitiveness and to nurture top researchers and engineers in chemical engineering.
The research center will lead the development of next-generation petrochemical materials and manufacturing technology and the establishment of pure high-refining processes which are more energy-efficient and environmentally friendly. KAIST and Hanwha will strive to secure new technologies that have the greatest commercialization potential in the global market. They will also establish a scholarship fund for 15 KAIST doctoral students in the Department of Chemical and Biomolecular Engineering.
Many professors from the Chemical and Biomolecular Engineering Department including Distinguished Professor Sang Yup Lee, who was listed in the Top 20 Translational Researchers of 2014 by Nature Biotechnology this year, and Professor Hyunjoo Lee who received the Woman Scholar award at the 2015 World Chemistry Conference, will work at the research center.
Professor Lee, the head of the research center, said, “Collaborating with Hanwha will give us a strong basis for our efforts to carry out original research and train the best researchers in the field.”
Chang-Bum Kim, the Chief Executive Officer (CEO) of Hanwha Chemical, said,
“We hope our collaborations with KAIST will go beyond the typical industry and university cooperation. The two organizations will indeed jointly operate the research center, and this will become a new model for industry and university cooperation. We expect that the research center will play a crucial role in the development of new products and technologies to grow the Korean chemical industry.”
In the photo, President Steve Kang of KAIST (fourth from left) and CEO Chang-Bum Kim of Hanwha Chemical (fifth from left) hold the MOU together.
2015.11.01 View 13202 -
 Establishment of System Metabolic Engineering Strategies
Although conventional petrochemical processes have generated chemicals and materials which have been useful to mankind, they have also triggered a variety of environmental problems including climate change and relied too much on nonrenewable natural resources. To ameliorate this, researchers have actively pursued the development of industrial microbial strains around the globe in order to overproduce industrially useful chemicals and materials from microbes using renewable biomass. This discipline is called metabolic engineering.
Thanks to advances in genetic engineering and our knowledge of cellular metabolism, conventional metabolic engineering efforts have succeeded to a certain extent in developing microbial strains that overproduce bioproducts at an industrial level. However, many metabolic engineering projects launched in academic labs do not reach commercial markets due to a failure to fully integrate industrial bioprocesses.
In response to this, Distinguished Professor Sang Yup Lee and Dr. Hyun Uk Kim, both from the Department of Chemical and Biomolecular Engineering at KAIST, have recently suggested ten general strategies of systems metabolic engineering to successfully develop industrial microbial strains. Systems metabolic engineering differs from conventional metabolic engineering by incorporating traditional metabolic engineering approaches along with tools of other fields, such as systems biology, synthetic biology, and molecular evolution.
The ten strategies of systems metabolic engineering have been featured in Nature Biotechnology released online in October 2015, which is entitled "Systems strategies for developing industrial microbial strains."
The strategies cover economic, state-of-the-art biological techniques and traditional bioprocess aspects. Specifically, they consist of: 1) project design including economic evaluation of a target bioproduct; 2) selection of host strains to be used for overproduction of a bioproduct; 3) metabolic pathway reconstruction for bioproducts that are not naturally produced in the selected host strains; 4) increasing tolerance of a host strain against the bioproduct; 5) removing negative regulatory circuits in the microbial host limiting overproduction of a bioproduct; 6) rerouting intracellular fluxes to optimize cofactor and precursor availability necessary for the bioproduct formation; 7) diagnosing and optimizing metabolic fluxes towards product formation; 8) diagnosis and optimization of microbial culture conditions including carbon sources; 9) system-wide gene manipulation to further increase the host strain's production performance using high-throughput genome-scale engineering and computational tools; and 10) scale-up fermentation of the developed strain and diagnosis for the reproducibility of the strain's production performance.
These ten strategies were articulated with successful examples of the production of L-arginine using Corynebacterium glutamicum, 1,4-butanediol using Escherichia coli, and L-lysine and bio-nylon using C. glutamicum.
Professor Sang Yup Lee said, "At the moment, the chance of commercializing microbial strains developed in academic labs is very low. The strategies of systems metabolic engineering outlined in this analysis can serve as guidelines when developing industrial microbial strains. We hope that these strategies contribute to improving opportunities to commercialize microbial strains developed in academic labs with drastically reduced costs and efforts, and that a large fraction of petroleum-based processes will be replaced with sustainable bioprocesses."
Lee S. Y. & Kim, H. U. Systems Strategies for Developing Industrial Microbial Strains. Nature Biotechnology (2015).
This work was supported by the Technology Development Program to Solve Climate Change on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556) and by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) from the Ministry of Science, ICT and Future Planning (MSIP), Korea, and through the National Research Foundation (NRF) of Korea. This work was also supported by the Novo Nordisk Foundation.
Picture: Concept of the Systems Metabolic Engineering Framework
(a) Three major bioprocess stages (b) Considerations in systems metabolic engineering to optimize the whole bioprocess. List of considerations for the strain development and fermentation contribute to improving microbial strain's production performance (red), whereas those for the separation and purification help in reducing overall operation costs by facilitating the downstream process (blue). Some of the considerations can be repeated in the course of systems metabolic engineering.
2015.10.19 View 11892
Establishment of System Metabolic Engineering Strategies
Although conventional petrochemical processes have generated chemicals and materials which have been useful to mankind, they have also triggered a variety of environmental problems including climate change and relied too much on nonrenewable natural resources. To ameliorate this, researchers have actively pursued the development of industrial microbial strains around the globe in order to overproduce industrially useful chemicals and materials from microbes using renewable biomass. This discipline is called metabolic engineering.
Thanks to advances in genetic engineering and our knowledge of cellular metabolism, conventional metabolic engineering efforts have succeeded to a certain extent in developing microbial strains that overproduce bioproducts at an industrial level. However, many metabolic engineering projects launched in academic labs do not reach commercial markets due to a failure to fully integrate industrial bioprocesses.
In response to this, Distinguished Professor Sang Yup Lee and Dr. Hyun Uk Kim, both from the Department of Chemical and Biomolecular Engineering at KAIST, have recently suggested ten general strategies of systems metabolic engineering to successfully develop industrial microbial strains. Systems metabolic engineering differs from conventional metabolic engineering by incorporating traditional metabolic engineering approaches along with tools of other fields, such as systems biology, synthetic biology, and molecular evolution.
The ten strategies of systems metabolic engineering have been featured in Nature Biotechnology released online in October 2015, which is entitled "Systems strategies for developing industrial microbial strains."
The strategies cover economic, state-of-the-art biological techniques and traditional bioprocess aspects. Specifically, they consist of: 1) project design including economic evaluation of a target bioproduct; 2) selection of host strains to be used for overproduction of a bioproduct; 3) metabolic pathway reconstruction for bioproducts that are not naturally produced in the selected host strains; 4) increasing tolerance of a host strain against the bioproduct; 5) removing negative regulatory circuits in the microbial host limiting overproduction of a bioproduct; 6) rerouting intracellular fluxes to optimize cofactor and precursor availability necessary for the bioproduct formation; 7) diagnosing and optimizing metabolic fluxes towards product formation; 8) diagnosis and optimization of microbial culture conditions including carbon sources; 9) system-wide gene manipulation to further increase the host strain's production performance using high-throughput genome-scale engineering and computational tools; and 10) scale-up fermentation of the developed strain and diagnosis for the reproducibility of the strain's production performance.
These ten strategies were articulated with successful examples of the production of L-arginine using Corynebacterium glutamicum, 1,4-butanediol using Escherichia coli, and L-lysine and bio-nylon using C. glutamicum.
Professor Sang Yup Lee said, "At the moment, the chance of commercializing microbial strains developed in academic labs is very low. The strategies of systems metabolic engineering outlined in this analysis can serve as guidelines when developing industrial microbial strains. We hope that these strategies contribute to improving opportunities to commercialize microbial strains developed in academic labs with drastically reduced costs and efforts, and that a large fraction of petroleum-based processes will be replaced with sustainable bioprocesses."
Lee S. Y. & Kim, H. U. Systems Strategies for Developing Industrial Microbial Strains. Nature Biotechnology (2015).
This work was supported by the Technology Development Program to Solve Climate Change on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556) and by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) from the Ministry of Science, ICT and Future Planning (MSIP), Korea, and through the National Research Foundation (NRF) of Korea. This work was also supported by the Novo Nordisk Foundation.
Picture: Concept of the Systems Metabolic Engineering Framework
(a) Three major bioprocess stages (b) Considerations in systems metabolic engineering to optimize the whole bioprocess. List of considerations for the strain development and fermentation contribute to improving microbial strain's production performance (red), whereas those for the separation and purification help in reducing overall operation costs by facilitating the downstream process (blue). Some of the considerations can be repeated in the course of systems metabolic engineering.
2015.10.19 View 11892 -
 Discovery of Redox-Switch of KEenzyme Involved in N-Butanol Biosynthesis
Research teams at KAIST and Kyungpook National University (KNU) have succeeded in uncovering the redox-switch of thiolase, a key enzyme for n-butanol production in Clostridium acetobutylicum, one of the best known butanol-producing bacteria.
Biological n-butanol production was first reported by Louis Pasteur in 1861, and the bioprocess was industrialized usingClostridium acetobutylicum. The fermentation process by Clostridium strains has been known to be the most efficient one for n-butanol production. Due to growing world-wide issues such as energy security and climate change, the biological production of n-butanol has been receiving much renewed interest. This is because n-butanol possesses much better fuel characteristics compared to ethanol, such as higher energy content (29.2 MJ/L vs 19.6 MJ/L), less corrosiveness, less hygroscopy, and the ease with which it can be blended with gasoline and diesel.
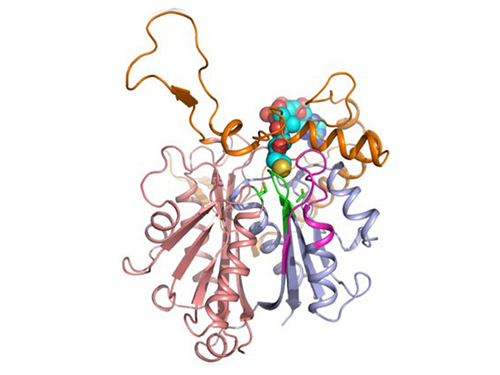
In the paper published in Nature Communications, a broad-scope, online-only, and open access journal issued by the Nature Publishing Group (NPG), on September 22, 2015, Professor Kyung-Jin Kim at the School of Life Sciences, KNU, and Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering, KAIST, have proved that the redox-switch of thiolase plays a role in a regulation of metabolic flux in C. acetobutylicum by using in silico modeling and simulation tools.
The research team has redesigned thiolase with enhanced activity on the basis of the 3D structure of the wild-type enzyme. To reinforce a metabolic flux toward butanol production, the metabolic network of C. acetobutylicum strain was engineered with the redesigned enzyme. The combination of the discovery of 3D enzyme structure and systems metabolic engineering approaches resulted in increased n-butanol production in C. acetobutylicum, which allows the production of this important industrial chemical to be cost competitive.
Professors Kim and Lee said, "We have reported the 3D structure of C. acetobutylicum thiolase-a key enzyme involved in n-butanol biosynthesis, for the first time. Further study will be done to produce butanol more economically on the basis of the 3D structure of C. acetobutylicum thiolase."
This work was published online in Nature Communications on September 22, 2015.
Reference: Kim et al. "Redox-switch regulatory mechanism of thiolase from Clostridium acetobutylicum," Nature Communications
This research was supported by the Technology Development Program to Solve Climate Changes from the Ministry of Education, Science and Technology (MEST), Korea, the National Research Foundation of Korea, and the Advanced Biomass Center through the Global Frontier Research Program of the MEST, Korea.
For further information, contact Dr. Sang Yup Lee, Distinguished Professor, KAIST, Daejeon, Korea (leesy@kaist.ac.kr, +82-42-350-3930); and Dr. Kyung-Jin Kim, Professor, KNU, Daegu, Korea (kkim@knu.ac.kr, +82-53-950-6088).
Figure 1: A redox-switch of thiolase involves in butanol biosynthesis in Clostridium acetobutylicum. Thiolase condenses two acetyl-CoA molecules for initiating four carbon flux towards butanol.
Figure 2: Thiolase catalyzes the condensation reaction of acetyl-CoA to acetoacetyl-CoA. Two catalytic cysteine residues at 88th and 378th are oxidized and formed an intermolecular disulfide bond in an oxidized status, which results in inactivation of the enzyme for n-butanol biosynthesis. The intermolecular disulfide bond is broken enabling the n-butanol biosynthesis, when the environment status is reduced.
2015.09.23 View 11708
Discovery of Redox-Switch of KEenzyme Involved in N-Butanol Biosynthesis
Research teams at KAIST and Kyungpook National University (KNU) have succeeded in uncovering the redox-switch of thiolase, a key enzyme for n-butanol production in Clostridium acetobutylicum, one of the best known butanol-producing bacteria.
Biological n-butanol production was first reported by Louis Pasteur in 1861, and the bioprocess was industrialized usingClostridium acetobutylicum. The fermentation process by Clostridium strains has been known to be the most efficient one for n-butanol production. Due to growing world-wide issues such as energy security and climate change, the biological production of n-butanol has been receiving much renewed interest. This is because n-butanol possesses much better fuel characteristics compared to ethanol, such as higher energy content (29.2 MJ/L vs 19.6 MJ/L), less corrosiveness, less hygroscopy, and the ease with which it can be blended with gasoline and diesel.
In the paper published in Nature Communications, a broad-scope, online-only, and open access journal issued by the Nature Publishing Group (NPG), on September 22, 2015, Professor Kyung-Jin Kim at the School of Life Sciences, KNU, and Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering, KAIST, have proved that the redox-switch of thiolase plays a role in a regulation of metabolic flux in C. acetobutylicum by using in silico modeling and simulation tools.
The research team has redesigned thiolase with enhanced activity on the basis of the 3D structure of the wild-type enzyme. To reinforce a metabolic flux toward butanol production, the metabolic network of C. acetobutylicum strain was engineered with the redesigned enzyme. The combination of the discovery of 3D enzyme structure and systems metabolic engineering approaches resulted in increased n-butanol production in C. acetobutylicum, which allows the production of this important industrial chemical to be cost competitive.
Professors Kim and Lee said, "We have reported the 3D structure of C. acetobutylicum thiolase-a key enzyme involved in n-butanol biosynthesis, for the first time. Further study will be done to produce butanol more economically on the basis of the 3D structure of C. acetobutylicum thiolase."
This work was published online in Nature Communications on September 22, 2015.
Reference: Kim et al. "Redox-switch regulatory mechanism of thiolase from Clostridium acetobutylicum," Nature Communications
This research was supported by the Technology Development Program to Solve Climate Changes from the Ministry of Education, Science and Technology (MEST), Korea, the National Research Foundation of Korea, and the Advanced Biomass Center through the Global Frontier Research Program of the MEST, Korea.
For further information, contact Dr. Sang Yup Lee, Distinguished Professor, KAIST, Daejeon, Korea (leesy@kaist.ac.kr, +82-42-350-3930); and Dr. Kyung-Jin Kim, Professor, KNU, Daegu, Korea (kkim@knu.ac.kr, +82-53-950-6088).
Figure 1: A redox-switch of thiolase involves in butanol biosynthesis in Clostridium acetobutylicum. Thiolase condenses two acetyl-CoA molecules for initiating four carbon flux towards butanol.
Figure 2: Thiolase catalyzes the condensation reaction of acetyl-CoA to acetoacetyl-CoA. Two catalytic cysteine residues at 88th and 378th are oxidized and formed an intermolecular disulfide bond in an oxidized status, which results in inactivation of the enzyme for n-butanol biosynthesis. The intermolecular disulfide bond is broken enabling the n-butanol biosynthesis, when the environment status is reduced.
2015.09.23 View 11708 -
 'Engineered Bacterium Produces 1,3-Diaminopropane'
A research team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported, for the first time, the production of 1,3-diaminopropane via fermentation of an engineered bacterium.
1,3-Diaminopropane is a three carbon diamine, which has a wide range of industrial applications including epoxy resin and cross-linking agents, as well as precursors for pharmaceuticals, agrochemicals, and organic chemicals. It can also be polymerized with dicarboxylic acids to make polyamides (nylons) for use as engineering plastics, medical materials, and adhesives.
Traditionally, 1,3-diaminopropane is derived from petroleum-based processes. In effort to address critical problems such as the depletion of petroleum and environmental issues inherent to the petroleum-based processes, the research team has developed an Escherichia coli (E. coli) strain capable of producing 1,3-diaminopropane. Using this technology, 1,3-diaminopropane can now be produced from renewable biomass instead of petroleum.
E. coli as found in nature is unable to produce 1,3-diaminopropane. Metabolic engineering, a technology to transform microorganisms into highly efficient microbial cell factories capable of producing chemical compounds of interest, was utilized to engineer the E. coli strain. First, naturally existing metabolic pathways for the biosynthesis of 1,3-diaminopropane were introduced into a virtual cell in silico to determine the most efficient metabolic pathway for the 1,3-diaminopropane production. The metabolic pathway selected was then introduced into an E. coli strain and successfully produced 1,3-diaminopropane for the first time in the world.
The research team applied metabolic engineering additionally, and the production titer of 1,3-diaminopropane increased about 21 fold. The Fed-batch fermentation of the engineered E. coli strain produced 13 grams per liter of 1,3-diaminoproapne. With this technology, 1,3-diaminopropane can be produced using renewable biomass, and it will be the starting point for replacing the current petroleum-based processes with bio-based processes.
Professor Lee said, “Our study suggested a possibility to produce 1,3-diaminopropane based on biorefinery. Further study will be done to increase the titer and productivity of 1,3-diaminopropane.”
This work was published online in Scientific Reports on August 11, 2015.
Reference: Chae, T.U. et al. "Metabolic engineering of Escherichia coli for the production of 1,3-diaminopropane, a three carbon diamine," Scientific Reports:
http://www.nature.com/articles/srep13040
This research was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from Ministry of Science, ICT and Future Planning (MSIP) through the National Research Foundation (NRF) of Korea.
Figure 1: Metabolic engineering strategies for 1,3-diaminopropane production using C4 pathway
Figure 2: Fed-batch fermentation profiles of two final engineered E. coli strains
2015.08.12 View 11898
'Engineered Bacterium Produces 1,3-Diaminopropane'
A research team led by Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST reported, for the first time, the production of 1,3-diaminopropane via fermentation of an engineered bacterium.
1,3-Diaminopropane is a three carbon diamine, which has a wide range of industrial applications including epoxy resin and cross-linking agents, as well as precursors for pharmaceuticals, agrochemicals, and organic chemicals. It can also be polymerized with dicarboxylic acids to make polyamides (nylons) for use as engineering plastics, medical materials, and adhesives.
Traditionally, 1,3-diaminopropane is derived from petroleum-based processes. In effort to address critical problems such as the depletion of petroleum and environmental issues inherent to the petroleum-based processes, the research team has developed an Escherichia coli (E. coli) strain capable of producing 1,3-diaminopropane. Using this technology, 1,3-diaminopropane can now be produced from renewable biomass instead of petroleum.
E. coli as found in nature is unable to produce 1,3-diaminopropane. Metabolic engineering, a technology to transform microorganisms into highly efficient microbial cell factories capable of producing chemical compounds of interest, was utilized to engineer the E. coli strain. First, naturally existing metabolic pathways for the biosynthesis of 1,3-diaminopropane were introduced into a virtual cell in silico to determine the most efficient metabolic pathway for the 1,3-diaminopropane production. The metabolic pathway selected was then introduced into an E. coli strain and successfully produced 1,3-diaminopropane for the first time in the world.
The research team applied metabolic engineering additionally, and the production titer of 1,3-diaminopropane increased about 21 fold. The Fed-batch fermentation of the engineered E. coli strain produced 13 grams per liter of 1,3-diaminoproapne. With this technology, 1,3-diaminopropane can be produced using renewable biomass, and it will be the starting point for replacing the current petroleum-based processes with bio-based processes.
Professor Lee said, “Our study suggested a possibility to produce 1,3-diaminopropane based on biorefinery. Further study will be done to increase the titer and productivity of 1,3-diaminopropane.”
This work was published online in Scientific Reports on August 11, 2015.
Reference: Chae, T.U. et al. "Metabolic engineering of Escherichia coli for the production of 1,3-diaminopropane, a three carbon diamine," Scientific Reports:
http://www.nature.com/articles/srep13040
This research was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from Ministry of Science, ICT and Future Planning (MSIP) through the National Research Foundation (NRF) of Korea.
Figure 1: Metabolic engineering strategies for 1,3-diaminopropane production using C4 pathway
Figure 2: Fed-batch fermentation profiles of two final engineered E. coli strains
2015.08.12 View 11898