network
-
 KAIST Accelerates Synthetic Microbe Design by Discovering Novel Enzymes Using AI
< (From left) Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering (top), Hongkeun Ji, PhD candidate of the Department of Chemical and Biomolecular Engineering (top), Ha Rim Kim, PhD candidate of the Department of Chemical and Biomolecular Engineering, and Dr. Gi Bae Kim of the BioProcess Engineering Research Center >
Enzymes are proteins that catalyze biochemical reactions within cells and play a pivotal role in metabolic processes. Accordingly, identifying the functions of novel enzymes is a critical task in the construction of microbial cell factories.
A KAIST research team has leveraged artificial intelligence (AI) to design novel enzymes that do not exist in nature, significantly accelerating microbial cell factory development and boosting the potential for next-generation biotechnological applications such as drug development and biofuel production.
KAIST (represented by President Kwang-Hyung Lee) announced on the 21st of April that Distinguished Professor Sang Yup Lee and his team from the Department of Chemical and Biomolecular Engineering have published a review titled “Enzyme Functional Classification Using Artificial Intelligence,” which outlines the advancement of AI-based enzyme function prediction technologies and analyzes how AI has contributed to the discovery and design of new enzymes.
Professor Lee’s team systematically reviewed the development of enzyme function prediction technologies utilizing machine learning and deep learning, offering a comprehensive analysis.
From sequence similarity-based prediction methods to the integration of convolutional neural networks (CNNs), recurrent neural networks (RNNs), graph neural networks (GNNs), and transformer-based large language models, the paper covers a broad range of AI applications. It analyzes how these technologies extract meaningful information from protein sequences and enhance prediction accuracy.
In particular, enzyme function prediction using deep learning goes beyond simple sequence similarity analysis. By automatically extracting structural and evolutionary features embedded in amino acid sequences, deep learning enables more precise predictions of catalytic functions.
This highlights the unique advantages of AI models compared to traditional bioinformatics approaches.
Moreover, the review suggests that the advancement of generative AI will move future research beyond predicting existing functions to generating entirely new enzymes with functions not found in nature. This shift is expected to profoundly impact the trajectory of biotechnology and synthetic biology.
< Figure 1. Extraction of enzyme characteristics and function prediction using various deep learning structures >
Ha Rim Kim, a Ph.D. candidate and co-first author from the Department of Chemical and Biomolecular Engineering, stated, “AI-based enzyme function prediction and enzyme design are highly important across various fields including metabolic engineering, synthetic biology, and healthcare.”
Distinguished Professor Sang Yup Lee added, “AI-powered enzyme function prediction shows the potential to solve diverse biological problems and will significantly contribute to accelerating research across the entire field.”
The review was published on March 28 in Trends in Biotechnology, a leading biotechnology journal issued by Cell Press.
※ Title: Enzyme Functional Classification Using Artificial Intelligence
※DOI: https://doi.org/10.1016/j.tibtech.2025.03.003
※ Author Information: Ha Rim Kim (KAIST, Co-first author), Hongkeun Ji (KAIST, Co-first author), Gi Bae Kim (KAIST, Third author), Sang Yup Lee (KAIST, Corresponding author)
This research was supported by the Ministry of Science and ICT under the project Development of Core Technologies for Advanced Synthetic Biology to Lead the Bio-Manufacturing Industry (aimed at replacing petroleum-based chemicals), and also by joint support from the Ministry of Science and ICT and the Ministry of Health and Welfare for the project Development of Novel Antibiotic Structures Using Deep Learning-Based Synthetic Biology.
2025.04.07 View 4879
KAIST Accelerates Synthetic Microbe Design by Discovering Novel Enzymes Using AI
< (From left) Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering (top), Hongkeun Ji, PhD candidate of the Department of Chemical and Biomolecular Engineering (top), Ha Rim Kim, PhD candidate of the Department of Chemical and Biomolecular Engineering, and Dr. Gi Bae Kim of the BioProcess Engineering Research Center >
Enzymes are proteins that catalyze biochemical reactions within cells and play a pivotal role in metabolic processes. Accordingly, identifying the functions of novel enzymes is a critical task in the construction of microbial cell factories.
A KAIST research team has leveraged artificial intelligence (AI) to design novel enzymes that do not exist in nature, significantly accelerating microbial cell factory development and boosting the potential for next-generation biotechnological applications such as drug development and biofuel production.
KAIST (represented by President Kwang-Hyung Lee) announced on the 21st of April that Distinguished Professor Sang Yup Lee and his team from the Department of Chemical and Biomolecular Engineering have published a review titled “Enzyme Functional Classification Using Artificial Intelligence,” which outlines the advancement of AI-based enzyme function prediction technologies and analyzes how AI has contributed to the discovery and design of new enzymes.
Professor Lee’s team systematically reviewed the development of enzyme function prediction technologies utilizing machine learning and deep learning, offering a comprehensive analysis.
From sequence similarity-based prediction methods to the integration of convolutional neural networks (CNNs), recurrent neural networks (RNNs), graph neural networks (GNNs), and transformer-based large language models, the paper covers a broad range of AI applications. It analyzes how these technologies extract meaningful information from protein sequences and enhance prediction accuracy.
In particular, enzyme function prediction using deep learning goes beyond simple sequence similarity analysis. By automatically extracting structural and evolutionary features embedded in amino acid sequences, deep learning enables more precise predictions of catalytic functions.
This highlights the unique advantages of AI models compared to traditional bioinformatics approaches.
Moreover, the review suggests that the advancement of generative AI will move future research beyond predicting existing functions to generating entirely new enzymes with functions not found in nature. This shift is expected to profoundly impact the trajectory of biotechnology and synthetic biology.
< Figure 1. Extraction of enzyme characteristics and function prediction using various deep learning structures >
Ha Rim Kim, a Ph.D. candidate and co-first author from the Department of Chemical and Biomolecular Engineering, stated, “AI-based enzyme function prediction and enzyme design are highly important across various fields including metabolic engineering, synthetic biology, and healthcare.”
Distinguished Professor Sang Yup Lee added, “AI-powered enzyme function prediction shows the potential to solve diverse biological problems and will significantly contribute to accelerating research across the entire field.”
The review was published on March 28 in Trends in Biotechnology, a leading biotechnology journal issued by Cell Press.
※ Title: Enzyme Functional Classification Using Artificial Intelligence
※DOI: https://doi.org/10.1016/j.tibtech.2025.03.003
※ Author Information: Ha Rim Kim (KAIST, Co-first author), Hongkeun Ji (KAIST, Co-first author), Gi Bae Kim (KAIST, Third author), Sang Yup Lee (KAIST, Corresponding author)
This research was supported by the Ministry of Science and ICT under the project Development of Core Technologies for Advanced Synthetic Biology to Lead the Bio-Manufacturing Industry (aimed at replacing petroleum-based chemicals), and also by joint support from the Ministry of Science and ICT and the Ministry of Health and Welfare for the project Development of Novel Antibiotic Structures Using Deep Learning-Based Synthetic Biology.
2025.04.07 View 4879 -
 KAIST Discovers Molecular Switch that Reverses Cancerous Transformation at the Critical Moment of Transition
< (From left) PhD student Seoyoon D. Jeong, (bottom) Professor Kwang-Hyun Cho, (top) Dr. Dongkwan Shin, Dr. Jeong-Ryeol Gong >
Professor Kwang-Hyun Cho’s research team has recently been highlighted for their work on developing an original technology for cancer reversal treatment that does not kill cancer cells but only changes their characteristics to reverse them to a state similar to normal cells. This time, they have succeeded in revealing for the first time that a molecular switch that can induce cancer reversal at the moment when normal cells change into cancer cells is hidden in the genetic network.
KAIST (President Kwang-Hyung Lee) announced on the 5th of February that Professor Kwang-Hyun Cho's research team of the Department of Bio and Brain Engineering has succeeded in developing a fundamental technology to capture the critical transition phenomenon at the moment when normal cells change into cancer cells and analyze it to discover a molecular switch that can revert cancer cells back into normal cells.
A critical transition is a phenomenon in which a sudden change in state occurs at a specific point in time, like water changing into steam at 100℃. This critical transition phenomenon also occurs in the process in which normal cells change into cancer cells at a specific point in time due to the accumulation of genetic and epigenetic changes.
The research team discovered that normal cells can enter an unstable critical transition state where normal cells and cancer cells coexist just before they change into cancer cells during tumorigenesis, the production or development of tumors, and analyzed this critical transition state using a systems biology method to develop a cancer reversal molecular switch identification technology that can reverse the cancerization process. They then applied this to colon cancer cells and confirmed through molecular cell experiments that cancer cells can recover the characteristics of normal cells.
This is an original technology that automatically infers a computer model of the genetic network that controls the critical transition of cancer development from single-cell RNA sequencing data, and systematically finds molecular switches for cancer reversion by simulation analysis. It is expected that this technology will be applied to the development of reversion therapies for other cancers in the future.
Professor Kwang-Hyun Cho said, "We have discovered a molecular switch that can revert the fate of cancer cells back to a normal state by capturing the moment of critical transition right before normal cells are changed into an irreversible cancerous state."
< Figure 1. Overall conceptual framework of the technology that automatically constructs a molecular regulatory network from single-cell RNA sequencing data of colon cancer cells to discover molecular switches for cancer reversion through computer simulation analysis. Professor Kwang-Hyun Cho's research team established a fundamental technology for automatic construction of a computer model of a core gene network by analyzing the entire process of tumorigenesis of colon cells turning into cancer cells, and developed an original technology for discovering the molecular switches that can induce cancer cell reversal through attractor landscape analysis. >
He continued, "In particular, this study has revealed in detail, at the genetic network level, what changes occur within cells behind the process of cancer development, which has been considered a mystery until now." He emphasized, "This is the first study to reveal that an important clue that can revert the fate of tumorigenesis is hidden at this very critical moment of change."
< Figure 2. Identification of tumor transition state using single-cell RNA sequencing data from colorectal cancer. Using single-cell RNA sequencing data from colorectal cancer patient-derived organoids for normal and cancerous tissues, a critical transition was identified in which normal and cancerous cells coexist and instability increases (a-d). The critical transition was confirmed to show intermediate levels of major phenotypic features related to cancer or normal tissues that are indicative of the states between the normal and cancerous cells (e). >
The results of this study, conducted by KAIST Dr. Dongkwan Shin (currently at the National Cancer Center), Dr. Jeong-Ryeol Gong, and doctoral student Seoyoon D. Jeong jointly with a research team at Seoul National University that provided the organoids (in vitro cultured tissues) from colon cancer patient, were published as an online paper in the international journal ‘Advanced Science’ published by Wiley on January 22nd.
(Paper title: Attractor landscape analysis reveals a reversion switch in the transition of colorectal tumorigenesis) (DOI: https://doi.org/10.1002/advs.202412503)
< Figure 3. Reconstruction of a dynamic network model for the transition state of colorectal cancer.
A new technology was established to build a gene network computer model that can simulate the dynamic changes between genes by integrating single-cell RNA sequencing data and existing experimental results on gene-to-gene interactions in the critical transition of cancer. (a). Using this technology, a gene network computer model for the critical transition of colorectal cancer was constructed, and the distribution of attractors representing normal and cancer cell phenotypes was investigated through attractor landscape analysis (b-e). >
This study was conducted with the support of the National Research Foundation of Korea under the Ministry of Science and ICT through the Mid-Career Researcher Program and Basic Research Laboratory Program and the Disease-Centered Translational Research Project of the Korea Health Industry Development Institute (KHIDI) of the Ministry of Health and Welfare.
< Figure 4. Quantification of attractor landscapes and discovery of transcription factors for cancer reversibility through perturbation simulation analysis. A methodology for implementing discontinuous attractor landscapes continuously from a computer model of gene networks and quantifying them as cancer scores was introduced (a), and attractor landscapes for the critical transition of colorectal cancer were secured (b-d). By tracking the change patterns of normal and cancer cell attractors through perturbation simulation analysis for each gene, the optimal combination of transcription factors for cancer reversion was discovered (e-h). This was confirmed in various parameter combinations as well (i). >
< Figure 5. Identification and experimental validation of the optimal target gene for cancer reversion. Among the common target genes of the discovered transcription factor combinations, we identified cancer reversing molecular switches that are predicted to suppress cancer cell proliferation and restore the characteristics of normal colon cells (a-d). When inhibitors for the molecular switches were treated to organoids derived from colon cancer patients, it was confirmed that cancer cell proliferation was suppressed and the expression of key genes related to cancer development was inhibited (e-h), and a group of genes related to normal colon epithelium was activated and transformed into a state similar to normal colon cells (i-j). >
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed an original technology to systematically discover key molecular switches that can induce reversion of colon cancer cells through a systems biology approach using an attractor landscape analysis of a genetic network model for the critical transition at the moment of transformation from normal cells to cancer cells, and verified the reversing effect of actual colon cancer through cellular experiments. >
2025.02.05 View 29896
KAIST Discovers Molecular Switch that Reverses Cancerous Transformation at the Critical Moment of Transition
< (From left) PhD student Seoyoon D. Jeong, (bottom) Professor Kwang-Hyun Cho, (top) Dr. Dongkwan Shin, Dr. Jeong-Ryeol Gong >
Professor Kwang-Hyun Cho’s research team has recently been highlighted for their work on developing an original technology for cancer reversal treatment that does not kill cancer cells but only changes their characteristics to reverse them to a state similar to normal cells. This time, they have succeeded in revealing for the first time that a molecular switch that can induce cancer reversal at the moment when normal cells change into cancer cells is hidden in the genetic network.
KAIST (President Kwang-Hyung Lee) announced on the 5th of February that Professor Kwang-Hyun Cho's research team of the Department of Bio and Brain Engineering has succeeded in developing a fundamental technology to capture the critical transition phenomenon at the moment when normal cells change into cancer cells and analyze it to discover a molecular switch that can revert cancer cells back into normal cells.
A critical transition is a phenomenon in which a sudden change in state occurs at a specific point in time, like water changing into steam at 100℃. This critical transition phenomenon also occurs in the process in which normal cells change into cancer cells at a specific point in time due to the accumulation of genetic and epigenetic changes.
The research team discovered that normal cells can enter an unstable critical transition state where normal cells and cancer cells coexist just before they change into cancer cells during tumorigenesis, the production or development of tumors, and analyzed this critical transition state using a systems biology method to develop a cancer reversal molecular switch identification technology that can reverse the cancerization process. They then applied this to colon cancer cells and confirmed through molecular cell experiments that cancer cells can recover the characteristics of normal cells.
This is an original technology that automatically infers a computer model of the genetic network that controls the critical transition of cancer development from single-cell RNA sequencing data, and systematically finds molecular switches for cancer reversion by simulation analysis. It is expected that this technology will be applied to the development of reversion therapies for other cancers in the future.
Professor Kwang-Hyun Cho said, "We have discovered a molecular switch that can revert the fate of cancer cells back to a normal state by capturing the moment of critical transition right before normal cells are changed into an irreversible cancerous state."
< Figure 1. Overall conceptual framework of the technology that automatically constructs a molecular regulatory network from single-cell RNA sequencing data of colon cancer cells to discover molecular switches for cancer reversion through computer simulation analysis. Professor Kwang-Hyun Cho's research team established a fundamental technology for automatic construction of a computer model of a core gene network by analyzing the entire process of tumorigenesis of colon cells turning into cancer cells, and developed an original technology for discovering the molecular switches that can induce cancer cell reversal through attractor landscape analysis. >
He continued, "In particular, this study has revealed in detail, at the genetic network level, what changes occur within cells behind the process of cancer development, which has been considered a mystery until now." He emphasized, "This is the first study to reveal that an important clue that can revert the fate of tumorigenesis is hidden at this very critical moment of change."
< Figure 2. Identification of tumor transition state using single-cell RNA sequencing data from colorectal cancer. Using single-cell RNA sequencing data from colorectal cancer patient-derived organoids for normal and cancerous tissues, a critical transition was identified in which normal and cancerous cells coexist and instability increases (a-d). The critical transition was confirmed to show intermediate levels of major phenotypic features related to cancer or normal tissues that are indicative of the states between the normal and cancerous cells (e). >
The results of this study, conducted by KAIST Dr. Dongkwan Shin (currently at the National Cancer Center), Dr. Jeong-Ryeol Gong, and doctoral student Seoyoon D. Jeong jointly with a research team at Seoul National University that provided the organoids (in vitro cultured tissues) from colon cancer patient, were published as an online paper in the international journal ‘Advanced Science’ published by Wiley on January 22nd.
(Paper title: Attractor landscape analysis reveals a reversion switch in the transition of colorectal tumorigenesis) (DOI: https://doi.org/10.1002/advs.202412503)
< Figure 3. Reconstruction of a dynamic network model for the transition state of colorectal cancer.
A new technology was established to build a gene network computer model that can simulate the dynamic changes between genes by integrating single-cell RNA sequencing data and existing experimental results on gene-to-gene interactions in the critical transition of cancer. (a). Using this technology, a gene network computer model for the critical transition of colorectal cancer was constructed, and the distribution of attractors representing normal and cancer cell phenotypes was investigated through attractor landscape analysis (b-e). >
This study was conducted with the support of the National Research Foundation of Korea under the Ministry of Science and ICT through the Mid-Career Researcher Program and Basic Research Laboratory Program and the Disease-Centered Translational Research Project of the Korea Health Industry Development Institute (KHIDI) of the Ministry of Health and Welfare.
< Figure 4. Quantification of attractor landscapes and discovery of transcription factors for cancer reversibility through perturbation simulation analysis. A methodology for implementing discontinuous attractor landscapes continuously from a computer model of gene networks and quantifying them as cancer scores was introduced (a), and attractor landscapes for the critical transition of colorectal cancer were secured (b-d). By tracking the change patterns of normal and cancer cell attractors through perturbation simulation analysis for each gene, the optimal combination of transcription factors for cancer reversion was discovered (e-h). This was confirmed in various parameter combinations as well (i). >
< Figure 5. Identification and experimental validation of the optimal target gene for cancer reversion. Among the common target genes of the discovered transcription factor combinations, we identified cancer reversing molecular switches that are predicted to suppress cancer cell proliferation and restore the characteristics of normal colon cells (a-d). When inhibitors for the molecular switches were treated to organoids derived from colon cancer patients, it was confirmed that cancer cell proliferation was suppressed and the expression of key genes related to cancer development was inhibited (e-h), and a group of genes related to normal colon epithelium was activated and transformed into a state similar to normal colon cells (i-j). >
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed an original technology to systematically discover key molecular switches that can induce reversion of colon cancer cells through a systems biology approach using an attractor landscape analysis of a genetic network model for the critical transition at the moment of transformation from normal cells to cancer cells, and verified the reversing effect of actual colon cancer through cellular experiments. >
2025.02.05 View 29896 -
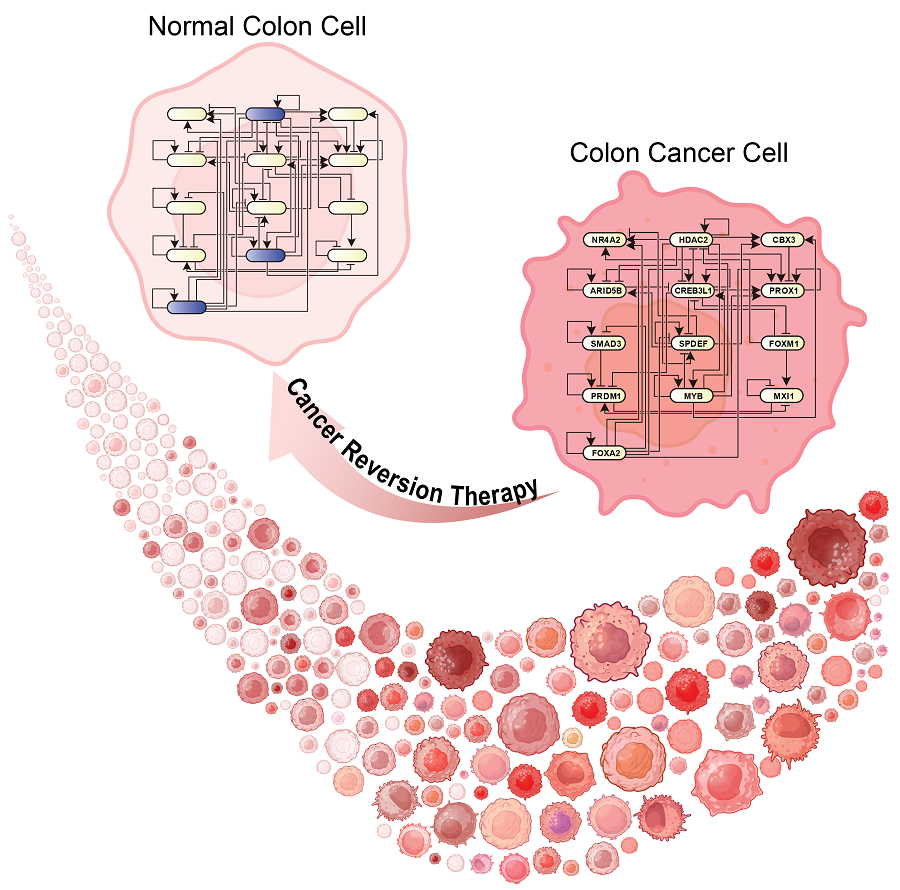 KAIST Develops Foundational Technology to Revert Cancer Cells to Normal Cells
Despite the development of numerous cancer treatment technologies, the common goal of current cancer therapies is to eliminate cancer cells. This approach, however, faces fundamental limitations, including cancer cells developing resistance and returning, as well as severe side effects from the destruction of healthy cells.
< (From top left) Bio and Brain Engineering PhD candidates Juhee Kim, Jeong-Ryeol Gong, Chun-Kyung Lee, and Hoon-Min Kim posed for a group photo with Professor Kwang-Hyun Cho >
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of December that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering has developed a groundbreaking technology that can treat colon cancer by converting cancer cells into a state resembling normal colon cells without killing them, thus avoiding side effects.
The research team focused on the observation that during the oncogenesis process, normal cells regress along their differentiation trajectory. Building on this insight, they developed a technology to create a digital twin of the gene network associated with the differentiation trajectory of normal cells.
< Figure 1. Technology for creating a digital twin of a gene network from single-cell transcriptome data of a normal cell differentiation trajectory. Professor Kwang-Hyun Cho's research team developed a digital twin creation technology that precisely observes the dynamics of gene regulatory relationships during the process of normal cells differentiating along a differentiation trajectory and analyzes the relationships among key genes to build a mathematical model that can be simulated (A-F). In addition, they developed a technology to discover key regulatory factors that control the differentiation trajectory of normal cells by simulating and analyzing this digital twin. >
< Figure 2. Digital twin simulation simulating the differentiation trajectory of normal colon cells. The dynamics of single-cell transcriptome data for the differentiation trajectory of normal colon cells were analyzed (A) and a digital twin of the gene network was developed representing the regulatory relationships of key genes in this differentiation trajectory (B). The simulation results of the digital twin confirm that it readily reproduces the dynamics of single-cell transcriptome data (C, D). >
Through simulation analysis, the team systematically identified master molecular switches that induce normal cell differentiation. When these switches were applied to colon cancer cells, the cancer cells reverted to a normal-like state, a result confirmed through molecular and cellular experiments as well as animal studies.
< Figure 3. Discovery of top-level key control factors that induce differentiation of normal colon cells. By applying control factor discovery technology to the digital twin model, three genes, HDAC2, FOXA2, and MYB, were discovered as key control factors that induce differentiation of normal colon cells (A, B). The results of simulation analysis of the regulatory effects of the discovered control factors through the digital twin confirmed that they could induce complete differentiation of colon cells (C). >
< Figure 4. Verification of the effect of the key control factors discovered using colon cancer cells and animal experiments on the reversibility of colon cancer. The key control factors of the normal colon cell differentiation trajectory discovered through digital twin simulation analysis were applied to actual colon cancer cells and colon cancer mouse animal models to experimentally verify the effect of cancer reversibility. The key control factors significantly reduced the proliferation of three colon cancer cell lines (A), and this was confirmed in the same way in animal models (B-D). >
This research demonstrates that cancer cell reversion can be systematically achieved by analyzing and utilizing the digital twin of the cancer cell gene network, rather than relying on serendipitous discoveries. The findings hold significant promise for developing reversible cancer therapies that can be applied to various types of cancer.
< Figure 5. The change in overall gene expression was confirmed through the regulation of the identified key regulatory factors, which converted the state of colon cancer cells to that of normal colon cells. The transcriptomes of colon cancer tissues and normal colon tissues from more than 400 colon cancer patients were compared with the transcriptomes of colon cancer cell lines and reversible colon cancer cell lines, respectively. The comparison results confirmed that the regulation of the identified key regulatory factors converted all three colon cancer cell lines to a state similar to the transcriptome expression of normal colon tissues. >
Professor Kwang-Hyun Cho remarked, "The fact that cancer cells can be converted back to normal cells is an astonishing phenomenon. This study proves that such reversion can be systematically induced."
He further emphasized, "This research introduces the novel concept of reversible cancer therapy by reverting cancer cells to normal cells. It also develops foundational technology for identifying targets for cancer reversion through the systematic analysis of normal cell differentiation trajectories."
This research included contributions from Jeong-Ryeol Gong, Chun-Kyung Lee, Hoon-Min Kim, Juhee Kim, and Jaeog Jeon, and was published in the online edition of the international journal Advanced Science by Wiley on December 11. (Title: “Control of Cellular Differentiation Trajectories for Cancer Reversion”) DOI: https://doi.org/10.1002/advs.202402132
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed a source technology to systematically discover key control factors that can induce reversibility of colon cancer cells through a systems biology approach and a digital twin simulation analysis of the differentiation trajectory of normal colon cells, and verified the effects of reversion on actual colon cancer through molecular cell experiments and animal experiments. >
The study was supported by the Ministry of Science and ICT and the National Research Foundation of Korea through the Mid-Career Researcher Program and Basic Research Laboratory Program. The research findings have been transferred to BioRevert Inc., where they will be used for the development of practical cancer reversion therapies.
2024.12.23 View 107179
KAIST Develops Foundational Technology to Revert Cancer Cells to Normal Cells
Despite the development of numerous cancer treatment technologies, the common goal of current cancer therapies is to eliminate cancer cells. This approach, however, faces fundamental limitations, including cancer cells developing resistance and returning, as well as severe side effects from the destruction of healthy cells.
< (From top left) Bio and Brain Engineering PhD candidates Juhee Kim, Jeong-Ryeol Gong, Chun-Kyung Lee, and Hoon-Min Kim posed for a group photo with Professor Kwang-Hyun Cho >
KAIST (represented by President Kwang Hyung Lee) announced on the 20th of December that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering has developed a groundbreaking technology that can treat colon cancer by converting cancer cells into a state resembling normal colon cells without killing them, thus avoiding side effects.
The research team focused on the observation that during the oncogenesis process, normal cells regress along their differentiation trajectory. Building on this insight, they developed a technology to create a digital twin of the gene network associated with the differentiation trajectory of normal cells.
< Figure 1. Technology for creating a digital twin of a gene network from single-cell transcriptome data of a normal cell differentiation trajectory. Professor Kwang-Hyun Cho's research team developed a digital twin creation technology that precisely observes the dynamics of gene regulatory relationships during the process of normal cells differentiating along a differentiation trajectory and analyzes the relationships among key genes to build a mathematical model that can be simulated (A-F). In addition, they developed a technology to discover key regulatory factors that control the differentiation trajectory of normal cells by simulating and analyzing this digital twin. >
< Figure 2. Digital twin simulation simulating the differentiation trajectory of normal colon cells. The dynamics of single-cell transcriptome data for the differentiation trajectory of normal colon cells were analyzed (A) and a digital twin of the gene network was developed representing the regulatory relationships of key genes in this differentiation trajectory (B). The simulation results of the digital twin confirm that it readily reproduces the dynamics of single-cell transcriptome data (C, D). >
Through simulation analysis, the team systematically identified master molecular switches that induce normal cell differentiation. When these switches were applied to colon cancer cells, the cancer cells reverted to a normal-like state, a result confirmed through molecular and cellular experiments as well as animal studies.
< Figure 3. Discovery of top-level key control factors that induce differentiation of normal colon cells. By applying control factor discovery technology to the digital twin model, three genes, HDAC2, FOXA2, and MYB, were discovered as key control factors that induce differentiation of normal colon cells (A, B). The results of simulation analysis of the regulatory effects of the discovered control factors through the digital twin confirmed that they could induce complete differentiation of colon cells (C). >
< Figure 4. Verification of the effect of the key control factors discovered using colon cancer cells and animal experiments on the reversibility of colon cancer. The key control factors of the normal colon cell differentiation trajectory discovered through digital twin simulation analysis were applied to actual colon cancer cells and colon cancer mouse animal models to experimentally verify the effect of cancer reversibility. The key control factors significantly reduced the proliferation of three colon cancer cell lines (A), and this was confirmed in the same way in animal models (B-D). >
This research demonstrates that cancer cell reversion can be systematically achieved by analyzing and utilizing the digital twin of the cancer cell gene network, rather than relying on serendipitous discoveries. The findings hold significant promise for developing reversible cancer therapies that can be applied to various types of cancer.
< Figure 5. The change in overall gene expression was confirmed through the regulation of the identified key regulatory factors, which converted the state of colon cancer cells to that of normal colon cells. The transcriptomes of colon cancer tissues and normal colon tissues from more than 400 colon cancer patients were compared with the transcriptomes of colon cancer cell lines and reversible colon cancer cell lines, respectively. The comparison results confirmed that the regulation of the identified key regulatory factors converted all three colon cancer cell lines to a state similar to the transcriptome expression of normal colon tissues. >
Professor Kwang-Hyun Cho remarked, "The fact that cancer cells can be converted back to normal cells is an astonishing phenomenon. This study proves that such reversion can be systematically induced."
He further emphasized, "This research introduces the novel concept of reversible cancer therapy by reverting cancer cells to normal cells. It also develops foundational technology for identifying targets for cancer reversion through the systematic analysis of normal cell differentiation trajectories."
This research included contributions from Jeong-Ryeol Gong, Chun-Kyung Lee, Hoon-Min Kim, Juhee Kim, and Jaeog Jeon, and was published in the online edition of the international journal Advanced Science by Wiley on December 11. (Title: “Control of Cellular Differentiation Trajectories for Cancer Reversion”) DOI: https://doi.org/10.1002/advs.202402132
< Figure 6. Schematic diagram of the research results. Professor Kwang-Hyun Cho's research team developed a source technology to systematically discover key control factors that can induce reversibility of colon cancer cells through a systems biology approach and a digital twin simulation analysis of the differentiation trajectory of normal colon cells, and verified the effects of reversion on actual colon cancer through molecular cell experiments and animal experiments. >
The study was supported by the Ministry of Science and ICT and the National Research Foundation of Korea through the Mid-Career Researcher Program and Basic Research Laboratory Program. The research findings have been transferred to BioRevert Inc., where they will be used for the development of practical cancer reversion therapies.
2024.12.23 View 107179 -
 KAIST Proposes a New Way to Circumvent a Long-time Frustration in Neural Computing
The human brain begins learning through spontaneous random activities even before it receives sensory information from the external world. The technology developed by the KAIST research team enables much faster and more accurate learning when exposed to actual data by pre-learning random information in a brain-mimicking artificial neural network, and is expected to be a breakthrough in the development of brain-based artificial intelligence and neuromorphic computing technology in the future.
KAIST (President Kwang-Hyung Lee) announced on the 16th of December that Professor Se-Bum Paik 's research team in the Department of Brain Cognitive Sciences solved the weight transport problem*, a long-standing challenge in neural network learning, and through this, explained the principles that enable resource-efficient learning in biological brain neural networks.
*Weight transport problem: This is the biggest obstacle to the development of artificial intelligence that mimics the biological brain. It is the fundamental reason why large-scale memory and computational work are required in the learning of general artificial neural networks, unlike biological brains.
Over the past several decades, the development of artificial intelligence has been based on error backpropagation learning proposed by Geoffery Hinton, who won the Nobel Prize in Physics this year. However, error backpropagation learning was thought to be impossible in biological brains because it requires the unrealistic assumption that individual neurons must know all the connected information across multiple layers in order to calculate the error signal for learning.
< Figure 1. Illustration depicting the method of random noise training and its effects >
This difficult problem, called the weight transport problem, was raised by Francis Crick, who won the Nobel Prize in Physiology or Medicine for the discovery of the structure of DNA, after the error backpropagation learning was proposed by Hinton in 1986. Since then, it has been considered the reason why the operating principles of natural neural networks and artificial neural networks will forever be fundamentally different.
At the borderline of artificial intelligence and neuroscience, researchers including Hinton have continued to attempt to create biologically plausible models that can implement the learning principles of the brain by solving the weight transport problem.
In 2016, a joint research team from Oxford University and DeepMind in the UK first proposed the concept of error backpropagation learning being possible without weight transport, drawing attention from the academic world. However, biologically plausible error backpropagation learning without weight transport was inefficient, with slow learning speeds and low accuracy, making it difficult to apply in reality.
KAIST research team noted that the biological brain begins learning through internal spontaneous random neural activity even before experiencing external sensory experiences. To mimic this, the research team pre-trained a biologically plausible neural network without weight transport with meaningless random information (random noise).
As a result, they showed that the symmetry of the forward and backward neural cell connections of the neural network, which is an essential condition for error backpropagation learning, can be created. In other words, learning without weight transport is possible through random pre-training.
< Figure 2. Illustration depicting the meta-learning effect of random noise training >
The research team revealed that learning random information before learning actual data has the property of meta-learning, which is ‘learning how to learn.’ It was shown that neural networks that pre-learned random noise perform much faster and more accurate learning when exposed to actual data, and can achieve high learning efficiency without weight transport.
< Figure 3. Illustration depicting research on understanding the brain's operating principles through artificial neural networks >
Professor Se-Bum Paik said, “It breaks the conventional understanding of existing machine learning that only data learning is important, and provides a new perspective that focuses on the neuroscience principles of creating appropriate conditions before learning,” and added, “It is significant in that it solves important problems in artificial neural network learning through clues from developmental neuroscience, and at the same time provides insight into the brain’s learning principles through artificial neural network models.”
This study, in which Jeonghwan Cheon, a Master’s candidate of KAIST Department of Brain and Cognitive Sciences participated as the first author and Professor Sang Wan Lee of the same department as a co-author, was presented at the 38th Neural Information Processing Systems (NeurIPS), the world's top artificial intelligence conference, on December 14th in Vancouver, Canada. (Paper title: Pretraining with random noise for fast and robust learning without weight transport)
This study was conducted with the support of the National Research Foundation of Korea's Basic Research Program in Science and Engineering, the Information and Communications Technology Planning and Evaluation Institute's Talent Development Program, and the KAIST Singularity Professor Program.
2024.12.16 View 9165
KAIST Proposes a New Way to Circumvent a Long-time Frustration in Neural Computing
The human brain begins learning through spontaneous random activities even before it receives sensory information from the external world. The technology developed by the KAIST research team enables much faster and more accurate learning when exposed to actual data by pre-learning random information in a brain-mimicking artificial neural network, and is expected to be a breakthrough in the development of brain-based artificial intelligence and neuromorphic computing technology in the future.
KAIST (President Kwang-Hyung Lee) announced on the 16th of December that Professor Se-Bum Paik 's research team in the Department of Brain Cognitive Sciences solved the weight transport problem*, a long-standing challenge in neural network learning, and through this, explained the principles that enable resource-efficient learning in biological brain neural networks.
*Weight transport problem: This is the biggest obstacle to the development of artificial intelligence that mimics the biological brain. It is the fundamental reason why large-scale memory and computational work are required in the learning of general artificial neural networks, unlike biological brains.
Over the past several decades, the development of artificial intelligence has been based on error backpropagation learning proposed by Geoffery Hinton, who won the Nobel Prize in Physics this year. However, error backpropagation learning was thought to be impossible in biological brains because it requires the unrealistic assumption that individual neurons must know all the connected information across multiple layers in order to calculate the error signal for learning.
< Figure 1. Illustration depicting the method of random noise training and its effects >
This difficult problem, called the weight transport problem, was raised by Francis Crick, who won the Nobel Prize in Physiology or Medicine for the discovery of the structure of DNA, after the error backpropagation learning was proposed by Hinton in 1986. Since then, it has been considered the reason why the operating principles of natural neural networks and artificial neural networks will forever be fundamentally different.
At the borderline of artificial intelligence and neuroscience, researchers including Hinton have continued to attempt to create biologically plausible models that can implement the learning principles of the brain by solving the weight transport problem.
In 2016, a joint research team from Oxford University and DeepMind in the UK first proposed the concept of error backpropagation learning being possible without weight transport, drawing attention from the academic world. However, biologically plausible error backpropagation learning without weight transport was inefficient, with slow learning speeds and low accuracy, making it difficult to apply in reality.
KAIST research team noted that the biological brain begins learning through internal spontaneous random neural activity even before experiencing external sensory experiences. To mimic this, the research team pre-trained a biologically plausible neural network without weight transport with meaningless random information (random noise).
As a result, they showed that the symmetry of the forward and backward neural cell connections of the neural network, which is an essential condition for error backpropagation learning, can be created. In other words, learning without weight transport is possible through random pre-training.
< Figure 2. Illustration depicting the meta-learning effect of random noise training >
The research team revealed that learning random information before learning actual data has the property of meta-learning, which is ‘learning how to learn.’ It was shown that neural networks that pre-learned random noise perform much faster and more accurate learning when exposed to actual data, and can achieve high learning efficiency without weight transport.
< Figure 3. Illustration depicting research on understanding the brain's operating principles through artificial neural networks >
Professor Se-Bum Paik said, “It breaks the conventional understanding of existing machine learning that only data learning is important, and provides a new perspective that focuses on the neuroscience principles of creating appropriate conditions before learning,” and added, “It is significant in that it solves important problems in artificial neural network learning through clues from developmental neuroscience, and at the same time provides insight into the brain’s learning principles through artificial neural network models.”
This study, in which Jeonghwan Cheon, a Master’s candidate of KAIST Department of Brain and Cognitive Sciences participated as the first author and Professor Sang Wan Lee of the same department as a co-author, was presented at the 38th Neural Information Processing Systems (NeurIPS), the world's top artificial intelligence conference, on December 14th in Vancouver, Canada. (Paper title: Pretraining with random noise for fast and robust learning without weight transport)
This study was conducted with the support of the National Research Foundation of Korea's Basic Research Program in Science and Engineering, the Information and Communications Technology Planning and Evaluation Institute's Talent Development Program, and the KAIST Singularity Professor Program.
2024.12.16 View 9165 -
 KAIST Proposes AI Training Method that will Drastically Shorten Time for Complex Quantum Mechanical Calculations
- Professor Yong-Hoon Kim's team from the School of Electrical Engineering succeeded for the first time in accelerating quantum mechanical electronic structure calculations using a convolutional neural network (CNN) model
- Presenting an AI learning principle of quantum mechanical 3D chemical bonding information, the work is expected to accelerate the computer-assisted designing of next-generation materials and devices
The close relationship between AI and high-performance scientific computing can be seen in the fact that both the 2024 Nobel Prizes in Physics and Chemistry were awarded to scientists for their AI-related research contributions in their respective fields of study. KAIST researchers succeeded in dramatically reducing the computation time for highly sophisticated quantum mechanical computer simulations by predicting atomic-level chemical bonding information distributed in 3D space using a novel AI approach.
KAIST (President Kwang-Hyung Lee) announced on the 30th of October that Professor Yong-Hoon Kim's team from the School of Electrical Engineering developed a 3D computer vision artificial neural network-based computation methodology that bypasses the complex algorithms required for atomic-level quantum mechanical calculations traditionally performed using supercomputers to derive the properties of materials.
< Figure 1. Various methodologies are utilized in the simulation of materials and materials, such as quantum mechanical calculations at the nanometer (nm) level, classical mechanical force fields at the scale of tens to hundreds of nanometers, continuum dynamics calculations at the macroscopic scale, and calculations that mix simulations at different scales. These simulations are already playing a key role in a wide range of basic research and application development fields in combination with informatics techniques. Recently, there have been active efforts to introduce machine learning techniques to radically accelerate simulations, but research on introducing machine learning techniques to quantum mechanical electronic structure calculations, which form the basis of high-scale simulations, is still insufficient. >
The quantum mechanical density functional theory (DFT) calculations using supercomputers have become an essential and standard tool in a wide range of research and development fields, including advanced materials and drug design, as they allow fast and accurate prediction of material properties.
*Density functional theory (DFT): A representative theory of ab initio (first principles) calculations that calculate quantum mechanical properties from the atomic level.
However, practical DFT calculations require generating 3D electron density and solving quantum mechanical equations through a complex, iterative self-consistent field (SCF)* process that must be repeated tens to hundreds of times. This restricts its application to systems with only a few hundred to a few thousand atoms.
*Self-consistent field (SCF): A scientific computing method widely used to solve complex many-body problems that must be described by a number of interconnected simultaneous differential equations.
Professor Yong-Hoon Kim’s research team questioned whether recent advancements in AI techniques could be used to bypass the SCF process. As a result, they developed the DeepSCF model, which accelerates calculations by learning chemical bonding information distributed in a 3D space using neural network algorithms from the field of computer vision.
< Figure 2. The deepSCF methodology developed in this study provides a way to rapidly accelerate DFT calculations by avoiding the self-consistent field process (orange box) that had to be performed repeatedly in traditional quantum mechanical electronic structure calculations through artificial neural network techniques (green box). The self-consistent field process is a process of predicting the 3D electron density, constructing the corresponding potential, and then solving the quantum mechanical Cohn-Sham equations, repeating tens to hundreds of times. The core idea of the deepSCF methodology is that the residual electron density (δρ), which is the difference between the electron density (ρ) and the sum of the electron densities of the constituent atoms (ρ0), corresponds to chemical bonding information, so the self-consistent field process is replaced with a 3D convolutional neural network model. >
The research team focused on the fact that, according to density functional theory, electron density contains all quantum mechanical information of electrons, and that the residual electron density — the difference between the total electron density and the sum of the electron densities of the constituent atoms — contains chemical bonding information. They used this as the target for machine learning.
They then adopted a dataset of organic molecules with various chemical bonding characteristics, and applied random rotations and deformations to the atomic structures of these molecules to further enhance the model’s accuracy and generalization capabilities. Ultimately, the research team demonstrated the validity and efficiency of the DeepSCF methodology on large, complex systems.
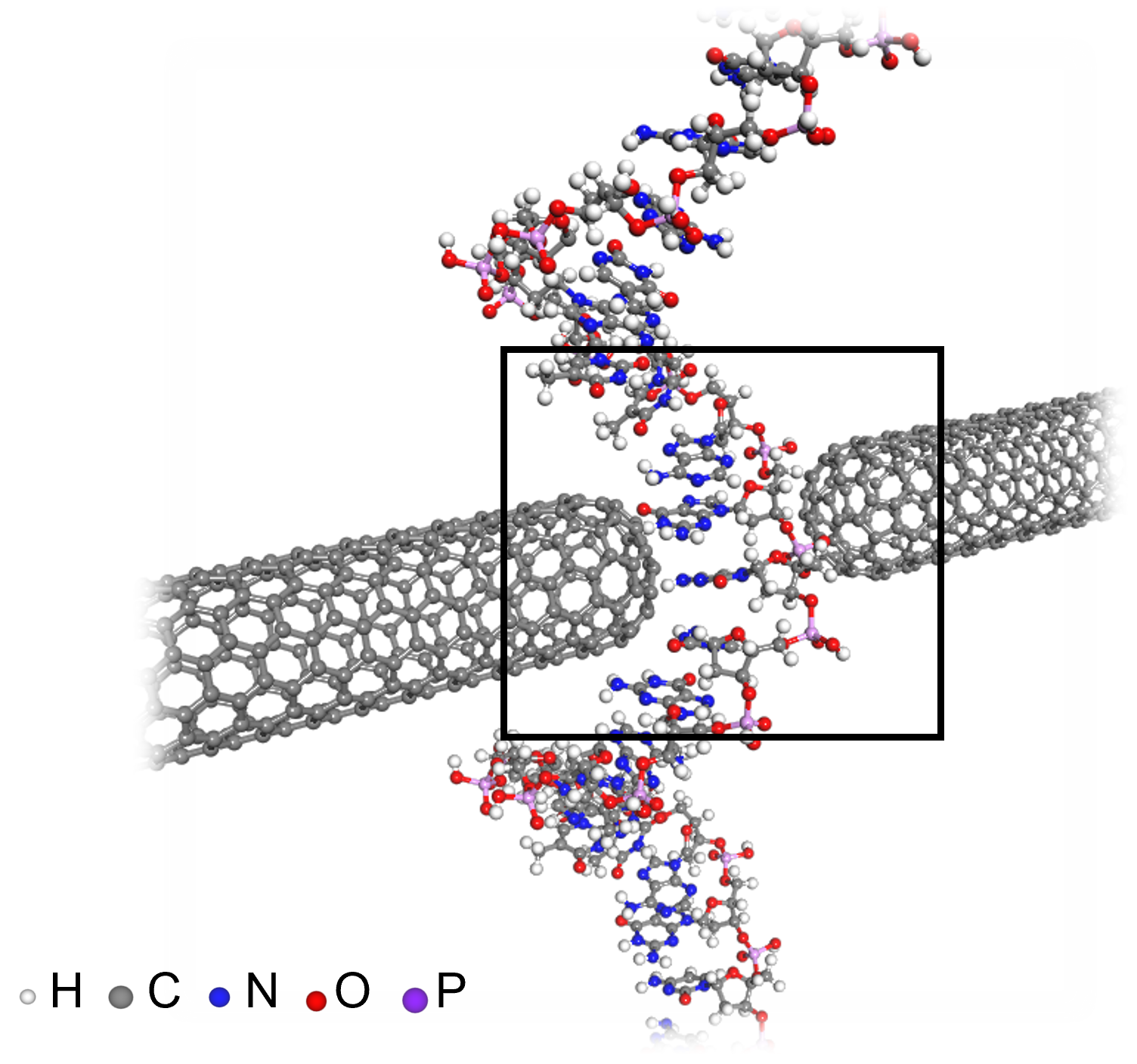
< Figure 3. An example of applying the deepSCF methodology to a carbon nanotube-based DNA sequence analysis device model (top left). In addition to classical mechanical interatomic forces (bottom right), the residual electron density (top right) and quantum mechanical electronic structure properties such as the electronic density of states (DOS) (bottom left) containing information on chemical bonding are rapidly predicted with an accuracy corresponding to the standard DFT calculation results that perform the SCF process. >
Professor Yong-Hoon Kim, who supervised the research, explained that his team had found a way to map quantum mechanical chemical bonding information in a 3D space onto artificial neural networks. He noted, “Since quantum mechanical electron structure calculations underpin materials simulations across all scales, this research establishes a foundational principle for accelerating material calculations using artificial intelligence.”
Ryong-Gyu Lee, a PhD candidate in the School of Electrical Engineering, served as the first author of this research, which was published online on October 24 in Npj Computational Materials, a prestigious journal in the field of material computation. (Paper title: “Convolutional network learning of self-consistent electron density via grid-projected atomic fingerprints”)
This research was conducted with support from the KAIST High-Risk Research Program for Graduate Students and the National Research Foundation of Korea’s Mid-career Researcher Support Program.
2024.10.30 View 7579
KAIST Proposes AI Training Method that will Drastically Shorten Time for Complex Quantum Mechanical Calculations
- Professor Yong-Hoon Kim's team from the School of Electrical Engineering succeeded for the first time in accelerating quantum mechanical electronic structure calculations using a convolutional neural network (CNN) model
- Presenting an AI learning principle of quantum mechanical 3D chemical bonding information, the work is expected to accelerate the computer-assisted designing of next-generation materials and devices
The close relationship between AI and high-performance scientific computing can be seen in the fact that both the 2024 Nobel Prizes in Physics and Chemistry were awarded to scientists for their AI-related research contributions in their respective fields of study. KAIST researchers succeeded in dramatically reducing the computation time for highly sophisticated quantum mechanical computer simulations by predicting atomic-level chemical bonding information distributed in 3D space using a novel AI approach.
KAIST (President Kwang-Hyung Lee) announced on the 30th of October that Professor Yong-Hoon Kim's team from the School of Electrical Engineering developed a 3D computer vision artificial neural network-based computation methodology that bypasses the complex algorithms required for atomic-level quantum mechanical calculations traditionally performed using supercomputers to derive the properties of materials.
< Figure 1. Various methodologies are utilized in the simulation of materials and materials, such as quantum mechanical calculations at the nanometer (nm) level, classical mechanical force fields at the scale of tens to hundreds of nanometers, continuum dynamics calculations at the macroscopic scale, and calculations that mix simulations at different scales. These simulations are already playing a key role in a wide range of basic research and application development fields in combination with informatics techniques. Recently, there have been active efforts to introduce machine learning techniques to radically accelerate simulations, but research on introducing machine learning techniques to quantum mechanical electronic structure calculations, which form the basis of high-scale simulations, is still insufficient. >
The quantum mechanical density functional theory (DFT) calculations using supercomputers have become an essential and standard tool in a wide range of research and development fields, including advanced materials and drug design, as they allow fast and accurate prediction of material properties.
*Density functional theory (DFT): A representative theory of ab initio (first principles) calculations that calculate quantum mechanical properties from the atomic level.
However, practical DFT calculations require generating 3D electron density and solving quantum mechanical equations through a complex, iterative self-consistent field (SCF)* process that must be repeated tens to hundreds of times. This restricts its application to systems with only a few hundred to a few thousand atoms.
*Self-consistent field (SCF): A scientific computing method widely used to solve complex many-body problems that must be described by a number of interconnected simultaneous differential equations.
Professor Yong-Hoon Kim’s research team questioned whether recent advancements in AI techniques could be used to bypass the SCF process. As a result, they developed the DeepSCF model, which accelerates calculations by learning chemical bonding information distributed in a 3D space using neural network algorithms from the field of computer vision.
< Figure 2. The deepSCF methodology developed in this study provides a way to rapidly accelerate DFT calculations by avoiding the self-consistent field process (orange box) that had to be performed repeatedly in traditional quantum mechanical electronic structure calculations through artificial neural network techniques (green box). The self-consistent field process is a process of predicting the 3D electron density, constructing the corresponding potential, and then solving the quantum mechanical Cohn-Sham equations, repeating tens to hundreds of times. The core idea of the deepSCF methodology is that the residual electron density (δρ), which is the difference between the electron density (ρ) and the sum of the electron densities of the constituent atoms (ρ0), corresponds to chemical bonding information, so the self-consistent field process is replaced with a 3D convolutional neural network model. >
The research team focused on the fact that, according to density functional theory, electron density contains all quantum mechanical information of electrons, and that the residual electron density — the difference between the total electron density and the sum of the electron densities of the constituent atoms — contains chemical bonding information. They used this as the target for machine learning.
They then adopted a dataset of organic molecules with various chemical bonding characteristics, and applied random rotations and deformations to the atomic structures of these molecules to further enhance the model’s accuracy and generalization capabilities. Ultimately, the research team demonstrated the validity and efficiency of the DeepSCF methodology on large, complex systems.
< Figure 3. An example of applying the deepSCF methodology to a carbon nanotube-based DNA sequence analysis device model (top left). In addition to classical mechanical interatomic forces (bottom right), the residual electron density (top right) and quantum mechanical electronic structure properties such as the electronic density of states (DOS) (bottom left) containing information on chemical bonding are rapidly predicted with an accuracy corresponding to the standard DFT calculation results that perform the SCF process. >
Professor Yong-Hoon Kim, who supervised the research, explained that his team had found a way to map quantum mechanical chemical bonding information in a 3D space onto artificial neural networks. He noted, “Since quantum mechanical electron structure calculations underpin materials simulations across all scales, this research establishes a foundational principle for accelerating material calculations using artificial intelligence.”
Ryong-Gyu Lee, a PhD candidate in the School of Electrical Engineering, served as the first author of this research, which was published online on October 24 in Npj Computational Materials, a prestigious journal in the field of material computation. (Paper title: “Convolutional network learning of self-consistent electron density via grid-projected atomic fingerprints”)
This research was conducted with support from the KAIST High-Risk Research Program for Graduate Students and the National Research Foundation of Korea’s Mid-career Researcher Support Program.
2024.10.30 View 7579 -
 KAIST Employs Image-recognition AI to Determine Battery Composition and Conditions
An international collaborative research team has developed an image recognition technology that can accurately determine the elemental composition and the number of charge and discharge cycles of a battery by examining only its surface morphology using AI learning.
KAIST (President Kwang-Hyung Lee) announced on July 2nd that Professor Seungbum Hong from the Department of Materials Science and Engineering, in collaboration with the Electronics and Telecommunications Research Institute (ETRI) and Drexel University in the United States, has developed a method to predict the major elemental composition and charge-discharge state of NCM cathode materials with 99.6% accuracy using convolutional neural networks (CNN)*.
*Convolutional Neural Network (CNN): A type of multi-layer, feed-forward, artificial neural network used for analyzing visual images.
The research team noted that while scanning electron microscopy (SEM) is used in semiconductor manufacturing to inspect wafer defects, it is rarely used in battery inspections. SEM is used for batteries to analyze the size of particles only at research sites, and reliability is predicted from the broken particles and the shape of the breakage in the case of deteriorated battery materials.
The research team decided that it would be groundbreaking if an automated SEM can be used in the process of battery production, just like in the semiconductor manufacturing, to inspect the surface of the cathode material to determine whether it was synthesized according to the desired composition and that the lifespan would be reliable, thereby reducing the defect rate.
< Figure 1. Example images of true cases and their grad-CAM overlays from the best trained network. >
The researchers trained a CNN-based AI applicable to autonomous vehicles to learn the surface images of battery materials, enabling it to predict the major elemental composition and charge-discharge cycle states of the cathode materials. They found that while the method could accurately predict the composition of materials with additives, it had lower accuracy for predicting charge-discharge states. The team plans to further train the AI with various battery material morphologies produced through different processes and ultimately use it for inspecting the compositional uniformity and predicting the lifespan of next-generation batteries.
Professor Joshua C. Agar, one of the collaborating researchers of the project from the Department of Mechanical Engineering and Mechanics of Drexel University, said, "In the future, artificial intelligence is expected to be applied not only to battery materials but also to various dynamic processes in functional materials synthesis, clean energy generation in fusion, and understanding foundations of particles and the universe."
Professor Seungbum Hong from KAIST, who led the research, stated, "This research is significant as it is the first in the world to develop an AI-based methodology that can quickly and accurately predict the major elemental composition and the state of the battery from the structural data of micron-scale SEM images. The methodology developed in this study for identifying the composition and state of battery materials based on microscopic images is expected to play a crucial role in improving the performance and quality of battery materials in the future."
< Figure 2. Accuracies of CNN Model predictions on SEM images of NCM cathode materials with additives under various conditions. >
This research was conducted by KAIST’s Materials Science and Engineering Department graduates Dr. Jimin Oh and Dr. Jiwon Yeom, the co-first authors, in collaboration with Professor Josh Agar and Dr. Kwang Man Kim from ETRI. It was supported by the National Research Foundation of Korea, the KAIST Global Singularity project, and international collaboration with the US research team. The results were published in the international journal npj Computational Materials on May 4. (Paper Title: “Composition and state prediction of lithium-ion cathode via convolutional neural network trained on scanning electron microscopy images”)
2024.07.02 View 8361
KAIST Employs Image-recognition AI to Determine Battery Composition and Conditions
An international collaborative research team has developed an image recognition technology that can accurately determine the elemental composition and the number of charge and discharge cycles of a battery by examining only its surface morphology using AI learning.
KAIST (President Kwang-Hyung Lee) announced on July 2nd that Professor Seungbum Hong from the Department of Materials Science and Engineering, in collaboration with the Electronics and Telecommunications Research Institute (ETRI) and Drexel University in the United States, has developed a method to predict the major elemental composition and charge-discharge state of NCM cathode materials with 99.6% accuracy using convolutional neural networks (CNN)*.
*Convolutional Neural Network (CNN): A type of multi-layer, feed-forward, artificial neural network used for analyzing visual images.
The research team noted that while scanning electron microscopy (SEM) is used in semiconductor manufacturing to inspect wafer defects, it is rarely used in battery inspections. SEM is used for batteries to analyze the size of particles only at research sites, and reliability is predicted from the broken particles and the shape of the breakage in the case of deteriorated battery materials.
The research team decided that it would be groundbreaking if an automated SEM can be used in the process of battery production, just like in the semiconductor manufacturing, to inspect the surface of the cathode material to determine whether it was synthesized according to the desired composition and that the lifespan would be reliable, thereby reducing the defect rate.
< Figure 1. Example images of true cases and their grad-CAM overlays from the best trained network. >
The researchers trained a CNN-based AI applicable to autonomous vehicles to learn the surface images of battery materials, enabling it to predict the major elemental composition and charge-discharge cycle states of the cathode materials. They found that while the method could accurately predict the composition of materials with additives, it had lower accuracy for predicting charge-discharge states. The team plans to further train the AI with various battery material morphologies produced through different processes and ultimately use it for inspecting the compositional uniformity and predicting the lifespan of next-generation batteries.
Professor Joshua C. Agar, one of the collaborating researchers of the project from the Department of Mechanical Engineering and Mechanics of Drexel University, said, "In the future, artificial intelligence is expected to be applied not only to battery materials but also to various dynamic processes in functional materials synthesis, clean energy generation in fusion, and understanding foundations of particles and the universe."
Professor Seungbum Hong from KAIST, who led the research, stated, "This research is significant as it is the first in the world to develop an AI-based methodology that can quickly and accurately predict the major elemental composition and the state of the battery from the structural data of micron-scale SEM images. The methodology developed in this study for identifying the composition and state of battery materials based on microscopic images is expected to play a crucial role in improving the performance and quality of battery materials in the future."
< Figure 2. Accuracies of CNN Model predictions on SEM images of NCM cathode materials with additives under various conditions. >
This research was conducted by KAIST’s Materials Science and Engineering Department graduates Dr. Jimin Oh and Dr. Jiwon Yeom, the co-first authors, in collaboration with Professor Josh Agar and Dr. Kwang Man Kim from ETRI. It was supported by the National Research Foundation of Korea, the KAIST Global Singularity project, and international collaboration with the US research team. The results were published in the international journal npj Computational Materials on May 4. (Paper Title: “Composition and state prediction of lithium-ion cathode via convolutional neural network trained on scanning electron microscopy images”)
2024.07.02 View 8361 -
 KAIST Research Team Breaks Down Musical Instincts with AI
Music, often referred to as the universal language, is known to be a common component in all cultures. Then, could ‘musical instinct’ be something that is shared to some degree despite the extensive environmental differences amongst cultures?
On January 16, a KAIST research team led by Professor Hawoong Jung from the Department of Physics announced to have identified the principle by which musical instincts emerge from the human brain without special learning using an artificial neural network model.
Previously, many researchers have attempted to identify the similarities and differences between the music that exist in various different cultures, and tried to understand the origin of the universality. A paper published in Science in 2019 had revealed that music is produced in all ethnographically distinct cultures, and that similar forms of beats and tunes are used. Neuroscientist have also previously found out that a specific part of the human brain, namely the auditory cortex, is responsible for processing musical information.
Professor Jung’s team used an artificial neural network model to show that cognitive functions for music forms spontaneously as a result of processing auditory information received from nature, without being taught music. The research team utilized AudioSet, a large-scale collection of sound data provided by Google, and taught the artificial neural network to learn the various sounds. Interestingly, the research team discovered that certain neurons within the network model would respond selectively to music. In other words, they observed the spontaneous generation of neurons that reacted minimally to various other sounds like those of animals, nature, or machines, but showed high levels of response to various forms of music including both instrumental and vocal.
The neurons in the artificial neural network model showed similar reactive behaviours to those in the auditory cortex of a real brain. For example, artificial neurons responded less to the sound of music that was cropped into short intervals and were rearranged. This indicates that the spontaneously-generated music-selective neurons encode the temporal structure of music. This property was not limited to a specific genre of music, but emerged across 25 different genres including classic, pop, rock, jazz, and electronic.
< Figure 1. Illustration of the musicality of the brain and artificial neural network (created with DALL·E3 AI based on the paper content) >
Furthermore, suppressing the activity of the music-selective neurons was found to greatly impede the cognitive accuracy for other natural sounds. That is to say, the neural function that processes musical information helps process other sounds, and that ‘musical ability’ may be an instinct formed as a result of an evolutionary adaptation acquired to better process sounds from nature.
Professor Hawoong Jung, who advised the research, said, “The results of our study imply that evolutionary pressure has contributed to forming the universal basis for processing musical information in various cultures.” As for the significance of the research, he explained, “We look forward for this artificially built model with human-like musicality to become an original model for various applications including AI music generation, musical therapy, and for research in musical cognition.” He also commented on its limitations, adding, “This research however does not take into consideration the developmental process that follows the learning of music, and it must be noted that this is a study on the foundation of processing musical information in early development.”
< Figure 2. The artificial neural network that learned to recognize non-musical natural sounds in the cyber space distinguishes between music and non-music. >
This research, conducted by first author Dr. Gwangsu Kim of the KAIST Department of Physics (current affiliation: MIT Department of Brain and Cognitive Sciences) and Dr. Dong-Kyum Kim (current affiliation: IBS) was published in Nature Communications under the title, “Spontaneous emergence of rudimentary music detectors in deep neural networks”.
This research was supported by the National Research Foundation of Korea.
2024.01.23 View 9134
KAIST Research Team Breaks Down Musical Instincts with AI
Music, often referred to as the universal language, is known to be a common component in all cultures. Then, could ‘musical instinct’ be something that is shared to some degree despite the extensive environmental differences amongst cultures?
On January 16, a KAIST research team led by Professor Hawoong Jung from the Department of Physics announced to have identified the principle by which musical instincts emerge from the human brain without special learning using an artificial neural network model.
Previously, many researchers have attempted to identify the similarities and differences between the music that exist in various different cultures, and tried to understand the origin of the universality. A paper published in Science in 2019 had revealed that music is produced in all ethnographically distinct cultures, and that similar forms of beats and tunes are used. Neuroscientist have also previously found out that a specific part of the human brain, namely the auditory cortex, is responsible for processing musical information.
Professor Jung’s team used an artificial neural network model to show that cognitive functions for music forms spontaneously as a result of processing auditory information received from nature, without being taught music. The research team utilized AudioSet, a large-scale collection of sound data provided by Google, and taught the artificial neural network to learn the various sounds. Interestingly, the research team discovered that certain neurons within the network model would respond selectively to music. In other words, they observed the spontaneous generation of neurons that reacted minimally to various other sounds like those of animals, nature, or machines, but showed high levels of response to various forms of music including both instrumental and vocal.
The neurons in the artificial neural network model showed similar reactive behaviours to those in the auditory cortex of a real brain. For example, artificial neurons responded less to the sound of music that was cropped into short intervals and were rearranged. This indicates that the spontaneously-generated music-selective neurons encode the temporal structure of music. This property was not limited to a specific genre of music, but emerged across 25 different genres including classic, pop, rock, jazz, and electronic.
< Figure 1. Illustration of the musicality of the brain and artificial neural network (created with DALL·E3 AI based on the paper content) >
Furthermore, suppressing the activity of the music-selective neurons was found to greatly impede the cognitive accuracy for other natural sounds. That is to say, the neural function that processes musical information helps process other sounds, and that ‘musical ability’ may be an instinct formed as a result of an evolutionary adaptation acquired to better process sounds from nature.
Professor Hawoong Jung, who advised the research, said, “The results of our study imply that evolutionary pressure has contributed to forming the universal basis for processing musical information in various cultures.” As for the significance of the research, he explained, “We look forward for this artificially built model with human-like musicality to become an original model for various applications including AI music generation, musical therapy, and for research in musical cognition.” He also commented on its limitations, adding, “This research however does not take into consideration the developmental process that follows the learning of music, and it must be noted that this is a study on the foundation of processing musical information in early development.”
< Figure 2. The artificial neural network that learned to recognize non-musical natural sounds in the cyber space distinguishes between music and non-music. >
This research, conducted by first author Dr. Gwangsu Kim of the KAIST Department of Physics (current affiliation: MIT Department of Brain and Cognitive Sciences) and Dr. Dong-Kyum Kim (current affiliation: IBS) was published in Nature Communications under the title, “Spontaneous emergence of rudimentary music detectors in deep neural networks”.
This research was supported by the National Research Foundation of Korea.
2024.01.23 View 9134 -
 Center for Global Strategies and Planning Hosts Successful Virtual KAIST U.S. Alumni Connection Event
< Screen capture of the KAIST U.S. Alumni meeting held online on December 8 >
On December 8th, the Center for Global Strategies and Planning at KAIST, led by Vice President Man-Sung Yim of the International Office, conducted a virtual event to bring together KAIST alumni in the United States. The purpose of this event was to showcase KAIST's current initiatives in the U.S., facilitate information exchanges among U.S. alumni, and foster networking opportunities. Over 130 KAIST alumni based in the U.S. registered and attended the event.
The event began with a warm welcome from President Kwang-Hyung Lee, followed by a presentation from Vice President Man-Sung Yim on the current status and vision of KAIST's U.S. collaboration project as well as that of KAIST U.S. Foundation, Inc. Additionally, a distinguished KAIST alumnus, Seok-Hyun Yun, a professor from Harvard Medical School, delivered a keynote speech that highlighted the development of collaborative projects between KAIST and the United States. Alumni Hyun Gook Yoon, a manager at Ford Motor Company, and Eunkwang Joo, CEO of Wasder, also presented recent technological trends in the fields of batteries and blockchain, respectively.
President Kwang-Hyung Lee said, "This event serves as a crucial opportunity to enhance exchanges between KAIST and the U.S., playing a pivotal role in expanding KAIST's global presence." The event also featured small group discussions and networking sessions focusing on revitalizing collaborative efforts between KAIST and the United States.
After the small group discussions, a KAIST alumna and the current president of the Boston KAIST Alumni Association, Jiyoung Lee, shared her belief that the event will provide a meaningful opportunity for KAIST alumni from across the U.S. to come together and build a strong alumni community. Vice President Man-Sung Yim said, "Because collaboration with KAIST alumni in the U.S. is essential for the development of KAIST and innovative science and technology at the global level, we are committed to sustainably organizing meaningful events."
This virtual event for KAIST U.S. alumni has set a new milestone for global networking, marking the beginning of future collaborations and development.
2023.12.08 View 7954
Center for Global Strategies and Planning Hosts Successful Virtual KAIST U.S. Alumni Connection Event
< Screen capture of the KAIST U.S. Alumni meeting held online on December 8 >
On December 8th, the Center for Global Strategies and Planning at KAIST, led by Vice President Man-Sung Yim of the International Office, conducted a virtual event to bring together KAIST alumni in the United States. The purpose of this event was to showcase KAIST's current initiatives in the U.S., facilitate information exchanges among U.S. alumni, and foster networking opportunities. Over 130 KAIST alumni based in the U.S. registered and attended the event.
The event began with a warm welcome from President Kwang-Hyung Lee, followed by a presentation from Vice President Man-Sung Yim on the current status and vision of KAIST's U.S. collaboration project as well as that of KAIST U.S. Foundation, Inc. Additionally, a distinguished KAIST alumnus, Seok-Hyun Yun, a professor from Harvard Medical School, delivered a keynote speech that highlighted the development of collaborative projects between KAIST and the United States. Alumni Hyun Gook Yoon, a manager at Ford Motor Company, and Eunkwang Joo, CEO of Wasder, also presented recent technological trends in the fields of batteries and blockchain, respectively.
President Kwang-Hyung Lee said, "This event serves as a crucial opportunity to enhance exchanges between KAIST and the U.S., playing a pivotal role in expanding KAIST's global presence." The event also featured small group discussions and networking sessions focusing on revitalizing collaborative efforts between KAIST and the United States.
After the small group discussions, a KAIST alumna and the current president of the Boston KAIST Alumni Association, Jiyoung Lee, shared her belief that the event will provide a meaningful opportunity for KAIST alumni from across the U.S. to come together and build a strong alumni community. Vice President Man-Sung Yim said, "Because collaboration with KAIST alumni in the U.S. is essential for the development of KAIST and innovative science and technology at the global level, we are committed to sustainably organizing meaningful events."
This virtual event for KAIST U.S. alumni has set a new milestone for global networking, marking the beginning of future collaborations and development.
2023.12.08 View 7954 -
 A KAIST research team develops a high-performance modular SSD system semiconductor
In recent years, there has been a rise in demand for large amounts of data to train AI models and, thus, data size has become increasingly important over time. Accordingly, solid state drives (SSDs, storage devices that use a semiconductor memory unit), which are core storage devices for data centers and cloud services, have also seen an increase in demand. However, the internal components of higher performing SSDs have become more tightly coupled, and this tightly-coupled structure limits SSD from maximized performance.
On June 15, a KAIST research team led by Professor Dongjun Kim (John Kim) from the School of Electrical Engineering (EE) announced the development of the first SSD system semiconductor structure that can increase the reading/writing performance of next generation SSDs and extend their lifespan through high-performance modular SSD systems.
Professor Kim’s team identified the limitations of the tightly-coupled structures in existing SSD designs and proposed a de-coupled structure that can maximize SSD performance by configuring an internal on-chip network specialized for flash memory. This technique utilizes on-chip network technology, which can freely send packet-based data within the chip and is often used to design non-memory system semiconductors like CPUs and GPUs. Through this, the team developed a ‘modular SSD’, which shows reduced interdependence between front-end and back-end designs, and allows their independent design and assembly.
*on-chip network: a packet-based connection structure for the internal components of system semiconductors like CPUs/GPUs. On-chip networks are one of the most critical design components for high-performing system semiconductors, and their importance grows with the size of the semiconductor chip.
Professor Kim’s team refers to the components nearer to the CPU as the front-end and the parts closer to the flash memory as back-end. They newly constructed an on-chip network specific to flash memory in order to allow data transmission between the back-end’s flash controller, proposing a de-coupled structure that can minimize performance drop.
The SSD can accelerate some functions of the flash translation layer, a critical element to drive the SSD, in order to allow flash memory to actively overcome its limitations. Another advantage of the de-coupled, modular structure is that the flash translation layer is not limited to the characteristics of specific flash memories. Instead, their front-end and back-end designs can be carried out independently. Through this, the team could produce 21-times faster response times compared to existing systems and extend SSD lifespan by 23% by also applying the DDS defect detection technique.
< Figure 1. Schematic diagram of the structure of a high-performance modular SSD system developed by Professor Dong-Jun Kim's team >
This research, conducted by first author and Ph.D. candidate Jiho Kim from the KAIST School of EE and co-author Professor Myoungsoo Jung, was presented on the 19th of June at the 50th IEEE/ACM International Symposium on Computer Architecture, the most prestigious academic conference in the field of computer architecture, held in Orlando, Florida. (Paper Title: Decoupled SSD: Rethinking SSD Architecture through Network-based Flash Controllers)
< Figure 2. Conceptual diagram of hardware acceleration through high-performance modular SSD system >
Professor Dongjun Kim, who led the research, said, “This research is significant in that it identified the structural limitations of existing SSDs, and showed that on-chip network technology based on system memory semiconductors like CPUs can drive the hardware to actively carry out the necessary actions. We expect this to contribute greatly to the next-generation high-performance SSD market.” He added, “The de-coupled architecture is a structure that can actively operate to extend devices’ lifespan. In other words, its significance is not limited to the level of performance and can, therefore, be used for various applications.”
KAIST commented that this research is also meaningful in that the results were reaped through a collaborative study between two world-renowned researchers: Professor Myeongsoo Jung, recognized in the field of computer system storage devices, and Professor Dongjun Kim, a leading researcher in computer architecture and interconnection networks.
This research was funded by the National Research Foundation of Korea, Samsung Electronics, the IC Design Education Center, and Next Generation Semiconductor Technology and Development granted by the Institute of Information & Communications Technology, Planning & Evaluation.
2023.06.23 View 9393
A KAIST research team develops a high-performance modular SSD system semiconductor
In recent years, there has been a rise in demand for large amounts of data to train AI models and, thus, data size has become increasingly important over time. Accordingly, solid state drives (SSDs, storage devices that use a semiconductor memory unit), which are core storage devices for data centers and cloud services, have also seen an increase in demand. However, the internal components of higher performing SSDs have become more tightly coupled, and this tightly-coupled structure limits SSD from maximized performance.
On June 15, a KAIST research team led by Professor Dongjun Kim (John Kim) from the School of Electrical Engineering (EE) announced the development of the first SSD system semiconductor structure that can increase the reading/writing performance of next generation SSDs and extend their lifespan through high-performance modular SSD systems.
Professor Kim’s team identified the limitations of the tightly-coupled structures in existing SSD designs and proposed a de-coupled structure that can maximize SSD performance by configuring an internal on-chip network specialized for flash memory. This technique utilizes on-chip network technology, which can freely send packet-based data within the chip and is often used to design non-memory system semiconductors like CPUs and GPUs. Through this, the team developed a ‘modular SSD’, which shows reduced interdependence between front-end and back-end designs, and allows their independent design and assembly.
*on-chip network: a packet-based connection structure for the internal components of system semiconductors like CPUs/GPUs. On-chip networks are one of the most critical design components for high-performing system semiconductors, and their importance grows with the size of the semiconductor chip.
Professor Kim’s team refers to the components nearer to the CPU as the front-end and the parts closer to the flash memory as back-end. They newly constructed an on-chip network specific to flash memory in order to allow data transmission between the back-end’s flash controller, proposing a de-coupled structure that can minimize performance drop.
The SSD can accelerate some functions of the flash translation layer, a critical element to drive the SSD, in order to allow flash memory to actively overcome its limitations. Another advantage of the de-coupled, modular structure is that the flash translation layer is not limited to the characteristics of specific flash memories. Instead, their front-end and back-end designs can be carried out independently. Through this, the team could produce 21-times faster response times compared to existing systems and extend SSD lifespan by 23% by also applying the DDS defect detection technique.
< Figure 1. Schematic diagram of the structure of a high-performance modular SSD system developed by Professor Dong-Jun Kim's team >
This research, conducted by first author and Ph.D. candidate Jiho Kim from the KAIST School of EE and co-author Professor Myoungsoo Jung, was presented on the 19th of June at the 50th IEEE/ACM International Symposium on Computer Architecture, the most prestigious academic conference in the field of computer architecture, held in Orlando, Florida. (Paper Title: Decoupled SSD: Rethinking SSD Architecture through Network-based Flash Controllers)
< Figure 2. Conceptual diagram of hardware acceleration through high-performance modular SSD system >
Professor Dongjun Kim, who led the research, said, “This research is significant in that it identified the structural limitations of existing SSDs, and showed that on-chip network technology based on system memory semiconductors like CPUs can drive the hardware to actively carry out the necessary actions. We expect this to contribute greatly to the next-generation high-performance SSD market.” He added, “The de-coupled architecture is a structure that can actively operate to extend devices’ lifespan. In other words, its significance is not limited to the level of performance and can, therefore, be used for various applications.”
KAIST commented that this research is also meaningful in that the results were reaped through a collaborative study between two world-renowned researchers: Professor Myeongsoo Jung, recognized in the field of computer system storage devices, and Professor Dongjun Kim, a leading researcher in computer architecture and interconnection networks.
This research was funded by the National Research Foundation of Korea, Samsung Electronics, the IC Design Education Center, and Next Generation Semiconductor Technology and Development granted by the Institute of Information & Communications Technology, Planning & Evaluation.
2023.06.23 View 9393 -
 A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 13323
A KAIST Research Team Identifies a Cancer Reversion Mechanism
Despite decades of intensive cancer research by numerous biomedical scientists, cancer still holds its place as the number one cause of death in Korea. The fundamental reason behind the limitations of current cancer treatment methods is the fact that they all aim to completely destroy cancer cells, which eventually allows the cancer cells to acquire immunity. In other words, recurrences and side-effects caused by the destruction of healthy cells are inevitable. To this end, some have suggested anticancer treatment methods based on cancer reversion, which can revert cancer cells back to normal or near-normal cells under certain conditions. However, the practical development of this idea has not yet been attempted.
On June 8, a KAIST research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering reported to have successfully identified the fundamental principle of a process that can revert cancer cells back to normal cells without killing the cells.
Professor Cho’s team focused on the fact that unlike normal cells, which react according to external stimuli, cancer cells tend to ignore such stimuli and only undergo uncontrolled cell division. Through computer simulation analysis, the team discovered that the input-output (I/O) relationships that were distorted by genetic mutations could be reverted back to normal I/O relationships under certain conditions. The team then demonstrated through molecular cell experiments that such I/O relationship recovery also occurred in real cancer cells.
The results of this study, written by Dr. Jae Il Joo and Dr. Hwa-Jeong Park, were published in Wiley’s Advanced Science online on June 2 under the title, "Normalizing input-output relationships of cancer networks for reversion therapy."
< Image 1. Input-output (I/O) relationships in gene regulatory networks >
Professor Kwang-Hyun Cho's research team classified genes into four types by simulation-analyzing the effect of gene mutations on the I/O relationship of gene regulatory networks. (Figure A-J) In addition, by analyzing 18 genes of the cancer-related gene regulatory network, it was confirmed that when mutations occur in more than half of the genes constituting each network, reversibility is possible through appropriate control. (Figure K)
Professor Cho’s team uncovered that the reason the distorted I/O relationships of cancer cells could be reverted back to normal ones was the robustness and redundancy of intracellular gene control networks that developed over the course of evolution. In addition, they found that some genes were more promising as targets for cancer reversion than others, and showed through molecular cell experiments that controlling such genes could revert the distorted I/O relationships of cancer cells back to normal ones.
< Image 2. Simulation results of restoration of bladder cancer gene regulation network and I/O relationship of bladder cancer cells. >
The research team classified the effects of gene mutations on the I/O relationship in the bladder cancer gene regulation network by simulation analysis and classified them into 4 types. (Figure A) Through this, it was found that the distorted input-output relationship between bladder cancer cell lines KU-1919 and HCT-1197 could be restored to normal. (Figure B)
< Image 3. Analysis of survival of bladder cancer patients according to reversible gene mutation and I/O recovery experiment of bladder cancer cells. >
As predicted through network simulation analysis, Professor Kwang-Hyun Cho's research team confirmed through molecular cell experiments that the response to TGF-b was normally restored when AKT and MAP3K1 were inhibited in the bladder cancer cell line KU-1919. (Figure A-G) In addition, it was confirmed that there is a difference in the survival rate of bladder cancer patients depending on the presence or absence of a reversible gene mutation. (Figure H)
The results of this research show that the reversion of real cancer cells does not happen by chance, and that it is possible to systematically explore targets that can induce this phenomenon, thereby creating the potential for the development of innovative anticancer drugs that can control such target genes.
< Image 4. Cancer cell reversibility principle >
The research team analyzed the reversibility, redundancy, and robustness of various networks and found that there was a positive correlation between them. From this, it was found that reversibility was additionally inherent in the process of evolution in which the gene regulatory network acquired redundancy and consistency.
Professor Cho said, “By uncovering the fundamental principles of a new cancer reversion treatment strategy that may overcome the unresolved limitations of existing chemotherapy, we have increased the possibility of developing new and innovative drugs that can improve both the prognosis and quality of life of cancer patients.”
< Image 5. Conceptual diagram of research results >
The research team identified the fundamental control principle of cancer cell reversibility through systems biology research. When the I/O relationship of the intracellular gene regulatory network is distorted by mutation, the distorted I/O relationship can be restored to a normal state by identifying and adjusting the reversible gene target based on the redundancy of the molecular circuit inherent in the complex network.
After Professor Cho’s team first suggested the concept of reversion treatment, they published their results for reverting colorectal cancer in January 2020, and in January 2022 they successfully re-programmed malignant breast cancer cells back into hormone-treatable ones. In January 2023, the team successfully removed the metastasis ability from lung cancer cells and reverted them back to a state that allowed improved drug reactivity. However, these results were case studies of specific types of cancer and did not reveal what common principle allowed cancer reversion across all cancer types, making this the first revelation of the general principle of cancer reversion and its evolutionary origins.
This research was funded by the Ministry of Science and ICT of the Republic of Korea and the National Research Foundation of Korea.
2023.06.20 View 13323 -
 KAIST team develops smart immune system that can pin down on malignant tumors
A joint research team led by Professor Jung Kyoon Choi of the KAIST Department of Bio and Brain Engineering and Professor Jong-Eun Park of the KAIST Graduate School of Medical Science and Engineering (GSMSE) announced the development of the key technologies to treat cancers using smart immune cells designed based on AI and big data analysis. This technology is expected to be a next-generation immunotherapy that allows precision targeting of tumor cells by having the chimeric antigen receptors (CARs) operate through a logical circuit. Professor Hee Jung An of CHA Bundang Medical Center and Professor Hae-Ock Lee of the Catholic University of Korea also participated in this research to contribute joint effort.
Professor Jung Kyoon Choi’s team built a gene expression database from millions of cells, and used this to successfully develop and verify a deep-learning algorithm that could detect the differences in gene expression patterns between tumor cells and normal cells through a logical circuit. CAR immune cells that were fitted with the logic circuits discovered through this methodology could distinguish between tumorous and normal cells as a computer would, and therefore showed potentials to strike only on tumor cells accurately without causing unwanted side effects.
This research, conducted by co-first authors Dr. Joonha Kwon of the KAIST Department of Bio and Brain Engineering and Ph.D. candidate Junho Kang of KAIST GSMSE, was published by Nature Biotechnology on February 16, under the title Single-cell mapping of combinatorial target antigens for CAR switches using logic gates.
An area in cancer research where the most attempts and advances have been made in recent years is immunotherapy. This field of treatment, which utilizes the patient’s own immune system in order to overcome cancer, has several methods including immune checkpoint inhibitors, cancer vaccines and cellular treatments. Immune cells like CAR-T or CAR-NK equipped with chimera antigen receptors, in particular, can recognize cancer antigens and directly destroy cancer cells.
Starting with its success in blood cancer treatment, scientists have been trying to expand the application of CAR cell therapy to treat solid cancer. But there have been difficulties to develop CAR cells with effective killing abilities against solid cancer cells with minimized side effects. Accordingly, in recent years, the development of smarter CAR engineering technologies, i.e., computational logic gates such as AND, OR, and NOT, to effectively target cancer cells has been underway.
At this point in time, the research team built a large-scale database for cancer and normal cells to discover the exact genes that are expressed only from cancer cells at a single-cell level. The team followed this up by developing an AI algorithm that could search for a combination of genes that best distinguishes cancer cells from normal cells. This algorithm, in particular, has been used to find a logic circuit that can specifically target cancer cells through cell-level simulations of all gene combinations. CAR-T cells equipped with logic circuits discovered through this methodology are expected to distinguish cancerous cells from normal cells like computers, thereby minimizing side effects and maximizing the effects of chemotherapy.
Dr. Joonha Kwon, who is the first author of this paper, said, “this research suggests a new method that hasn’t been tried before. What’s particularly noteworthy is the process in which we found the optimal CAR cell circuit through simulations of millions of individual tumors and normal cells.” He added, “This is an innovative technology that can apply AI and computer logic circuits to immune cell engineering. It would contribute greatly to expanding CAR therapy, which is being successfully used for blood cancer, to solid cancers as well.”
This research was funded by the Original Technology Development Project and Research Program for Next Generation Applied Omic of the Korea Research Foundation.
Figure 1. A schematic diagram of manufacturing and administration process of CAR therapy and of cancer cell-specific dual targeting using CAR.
Figure 2. Deep learning (convolutional neural networks, CNNs) algorithm for selection of dual targets based on gene combination (left) and algorithm for calculating expressing cell fractions by gene combination according to logical circuit (right).
2023.03.09 View 13271
KAIST team develops smart immune system that can pin down on malignant tumors
A joint research team led by Professor Jung Kyoon Choi of the KAIST Department of Bio and Brain Engineering and Professor Jong-Eun Park of the KAIST Graduate School of Medical Science and Engineering (GSMSE) announced the development of the key technologies to treat cancers using smart immune cells designed based on AI and big data analysis. This technology is expected to be a next-generation immunotherapy that allows precision targeting of tumor cells by having the chimeric antigen receptors (CARs) operate through a logical circuit. Professor Hee Jung An of CHA Bundang Medical Center and Professor Hae-Ock Lee of the Catholic University of Korea also participated in this research to contribute joint effort.
Professor Jung Kyoon Choi’s team built a gene expression database from millions of cells, and used this to successfully develop and verify a deep-learning algorithm that could detect the differences in gene expression patterns between tumor cells and normal cells through a logical circuit. CAR immune cells that were fitted with the logic circuits discovered through this methodology could distinguish between tumorous and normal cells as a computer would, and therefore showed potentials to strike only on tumor cells accurately without causing unwanted side effects.
This research, conducted by co-first authors Dr. Joonha Kwon of the KAIST Department of Bio and Brain Engineering and Ph.D. candidate Junho Kang of KAIST GSMSE, was published by Nature Biotechnology on February 16, under the title Single-cell mapping of combinatorial target antigens for CAR switches using logic gates.
An area in cancer research where the most attempts and advances have been made in recent years is immunotherapy. This field of treatment, which utilizes the patient’s own immune system in order to overcome cancer, has several methods including immune checkpoint inhibitors, cancer vaccines and cellular treatments. Immune cells like CAR-T or CAR-NK equipped with chimera antigen receptors, in particular, can recognize cancer antigens and directly destroy cancer cells.
Starting with its success in blood cancer treatment, scientists have been trying to expand the application of CAR cell therapy to treat solid cancer. But there have been difficulties to develop CAR cells with effective killing abilities against solid cancer cells with minimized side effects. Accordingly, in recent years, the development of smarter CAR engineering technologies, i.e., computational logic gates such as AND, OR, and NOT, to effectively target cancer cells has been underway.
At this point in time, the research team built a large-scale database for cancer and normal cells to discover the exact genes that are expressed only from cancer cells at a single-cell level. The team followed this up by developing an AI algorithm that could search for a combination of genes that best distinguishes cancer cells from normal cells. This algorithm, in particular, has been used to find a logic circuit that can specifically target cancer cells through cell-level simulations of all gene combinations. CAR-T cells equipped with logic circuits discovered through this methodology are expected to distinguish cancerous cells from normal cells like computers, thereby minimizing side effects and maximizing the effects of chemotherapy.
Dr. Joonha Kwon, who is the first author of this paper, said, “this research suggests a new method that hasn’t been tried before. What’s particularly noteworthy is the process in which we found the optimal CAR cell circuit through simulations of millions of individual tumors and normal cells.” He added, “This is an innovative technology that can apply AI and computer logic circuits to immune cell engineering. It would contribute greatly to expanding CAR therapy, which is being successfully used for blood cancer, to solid cancers as well.”
This research was funded by the Original Technology Development Project and Research Program for Next Generation Applied Omic of the Korea Research Foundation.
Figure 1. A schematic diagram of manufacturing and administration process of CAR therapy and of cancer cell-specific dual targeting using CAR.
Figure 2. Deep learning (convolutional neural networks, CNNs) algorithm for selection of dual targets based on gene combination (left) and algorithm for calculating expressing cell fractions by gene combination according to logical circuit (right).
2023.03.09 View 13271 -
 KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 20827
KAIST presents a fundamental technology to remove metastatic traits from lung cancer cells
KAIST (President Kwang Hyung Lee) announced on January 30th that a research team led by Professor Kwang-Hyun Cho from the Department of Bio and Brain Engineering succeeded in using systems biology research to change the properties of carcinogenic cells in the lungs and eliminate both drug resistance and their ability to proliferate out to other areas of the body.
As the incidences of cancer increase within aging populations, cancer has become the most lethal disease threatening healthy life. Fatality rates are especially high when early detection does not happen in time and metastasis has occurred in various organs. In order to resolve this problem, a series of attempts were made to remove or lower the ability of cancer cells to spread, but they resulted in cancer cells in the intermediate state becoming more unstable and even more malignant, which created serious treatment challenges.
Professor Kwang-Hyun Cho's research team simulated various cancer cell states in the Epithelial-to-Mesenchymal Transition (EMT) of lung cancer cells, between epithelial cells without metastatic ability and mesenchymal cells with metastatic ability. A mathematical model of molecular network was established, and key regulators that could reverse the state of invasive and drug resistant mesenchymal cells back to the epithelial state were discovered through computer simulation analysis and molecular cell experiments. In particular, this process succeeded in properly reverting the mesenchymal lung cancer cells to a state where they were sensitive to chemotherapy treatment while avoiding the unstable EMT hybrid cell state in the middle process, which had remained a difficult problem.
The results of this research, in which KAIST Ph.D. student Namhee Kim, Dr. Chae Young Hwang, Researcher Taeyoung Kim, and Ph.D. student Hyunjin Kim participated, were published as an online paper in the international journal “Cancer Research” published by the American Association for Cancer Research (AACR) on January 30th. (Paper title: A cell fate reprogramming strategy reverses epithelial-to-mesenchymal transition of lung cancer cells while avoiding hybrid states)
Cells in an EMT hybrid state, which are caused by incomplete transitions during the EMT process in cancer cells, have the characteristics of both epithelial cells and mesenchymal cells, and are known to have high drug resistance and metastatic potential by acquiring high stem cell capacity. In particular, EMT is further enhanced through factors such as transforming growth factor-beta (TGF-β) secreted from the tumor microenvironment (TME) and, as a result, various cell states with high plasticity appear. Due to the complexity of EMT, it has been very difficult to completely reverse the transitional process of the mesenchymal cancer cells to an epithelial cell state in which metastatic ability and drug resistance are eliminated while avoiding the EMT hybrid cell state with high metastatic ability and drug resistance.
Professor Kwang-Hyun Cho's research team established a mathematical model of the gene regulation network that governs the complex process of EMT, and then applied large-scale computer simulation analysis and complex system network control technology to identify and verify 'p53', 'SMAD4', and 'ERK1' and 'ERK 2' (collectively ERKs) through molecular cell experiments as the three key molecular targets that can transform lung cancer cells in the mesenchymal cell state, reversed back to an epithelial cell state that no longer demonstrates the ability to metastasize, while avoiding the EMT hybrid cell state.
In particular, by analyzing the molecular regulatory mechanism of the complex EMT process at the system level, the key pathways were identified that were linked to the positive feedback that plays an important role in completely returning cancer cells to an epithelial cell state in which metastatic ability and drug resistance are removed.
This discovery is significant in that it proved that mesenchymal cells can be reverted to the state of epithelial cells under conditions where TGF-β stimulation are present, like they are in the actual environment where cancer tissue forms in the human body.
Abnormal EMT in cancer cells leads to various malignant traits such as the migration and invasion of cancer cells, changes in responsiveness to chemotherapy treatment, enhanced stem cell function, and the dissemination of cancer. In particular, the acquisition of the metastatic ability of cancer cells is a key determinant factor for the prognosis of cancer patients. The EMT reversal technology in lung cancer cells developed in this research is a new anti-cancer treatment strategy that reprograms cancer cells to eliminate their high plasticity and metastatic potential and increase their responsiveness to chemotherapy.
Professor Kwang-Hyun Cho said, "By succeeding in reversing the state of lung cancer cells that acquired high metastatic traits and resistance to drugs and reverting them to a treatable epithelial cell state with renewed sensitivity to chemotherapy, the research findings propose a new strategy for treatments that can improve the prognosis of cancer patients.”
Professor Kwang-Hyun Cho's research team was the first to present the principle of reversal treatment to revert cancer cells to normal cells, following through with the announcement of the results of their study that reverted colon cancer cells to normal colon cells in January of 2020, and also presenting successful re-programming research where the most malignant basal type breast cancer cells turned into less-malignant luminal type breast cancer cells that were treatable with hormonal therapies in January of 2022. This latest research result is the third in the development of reversal technology where lung cancer cells that had acquired metastatic traits returned to a state in which their metastatic ability was removed and drug sensitivity was enhanced.
This research was carried out with support from the Ministry of Science and ICT and the National Research Foundation of Korea's Basic Research in Science & Engineering Program for Mid-Career Researchers.
< Figure 1. Construction of the mathematical model of the regulatory network to represent the EMT phenotype based on the interaction between various molecules related to EMT.
(A) Professor Kwang-Hyun Cho's research team investigated numerous literatures and databases related to complex EMT, and based on comparative analysis of cell line data showing epithelial and mesenchymal cell conditions, they extracted key signaling pathways related to EMT and built a mathematical model of regulatory network (B) By comparing the results of computer simulation analysis and the molecular cell experiments, it was verified how well the constructed mathematical model simulated the actual cellular phenomena. >
< Figure 2. Understanding of various EMT phenotypes through large-scale computer simulation analysis and complex system network control technology.
(A) Through computer simulation analysis and experiments, Professor Kwang-Hyun Cho's research team found that complete control of EMT is impossible with single-molecule control alone. In particular, through comparison of the relative stability of attractors, it was revealed that the cell state exhibiting EMT hybrid characteristics has unstable properties. (B), (C) Based on these results, Prof. Cho’s team identified two feedbacks (positive feedback consisting of Snail-miR-34 and ZEB1-miR-200) that play an important role in avoiding the EMT hybrid state that appeared in the TGF-β-ON state. It was found through computer simulation analysis that the two feedbacks restore relatively high stability when the excavated p53 and SMAD4 are regulated. In addition, molecular cell experiments demonstrated that the expression levels of E-cad and ZEB1, which are representative phenotypic markers of EMT, changed similarly to the expression profile in the epithelial cell state, despite the TGF-β-ON state. >
< Figure 3. Complex molecular network analysis and discovery of reprogramming molecular targets for intact elimination of EMT hybrid features.
(A) Controlling the expression of p53 and SMAD4 in lung cancer cell lines was expected to overcome drug resistance, but contrary to expectations, chemotherapy responsiveness was not restored. (B) Professor Kwang-Hyun Cho's research team additionally analyzed computer simulations, genome data, and experimental results and found that high expression levels of TWIST1 and EPCAM were related to drug resistance. (C) Prof. Cho’s team identified three key molecular targets: p53, SMAD4 and ERK1 & ERK2. (D), (E) Furthermore, they identified a key pathway that plays an important role in completely reversing into epithelial cells while avoiding EMT hybrid characteristics, and confirmed through network analysis and attractor analysis that high stability of the key pathway was restored when the proposed molecular target was controlled. >
< Figure 4. Verification through experiments with lung cancer cell lines.
When p53 was activated and SMAD4 and ERK1/2 were inhibited in lung cancer cell lines, (A), (B) E-cad protein expression increased and ZEB1 protein expression decreased, and (C) mesenchymal cell status including TWIST1 and EPCAM and gene expression of markers related to stem cell potential characteristics were completely inhibited. In addition, (D) it was confirmed that resistance to chemotherapy treatment was also overcome as the cell state was reversed by the regulated target. >
< Figure 5. A schematic representation of the research results.
Prof. Cho’s research team identified key molecular regulatory pathways to avoid high plasticity formed by abnormal EMT of cancer cells and reverse it to an epithelial cell state through systems biology research. From this analysis, a reprogramming molecular target that can reverse the state of mesenchymal cells with acquired invasiveness and drug resistance to the state of epithelial cells with restored drug responsiveness was discovered.
For lung cancer cells, when a drug that enhances the expression of p53, one of the molecular targets discovered, and inhibits the expression of SMAD4 and ERK1 & ERK2 is administered, the molecular network of genes in the state of mesenchymal cells is modified, eventually eliminating metastatic ability and it is reprogrammed to turn into epithelial cells without the resistance to chemotherapy treatments. >
2023.01.30 View 20827