Nature
-
 Making Graphene Using Laser-induced Phase Separation
IBS & KAIST researchers clarify how laser annealing technology can lead to the production of ultrathin nanomaterials
All our smart phones have shiny flat AMOLED (active-matrix organic light-emitting diode) displays. Behind each single pixel of these displays hides at least two silicon transistors which are mass-manufactured using laser annealing technology. While the traditional methods to make the transistors use temperature above 1,000°C, the laser technique reaches the same results at low temperatures even on plastic substrates (melting temperature below 300°C). Interestingly, a similar procedure can be used to generate crystals of graphene. Graphene is a strong and thin nano-material made of carbon, its electric and heat-conductive properties have attracted the attention of scientists worldwide.
Professor Keon Jae Lee of the Materials Science and Engineering Department at KAIST and his research group at the Center for Multidimensional Carbon Materials within the Institute for Basic Science (IBS), as well as Professor Sung-Yool Choi of the Electrical Engineering School at KAIST and his research team discovered graphene synthesis mechanism using laser-induced solid-state phase separation of single-crystal silicon carbide (SiC). This study, available in Nature Communications, clarifies how this laser technology can separate a complex compound (SiC) into its ultrathin elements of carbon and silicon.
Although several fundamental studies presented the effect of excimer lasers in transforming elemental materials like silicon, the laser interaction with more complex compounds like SiC has rarely been studied due to the complexity of compound phase transition and ultra-short processing time.
With high resolution microscope images and molecular dynamic simulations, scientists found that a single-pulse irradiation of xenon chloride excimer laser of 30 nanoseconds melts SiC, leading to the separation of a liquid SiC layer, a disordered carbon layer with graphitic domains (about 2.5 nm thick) on top surface and a polycrystalline silicon layer (about 5 nm) below carbon layer. Giving additional pulses causes the sublimation of the separated silicon, while the disordered carbon layer is transformed into a multilayer graphene.
"This research shows that the laser material interaction technology can be a powerful tool for the next generation of two dimensional nanomaterials," said Professor Lee.
Professor Choi added: "Using laser-induced phase separation of complex compounds, new types of two dimensional materials can be synthesized in the future."
High-resolution transmission electron microscopy shows that after just one laser pulse of 30 nanoseconds, the silicon carbide (SiC) substrate is melted and separates into a carbon and a silicon layer. More pulses cause the carbon layer to organize into graphene and the silicon to leave as gas.
Molecular dynamics simulates the graphene formation mechanism. The carbon layer on the top forms because the laser-induced liquid SiC (SiC (l)) is unstable.
(Press Release by Courtesy of the Institute for Basic Science (IBS))
2016.12.01 View 13284
Making Graphene Using Laser-induced Phase Separation
IBS & KAIST researchers clarify how laser annealing technology can lead to the production of ultrathin nanomaterials
All our smart phones have shiny flat AMOLED (active-matrix organic light-emitting diode) displays. Behind each single pixel of these displays hides at least two silicon transistors which are mass-manufactured using laser annealing technology. While the traditional methods to make the transistors use temperature above 1,000°C, the laser technique reaches the same results at low temperatures even on plastic substrates (melting temperature below 300°C). Interestingly, a similar procedure can be used to generate crystals of graphene. Graphene is a strong and thin nano-material made of carbon, its electric and heat-conductive properties have attracted the attention of scientists worldwide.
Professor Keon Jae Lee of the Materials Science and Engineering Department at KAIST and his research group at the Center for Multidimensional Carbon Materials within the Institute for Basic Science (IBS), as well as Professor Sung-Yool Choi of the Electrical Engineering School at KAIST and his research team discovered graphene synthesis mechanism using laser-induced solid-state phase separation of single-crystal silicon carbide (SiC). This study, available in Nature Communications, clarifies how this laser technology can separate a complex compound (SiC) into its ultrathin elements of carbon and silicon.
Although several fundamental studies presented the effect of excimer lasers in transforming elemental materials like silicon, the laser interaction with more complex compounds like SiC has rarely been studied due to the complexity of compound phase transition and ultra-short processing time.
With high resolution microscope images and molecular dynamic simulations, scientists found that a single-pulse irradiation of xenon chloride excimer laser of 30 nanoseconds melts SiC, leading to the separation of a liquid SiC layer, a disordered carbon layer with graphitic domains (about 2.5 nm thick) on top surface and a polycrystalline silicon layer (about 5 nm) below carbon layer. Giving additional pulses causes the sublimation of the separated silicon, while the disordered carbon layer is transformed into a multilayer graphene.
"This research shows that the laser material interaction technology can be a powerful tool for the next generation of two dimensional nanomaterials," said Professor Lee.
Professor Choi added: "Using laser-induced phase separation of complex compounds, new types of two dimensional materials can be synthesized in the future."
High-resolution transmission electron microscopy shows that after just one laser pulse of 30 nanoseconds, the silicon carbide (SiC) substrate is melted and separates into a carbon and a silicon layer. More pulses cause the carbon layer to organize into graphene and the silicon to leave as gas.
Molecular dynamics simulates the graphene formation mechanism. The carbon layer on the top forms because the laser-induced liquid SiC (SiC (l)) is unstable.
(Press Release by Courtesy of the Institute for Basic Science (IBS))
2016.12.01 View 13284 -
 Technology to Allow Non-Magnetic Materials to Have Magnetic Properties by Professor Chan-Ho Yang
Professor Chan-Ho Yang and his research team from the Department of Physics at KAIST have developed a technology that allows non-magnetic materials to have magnetic properties or, in reverse, to remove magnetic properties from a magnet using an electric field.
Based on this research, it is expected that if magnetic-material-based data storage is developed, applications for high-speed massive data transfer will be possible.
The results of this research, with Ph.D. candidate Byung-Kwon Jang as the first author, were published online in Nature Physics on October 3.
Very small magnets exist inside of any materials. If the direction of the minuscule magnets is dis-aligned, pointing multiple directions, it is non-magnetic. If the direction is aligned in a certain direction, the material holds magnetic property just like any magnet we normally see.
Data storage capacity technology has rapidly advanced to the point where we can easily get a portable hard disk drive (HDD) with terabyte-level storage; however, the increase in storage is inevitably followed by slower data access speed for a storage device. Although HDDs are currently the most widely used data storage devices, their technical applications are limited due to their slow data access speed.
Other methods such as solid-state drives (SSDs), floating gates, and resistive switching have been developed as alternatives. Yet, they leave tracks every time data is written, and this can cause fatigue cumulative damage.
There have been many attempts to compose cells—the smallest data storage space on a storage device—with magnetic materials as that would enable faster data access speeds and remove fatigue cumulative damage. Generally, the techniques tried by researchers were to use induced magnetic fields through current flow. However, magnetic fields are very difficult to shield and can affect a large area. As a result, they alternate the magnetic property of adjacent cells. Because each cell cannot be adjusted one by one, it cannot also be arranged in a certain direction, and therefore, it is hard to change the magnetic state.
Professor Yang and his team adjusted the magnetic state by using magnetoelectric interaction to deal with this issue. Instead of using magnetic fields, magnetoelectric interaction is a method that uses an electric field to adjust the magnetic state. It has the advantage of smaller energy consumption as well.
Professor Yang's team demonstrated that cells facing random directions can be arranged in a certain direction by only inducing an electric field. In addition, the reverse was also proved to be feasible.
Until this research, most cases of previous findings were only feasible at extremely low temperatures or high temperatures, but the technology developed by the research team is practicable at room temperature by manipulating chemical pressure. It allows for a reversible magnetic state, and moreover, is non-volatile. Therefore, the results of this research are expected to provide the basis for developing next-generation information storage device.
Professor Yang said, “The changes in the electric magnetic state will be accompanied by entropy changes” and added, “Our research is expected to open new potential for future applications not only for magnetoelectric devices, but also for thermoelectric effect.”
This research has been worked on jointly with Dr. Si-Yong Choi from the Korea Institute of Materials Science, Prof. Yoon-Hee Jeong from the Pohang University of Science and Technology, Dr. Tae-Yeong Koo from the Pohang Accelerator Laboratory, Dr. Kyung-Tae Ko from the Max Planck Institute for Chemical Physics of Solids, Dr. Jun-Sik Lee and Dr. Hendrik Ohldag from the SLAC National Accelerator Laboratory of the United States, and Prof. Jan Seidel from the University of New South Wales of Australia.
The research was supported by the Mid-Career Researcher Program of the National Research Foundation of Korea, Global Research Network Support Project, Leading Research Center Support Project (Condensed Quantum Coherence Research Center), Global Frontier Project (Hybrid Interface Materials Research Group), and others.
Picture: The concept graphic for the electric-field-induced magnetic phase switching the magnetic direction
2016.11.04 View 8944
Technology to Allow Non-Magnetic Materials to Have Magnetic Properties by Professor Chan-Ho Yang
Professor Chan-Ho Yang and his research team from the Department of Physics at KAIST have developed a technology that allows non-magnetic materials to have magnetic properties or, in reverse, to remove magnetic properties from a magnet using an electric field.
Based on this research, it is expected that if magnetic-material-based data storage is developed, applications for high-speed massive data transfer will be possible.
The results of this research, with Ph.D. candidate Byung-Kwon Jang as the first author, were published online in Nature Physics on October 3.
Very small magnets exist inside of any materials. If the direction of the minuscule magnets is dis-aligned, pointing multiple directions, it is non-magnetic. If the direction is aligned in a certain direction, the material holds magnetic property just like any magnet we normally see.
Data storage capacity technology has rapidly advanced to the point where we can easily get a portable hard disk drive (HDD) with terabyte-level storage; however, the increase in storage is inevitably followed by slower data access speed for a storage device. Although HDDs are currently the most widely used data storage devices, their technical applications are limited due to their slow data access speed.
Other methods such as solid-state drives (SSDs), floating gates, and resistive switching have been developed as alternatives. Yet, they leave tracks every time data is written, and this can cause fatigue cumulative damage.
There have been many attempts to compose cells—the smallest data storage space on a storage device—with magnetic materials as that would enable faster data access speeds and remove fatigue cumulative damage. Generally, the techniques tried by researchers were to use induced magnetic fields through current flow. However, magnetic fields are very difficult to shield and can affect a large area. As a result, they alternate the magnetic property of adjacent cells. Because each cell cannot be adjusted one by one, it cannot also be arranged in a certain direction, and therefore, it is hard to change the magnetic state.
Professor Yang and his team adjusted the magnetic state by using magnetoelectric interaction to deal with this issue. Instead of using magnetic fields, magnetoelectric interaction is a method that uses an electric field to adjust the magnetic state. It has the advantage of smaller energy consumption as well.
Professor Yang's team demonstrated that cells facing random directions can be arranged in a certain direction by only inducing an electric field. In addition, the reverse was also proved to be feasible.
Until this research, most cases of previous findings were only feasible at extremely low temperatures or high temperatures, but the technology developed by the research team is practicable at room temperature by manipulating chemical pressure. It allows for a reversible magnetic state, and moreover, is non-volatile. Therefore, the results of this research are expected to provide the basis for developing next-generation information storage device.
Professor Yang said, “The changes in the electric magnetic state will be accompanied by entropy changes” and added, “Our research is expected to open new potential for future applications not only for magnetoelectric devices, but also for thermoelectric effect.”
This research has been worked on jointly with Dr. Si-Yong Choi from the Korea Institute of Materials Science, Prof. Yoon-Hee Jeong from the Pohang University of Science and Technology, Dr. Tae-Yeong Koo from the Pohang Accelerator Laboratory, Dr. Kyung-Tae Ko from the Max Planck Institute for Chemical Physics of Solids, Dr. Jun-Sik Lee and Dr. Hendrik Ohldag from the SLAC National Accelerator Laboratory of the United States, and Prof. Jan Seidel from the University of New South Wales of Australia.
The research was supported by the Mid-Career Researcher Program of the National Research Foundation of Korea, Global Research Network Support Project, Leading Research Center Support Project (Condensed Quantum Coherence Research Center), Global Frontier Project (Hybrid Interface Materials Research Group), and others.
Picture: The concept graphic for the electric-field-induced magnetic phase switching the magnetic direction
2016.11.04 View 8944 -
 Doctoral Student Receives the Best Paper Award from the International Metabolic Engineering Conference 2016
So Young Choi, a Ph.D. candidate at the Department of Chemical and Biomolecular Engineering at KAIST, received the Student and Young Investigator Poster Award at the 11th International Metabolic Engineering Conference held in Awaji, Japan on June 26-30.
Choi received the award for her research on one-step fermentative production of Poly(lactate-co-glycolate) (PLGA) from carbohydrates in Escherichia coli, which was published in the April 2016 issue of Nature Biotechnology.
In her paper, she presented a novel technology to synthesize PLGA, a non-natural copolymer, through a biological production process. Because of its biodegradability, non-toxicity, and biocompatibility, PLGA is widely used in biomedical and therapeutic applications, including surgical sutures, prosthetic devices, drug delivery, and tissue engineering.
Employing a metabolic engineering approach, Choi manipulated the metabolic pathway of an Escherichia coli bacterium to convert glucose and xylose into the biosynthesis of PLGA within the cell. Previously, PLGA could be obtained only through chemical synthesis.
Choi said, “I’m thrilled to receive an award from a flagship conference of my research field. Mindful of this recognition, I will continue my research to produce meaningful results, thereby contributing to the development of science and technology in Korea.”
The International Metabolic Engineering Conference is a leading professional gathering where state-of-the-art developments and achievements made in the field of metabolic engineering are shared. With the participation of about 400 professionals from all around the world, the conference participants discussed this year’s theme of “Design, Synthesis and System Integration for Metabolic Engineering.”
2016.07.07 View 12637
Doctoral Student Receives the Best Paper Award from the International Metabolic Engineering Conference 2016
So Young Choi, a Ph.D. candidate at the Department of Chemical and Biomolecular Engineering at KAIST, received the Student and Young Investigator Poster Award at the 11th International Metabolic Engineering Conference held in Awaji, Japan on June 26-30.
Choi received the award for her research on one-step fermentative production of Poly(lactate-co-glycolate) (PLGA) from carbohydrates in Escherichia coli, which was published in the April 2016 issue of Nature Biotechnology.
In her paper, she presented a novel technology to synthesize PLGA, a non-natural copolymer, through a biological production process. Because of its biodegradability, non-toxicity, and biocompatibility, PLGA is widely used in biomedical and therapeutic applications, including surgical sutures, prosthetic devices, drug delivery, and tissue engineering.
Employing a metabolic engineering approach, Choi manipulated the metabolic pathway of an Escherichia coli bacterium to convert glucose and xylose into the biosynthesis of PLGA within the cell. Previously, PLGA could be obtained only through chemical synthesis.
Choi said, “I’m thrilled to receive an award from a flagship conference of my research field. Mindful of this recognition, I will continue my research to produce meaningful results, thereby contributing to the development of science and technology in Korea.”
The International Metabolic Engineering Conference is a leading professional gathering where state-of-the-art developments and achievements made in the field of metabolic engineering are shared. With the participation of about 400 professionals from all around the world, the conference participants discussed this year’s theme of “Design, Synthesis and System Integration for Metabolic Engineering.”
2016.07.07 View 12637 -
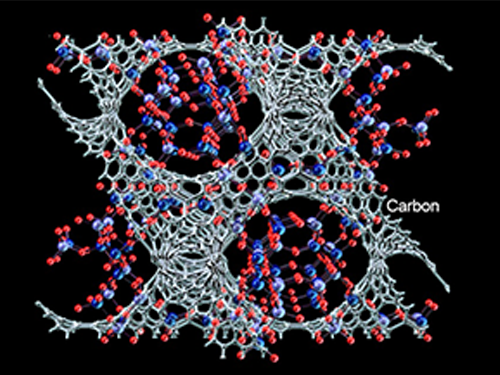 Synthesized Microporous 3D Graphene-like Carbons
Distinguished Professor Ryong Ryoo of the Chemistry Department at KAIST, who is also the Director of the Center for Nanomaterials and Carbon Materials at the Institute for Basic Science (IBS), and his research team have recently published their research results entitled "Lanthanum-catalysed Synthesis of Microporous 3D Graphene-like Carbons in a Zeolite Template" on June 29, 2016 in Nature on a new method to synthesize carbons having graphene structures with 3D periodic micropores, a trait resulted from using a zeolite as a template for the synthesis. The research team expects this technology to find a range of useful applications such as in batteries and catalysts.
Graphene, an allotrope of carbon, which was discovered more than a decade ago, has led to myriad research that seeks to unlock its vast potential. Zeolites, commonly used microporous solid catalysts in the petrochemical industry, have recently attracted attention in the field of material science as a template for carbon synthesis. Zeolites’ individual crystal is distinguished by its unique 1 nanometer (nm)-size pore structures. These structures facilitate the accommodation of carbon nanotubes inside zeolites. In their paper, the research team showed that these nanoporous systems are an ideal template for the carbon synthesis of three-dimensional (3D) graphene architecture, but zeolite pores are too small to accommodate bulky molecular compounds like polyaromatic and furfuryl alcohol that are often used in carbon synthesis. Small molecules like ethylene and acetylene can be used as a carbon source to achieve successful carbonization within zeolite pores, but it comes at a great cost. The high temperatures required for the synthesis cause reactions of carbons being deposited randomly on the external surfaces of zeolites as well as their internal pore walls, resulting in coke deposition and consequently, causing serious diffusion limitations in the zeolite pores.
The team from the IBS Center for Nanomaterials and Carbon Materials solved this conundrum with a novel approach. First author Dr. KIM Kyoungsoo explains: “The zeolite-template carbon synthesis has existed for a long time, but the problem with temperatures has foiled many scientists from extracting their full potential. Here, our team sought to find the answer by embedding lanthanum ions (La3+), a silvery-white metal element, in zeolite pores. This lowers the temperature required for the carbonization of ethylene or acetylene. Graphene-like sp2 carbon structures can be selectively formed inside the zeolite template, without carbon deposition at the external surfaces. After the zeolite template is removed, the carbon framework exhibits the electrical conductivity two orders of magnitude higher than amorphous mesoporous carbon, which is a pretty astonishing result. This highly efficient synthesis strategy based on the lanthanum ions renders the carbon framework to be formed in pores with a less than 1 nm diameter, just like as easily reproducible as in mesoporous templates. This provides a general method to synthesize carbon nanostructures with various topologies corresponding to the template zeolite pore topologies, such as FAU, EMT, beta, LTL, MFI, and LTA. Also, all the synthesis can be readily scaled up, which is important for practical applications in areas of batteries, fuel storage, and other zeolite-like catalyst supports.”
The research team began their experiment by utilizing La3+ ions. Dr. KIM elucidates why this silvery-white element proved so beneficial to the team, “La3+ ions are unreducible under carbonization process condition, so they can stay inside the zeolite pores instead of moving to the outer zeolite surface in the form of reduced metal particles. Within the pores, they can stabilize ethylene and the pyrocondensation intermediately to form a carbon framework in zeolites.”
In order to test this hypothesis, the team compared the amount of carbon deposited in La3+-containing form of Y zeolite (LaY) sample against a host of other samples such as NaY and HY. The experimental results indicate that all the LaY, NaY, and HY zeolite samples show rapid carbon deposition at 800°C. However, as the temperature decreases, there appears to be a dramatic difference between the different ionic forms of zeolites. At 600°C, the LaY zeolite is still active as a carbon deposition template. In contrast, both NaY and HY lose their carbon deposition functions almost completely.
The results, according to their paper published in Nature, highlight a catalytic effect of lanthanum for carbonization. By making graphene with 3D periodic nanoporous architectures, it promises a wide range of useful applications such as in batteries and catalysts but due to the lack of efficient synthetic strategies, such applications have not yet been successful. By taking advantage of the pore-selective carbon filling at decreased temperatures, the synthesis can readily be scaled up for studies requiring bulk quantities of carbon, in particular high electrical conductivity, which is a highly sought aspect for the production of batteries.
YouTube Link: https://youtu.be/lkNiHiB8lBk
Image 1: (Top to Bottom) Zeolite Template: Microporous Aluminosilicate; Zeolite ion exchanged with La3+ ions in aqueous solution; and Zeolite Template with La3+ ions
Image 2: (Top to Bottom) Catalytic carbonization progressed at La3+ ions-exchanged sites using ethylene as a carbon precursor. Carbon is highlighted in grey; Zeolite template removed in an acid solution (HF/ HCl); Microporous 3D graphene-like carbon
2016.07.01 View 10624
Synthesized Microporous 3D Graphene-like Carbons
Distinguished Professor Ryong Ryoo of the Chemistry Department at KAIST, who is also the Director of the Center for Nanomaterials and Carbon Materials at the Institute for Basic Science (IBS), and his research team have recently published their research results entitled "Lanthanum-catalysed Synthesis of Microporous 3D Graphene-like Carbons in a Zeolite Template" on June 29, 2016 in Nature on a new method to synthesize carbons having graphene structures with 3D periodic micropores, a trait resulted from using a zeolite as a template for the synthesis. The research team expects this technology to find a range of useful applications such as in batteries and catalysts.
Graphene, an allotrope of carbon, which was discovered more than a decade ago, has led to myriad research that seeks to unlock its vast potential. Zeolites, commonly used microporous solid catalysts in the petrochemical industry, have recently attracted attention in the field of material science as a template for carbon synthesis. Zeolites’ individual crystal is distinguished by its unique 1 nanometer (nm)-size pore structures. These structures facilitate the accommodation of carbon nanotubes inside zeolites. In their paper, the research team showed that these nanoporous systems are an ideal template for the carbon synthesis of three-dimensional (3D) graphene architecture, but zeolite pores are too small to accommodate bulky molecular compounds like polyaromatic and furfuryl alcohol that are often used in carbon synthesis. Small molecules like ethylene and acetylene can be used as a carbon source to achieve successful carbonization within zeolite pores, but it comes at a great cost. The high temperatures required for the synthesis cause reactions of carbons being deposited randomly on the external surfaces of zeolites as well as their internal pore walls, resulting in coke deposition and consequently, causing serious diffusion limitations in the zeolite pores.
The team from the IBS Center for Nanomaterials and Carbon Materials solved this conundrum with a novel approach. First author Dr. KIM Kyoungsoo explains: “The zeolite-template carbon synthesis has existed for a long time, but the problem with temperatures has foiled many scientists from extracting their full potential. Here, our team sought to find the answer by embedding lanthanum ions (La3+), a silvery-white metal element, in zeolite pores. This lowers the temperature required for the carbonization of ethylene or acetylene. Graphene-like sp2 carbon structures can be selectively formed inside the zeolite template, without carbon deposition at the external surfaces. After the zeolite template is removed, the carbon framework exhibits the electrical conductivity two orders of magnitude higher than amorphous mesoporous carbon, which is a pretty astonishing result. This highly efficient synthesis strategy based on the lanthanum ions renders the carbon framework to be formed in pores with a less than 1 nm diameter, just like as easily reproducible as in mesoporous templates. This provides a general method to synthesize carbon nanostructures with various topologies corresponding to the template zeolite pore topologies, such as FAU, EMT, beta, LTL, MFI, and LTA. Also, all the synthesis can be readily scaled up, which is important for practical applications in areas of batteries, fuel storage, and other zeolite-like catalyst supports.”
The research team began their experiment by utilizing La3+ ions. Dr. KIM elucidates why this silvery-white element proved so beneficial to the team, “La3+ ions are unreducible under carbonization process condition, so they can stay inside the zeolite pores instead of moving to the outer zeolite surface in the form of reduced metal particles. Within the pores, they can stabilize ethylene and the pyrocondensation intermediately to form a carbon framework in zeolites.”
In order to test this hypothesis, the team compared the amount of carbon deposited in La3+-containing form of Y zeolite (LaY) sample against a host of other samples such as NaY and HY. The experimental results indicate that all the LaY, NaY, and HY zeolite samples show rapid carbon deposition at 800°C. However, as the temperature decreases, there appears to be a dramatic difference between the different ionic forms of zeolites. At 600°C, the LaY zeolite is still active as a carbon deposition template. In contrast, both NaY and HY lose their carbon deposition functions almost completely.
The results, according to their paper published in Nature, highlight a catalytic effect of lanthanum for carbonization. By making graphene with 3D periodic nanoporous architectures, it promises a wide range of useful applications such as in batteries and catalysts but due to the lack of efficient synthetic strategies, such applications have not yet been successful. By taking advantage of the pore-selective carbon filling at decreased temperatures, the synthesis can readily be scaled up for studies requiring bulk quantities of carbon, in particular high electrical conductivity, which is a highly sought aspect for the production of batteries.
YouTube Link: https://youtu.be/lkNiHiB8lBk
Image 1: (Top to Bottom) Zeolite Template: Microporous Aluminosilicate; Zeolite ion exchanged with La3+ ions in aqueous solution; and Zeolite Template with La3+ ions
Image 2: (Top to Bottom) Catalytic carbonization progressed at La3+ ions-exchanged sites using ethylene as a carbon precursor. Carbon is highlighted in grey; Zeolite template removed in an acid solution (HF/ HCl); Microporous 3D graphene-like carbon
2016.07.01 View 10624 -
 Graphene-Based Transparent Electrodes for Highly Efficient Flexible OLEDs
A Korean research team developed an ideal electrode structure composed of graphene and layers of titanium dioxide and conducting polymers, resulting in highly flexible and efficient OLEDs.
The arrival of a thin and lightweight computer that even rolls up like a piece of paper will not be in the far distant future. Flexible organic light-emitting diodes (OLEDs), built upon a plastic substrate, have received greater attention lately for their use in next-generation displays that can be bent or rolled while still operating.
A Korean research team led by Professor Seunghyup Yoo from the School of Electrical Engineering, KAIST and Professor Tae-Woo Lee from the Department of Materials Science and Engineering, Pohang University of Science and Technology (POSTECH) has developed highly flexible OLEDs with excellent efficiency by using graphene as a transparent electrode (TE) which is placed in between titanium dioxide (TiO2) and conducting polymer layers. The research results were published online on June 2, 2016 in Nature Communications.
OLEDs are stacked in several ultra-thin layers on glass, foil, or plastic substrates, in which multi-layers of organic compounds are sandwiched between two electrodes (cathode and anode). When voltage is applied across the electrodes, electrons from the cathode and holes (positive charges) from the anode draw toward each other and meet in the emissive layer. OLEDs emit light as an electron recombines with a positive hole, releasing energy in the form of a photon. One of the electrodes in OLEDs is usually transparent, and depending on which electrode is transparent, OLEDs can either emit from the top or bottom.
In conventional bottom-emission OLEDs, an anode is transparent in order for the emitted photons to exit the device through its substrate. Indium-tin-oxide (ITO) is commonly used as a transparent anode because of its high transparency, low sheet resistance, and well-established manufacturing process. However, ITO can potentially be expensive, and moreover, is brittle, being susceptible to bending-induced formation of cracks.
Graphene, a two-dimensional thin layer of carbon atoms tightly bonded together in a hexagonal honeycomb lattice, has recently emerged as an alternative to ITO. With outstanding electrical, physical, and chemical properties, its atomic thinness leading to a high degree of flexibility and transparency makes it an ideal candidate for TEs. Nonetheless, the efficiency of graphene-based OLEDs reported to date has been, at best, about the same level of ITO-based OLEDs.
As a solution, the Korean research team, which further includes Professors Sung-Yool Choi (Electrical Engineering) and Taek-Soo Kim (Mechanical Engineering) of KAIST and their students, proposed a new device architecture that can maximize the efficiency of graphene-based OLEDs. They fabricated a transparent anode in a composite structure in which a TiO2 layer with a high refractive index (high-n) and a hole-injection layer (HIL) of conducting polymers with a low refractive index (low-n) sandwich graphene electrodes. This is an optical design that induces a synergistic collaboration between the high-n and low-n layers to increase the effective reflectance of TEs. As a result, the enhancement of the optical cavity resonance is maximized. The optical cavity resonance is related to the improvement of efficiency and color gamut in OLEDs. At the same time, the loss from surface plasmon polariton (SPP), a major cause for weak photon emissions in OLEDs, is also reduced due to the presence of the low-n conducting polymers.
Under this approach, graphene-based OLEDs exhibit 40.8% of ultrahigh external quantum efficiency (EQE) and 160.3 lm/W of power efficiency, which is unprecedented in those using graphene as a TE. Furthermore, these devices remain intact and operate well even after 1,000 bending cycles at a radius of curvature as small as 2.3 mm. This is a remarkable result for OLEDs containing oxide layers such as TiO2 because oxides are typically brittle and prone to bending-induced fractures even at a relatively low strain. The research team discovered that TiO2 has a crack-deflection toughening mechanism that tends to prevent bending-induced cracks from being formed easily.
Professor Yoo said, “What’s unique and advanced about this technology, compared with previous graphene-based OLEDs, is the synergistic collaboration of high- and low-index layers that enables optical management of both resonance effect and SPP loss, leading to significant enhancement in efficiency, all with little compromise in flexibility.” He added, “Our work was the achievement of collaborative research, transcending the boundaries of different fields, through which we have often found meaningful breakthroughs.”
Professor Lee said, “We expect that our technology will pave the way to develop an OLED light source for highly flexible and wearable displays, or flexible sensors that can be attached to the human body for health monitoring, for instance.”
The research paper is entitled “Synergistic Electrode Architecture for Efficient Graphene-based Flexible Organic Light-emitting Diodes” (DOI. 10.1038/NCOMMS11791). The lead authors are Jae-Ho Lee, a Ph.D. candidate at KAIST; Tae-Hee Han, a Ph.D. researcher at POSTECH; and Min-Ho Park, a Ph.D. candidate at POSTECH.
This study was supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF) through the Center for Advanced Flexible Display (CAFDC) funded by the Ministry of Science, ICT and Future Planning (MSIP); by the Center for Advanced Soft-Electronics funded by the MSIP as a Global Frontier Project; by the Graphene Research Center Program of KAIST; and by grants from the IT R&D Program of the Ministry of Trade, Industry and Energy of Korea (MOTIE).
Figure 1: Application of Graphene-based OLEDs
This picture shows an OLED with the composite structure of TiO2/graphene/conducting polymer electrode in operation. The OLED exhibits 40.8% of ultrahigh external quantum efficiency (EQE) and 160.3 lm/W of power efficiency. The device prepared on a plastic substrate shown in the right remains intact and operates well even after 1,000 bending cycles at a radius of curvature as small as 2.3 mm.
Figure 2: Schematic Device Structure of Graphene-based OLEDs
This picture shows the new architecture to develop highly flexible OLEDs with excellent efficiency by using graphene as a transparent electrode (TE).
2016.06.07 View 16114
Graphene-Based Transparent Electrodes for Highly Efficient Flexible OLEDs
A Korean research team developed an ideal electrode structure composed of graphene and layers of titanium dioxide and conducting polymers, resulting in highly flexible and efficient OLEDs.
The arrival of a thin and lightweight computer that even rolls up like a piece of paper will not be in the far distant future. Flexible organic light-emitting diodes (OLEDs), built upon a plastic substrate, have received greater attention lately for their use in next-generation displays that can be bent or rolled while still operating.
A Korean research team led by Professor Seunghyup Yoo from the School of Electrical Engineering, KAIST and Professor Tae-Woo Lee from the Department of Materials Science and Engineering, Pohang University of Science and Technology (POSTECH) has developed highly flexible OLEDs with excellent efficiency by using graphene as a transparent electrode (TE) which is placed in between titanium dioxide (TiO2) and conducting polymer layers. The research results were published online on June 2, 2016 in Nature Communications.
OLEDs are stacked in several ultra-thin layers on glass, foil, or plastic substrates, in which multi-layers of organic compounds are sandwiched between two electrodes (cathode and anode). When voltage is applied across the electrodes, electrons from the cathode and holes (positive charges) from the anode draw toward each other and meet in the emissive layer. OLEDs emit light as an electron recombines with a positive hole, releasing energy in the form of a photon. One of the electrodes in OLEDs is usually transparent, and depending on which electrode is transparent, OLEDs can either emit from the top or bottom.
In conventional bottom-emission OLEDs, an anode is transparent in order for the emitted photons to exit the device through its substrate. Indium-tin-oxide (ITO) is commonly used as a transparent anode because of its high transparency, low sheet resistance, and well-established manufacturing process. However, ITO can potentially be expensive, and moreover, is brittle, being susceptible to bending-induced formation of cracks.
Graphene, a two-dimensional thin layer of carbon atoms tightly bonded together in a hexagonal honeycomb lattice, has recently emerged as an alternative to ITO. With outstanding electrical, physical, and chemical properties, its atomic thinness leading to a high degree of flexibility and transparency makes it an ideal candidate for TEs. Nonetheless, the efficiency of graphene-based OLEDs reported to date has been, at best, about the same level of ITO-based OLEDs.
As a solution, the Korean research team, which further includes Professors Sung-Yool Choi (Electrical Engineering) and Taek-Soo Kim (Mechanical Engineering) of KAIST and their students, proposed a new device architecture that can maximize the efficiency of graphene-based OLEDs. They fabricated a transparent anode in a composite structure in which a TiO2 layer with a high refractive index (high-n) and a hole-injection layer (HIL) of conducting polymers with a low refractive index (low-n) sandwich graphene electrodes. This is an optical design that induces a synergistic collaboration between the high-n and low-n layers to increase the effective reflectance of TEs. As a result, the enhancement of the optical cavity resonance is maximized. The optical cavity resonance is related to the improvement of efficiency and color gamut in OLEDs. At the same time, the loss from surface plasmon polariton (SPP), a major cause for weak photon emissions in OLEDs, is also reduced due to the presence of the low-n conducting polymers.
Under this approach, graphene-based OLEDs exhibit 40.8% of ultrahigh external quantum efficiency (EQE) and 160.3 lm/W of power efficiency, which is unprecedented in those using graphene as a TE. Furthermore, these devices remain intact and operate well even after 1,000 bending cycles at a radius of curvature as small as 2.3 mm. This is a remarkable result for OLEDs containing oxide layers such as TiO2 because oxides are typically brittle and prone to bending-induced fractures even at a relatively low strain. The research team discovered that TiO2 has a crack-deflection toughening mechanism that tends to prevent bending-induced cracks from being formed easily.
Professor Yoo said, “What’s unique and advanced about this technology, compared with previous graphene-based OLEDs, is the synergistic collaboration of high- and low-index layers that enables optical management of both resonance effect and SPP loss, leading to significant enhancement in efficiency, all with little compromise in flexibility.” He added, “Our work was the achievement of collaborative research, transcending the boundaries of different fields, through which we have often found meaningful breakthroughs.”
Professor Lee said, “We expect that our technology will pave the way to develop an OLED light source for highly flexible and wearable displays, or flexible sensors that can be attached to the human body for health monitoring, for instance.”
The research paper is entitled “Synergistic Electrode Architecture for Efficient Graphene-based Flexible Organic Light-emitting Diodes” (DOI. 10.1038/NCOMMS11791). The lead authors are Jae-Ho Lee, a Ph.D. candidate at KAIST; Tae-Hee Han, a Ph.D. researcher at POSTECH; and Min-Ho Park, a Ph.D. candidate at POSTECH.
This study was supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF) through the Center for Advanced Flexible Display (CAFDC) funded by the Ministry of Science, ICT and Future Planning (MSIP); by the Center for Advanced Soft-Electronics funded by the MSIP as a Global Frontier Project; by the Graphene Research Center Program of KAIST; and by grants from the IT R&D Program of the Ministry of Trade, Industry and Energy of Korea (MOTIE).
Figure 1: Application of Graphene-based OLEDs
This picture shows an OLED with the composite structure of TiO2/graphene/conducting polymer electrode in operation. The OLED exhibits 40.8% of ultrahigh external quantum efficiency (EQE) and 160.3 lm/W of power efficiency. The device prepared on a plastic substrate shown in the right remains intact and operates well even after 1,000 bending cycles at a radius of curvature as small as 2.3 mm.
Figure 2: Schematic Device Structure of Graphene-based OLEDs
This picture shows the new architecture to develop highly flexible OLEDs with excellent efficiency by using graphene as a transparent electrode (TE).
2016.06.07 View 16114 -
 KAIST Team Develops Technology to Enable Unzipping of the Graphene Plane
Professor Sang-Wook Kim’s research team of the Material Science and Engineering Department has developed a technique, which enables unzipping of the graphene plane without uncontrollable damage. The research findings were published online on the January 22 issue of Nature Communications.
Graphene is a form of carbon in which its atoms form a honey-comb structure through chemical bonding. If this structure can be cut to a desired form, other carbon materials with nanostructure can be created. Many researchers have tried to obtain the accurate unzipping of graphene structures, but faced challenges doing so.
To break a very strong bond between carbon atoms, an equivalently strong chemical reaction must be induced. But the chemical reaction not only cuts out the desirable borders, but also damages the surrounding ones. Conventional techniques, which cut out graphene at once, damaged the chemical properties of the graphene structure after unzipping. This is similar to wearing out paper while manipulating it.
To solve this problem, the research team adopted “heteroatom doping.” The idea is similar to a sheet of paper being split following a groove drawn on the sheet. After making some regions of the structure unstable by doping other atoms such as nitrogen on a carbon plane, the regions are electrochemically stimulated to split the parts. Nitrogen or other atoms act as the groove on the grapheme plane.
The researchers finely controlled the amount of unzipping graphene by adjusting the amount of heteroatom dopants, from which they were able to create a quality nano graphene without any damage in its 2-dimensional crystalline structure. Using this technique, the researchers were able to obtain a capacitor with state-of-the-art energy transfer speed. The nano graphene can be combined with polymer, metal, and semiconductor nano molecules to form carbon composites.
Professor Kim said, “In order to commercialize this technique, heteroatom doping should be researched further. We plan to develop fabric-like carbon materials with excellent mechanical and electrical properties using this technique.”
Picture 1: Unzipped Carbon Nano Tube
Picture 2: Development of Nano Graphene from Carbon Nano Tube Using Heteroatom Dopants
Korean descriptions translated into English:
Unzipping Process of Graphene
Carbon Nano Tube → Nano Graphen
Heteroatom
This process is similar to a paper being split in two from a tiny hole punched therein.
2016.03.22 View 10091
KAIST Team Develops Technology to Enable Unzipping of the Graphene Plane
Professor Sang-Wook Kim’s research team of the Material Science and Engineering Department has developed a technique, which enables unzipping of the graphene plane without uncontrollable damage. The research findings were published online on the January 22 issue of Nature Communications.
Graphene is a form of carbon in which its atoms form a honey-comb structure through chemical bonding. If this structure can be cut to a desired form, other carbon materials with nanostructure can be created. Many researchers have tried to obtain the accurate unzipping of graphene structures, but faced challenges doing so.
To break a very strong bond between carbon atoms, an equivalently strong chemical reaction must be induced. But the chemical reaction not only cuts out the desirable borders, but also damages the surrounding ones. Conventional techniques, which cut out graphene at once, damaged the chemical properties of the graphene structure after unzipping. This is similar to wearing out paper while manipulating it.
To solve this problem, the research team adopted “heteroatom doping.” The idea is similar to a sheet of paper being split following a groove drawn on the sheet. After making some regions of the structure unstable by doping other atoms such as nitrogen on a carbon plane, the regions are electrochemically stimulated to split the parts. Nitrogen or other atoms act as the groove on the grapheme plane.
The researchers finely controlled the amount of unzipping graphene by adjusting the amount of heteroatom dopants, from which they were able to create a quality nano graphene without any damage in its 2-dimensional crystalline structure. Using this technique, the researchers were able to obtain a capacitor with state-of-the-art energy transfer speed. The nano graphene can be combined with polymer, metal, and semiconductor nano molecules to form carbon composites.
Professor Kim said, “In order to commercialize this technique, heteroatom doping should be researched further. We plan to develop fabric-like carbon materials with excellent mechanical and electrical properties using this technique.”
Picture 1: Unzipped Carbon Nano Tube
Picture 2: Development of Nano Graphene from Carbon Nano Tube Using Heteroatom Dopants
Korean descriptions translated into English:
Unzipping Process of Graphene
Carbon Nano Tube → Nano Graphen
Heteroatom
This process is similar to a paper being split in two from a tiny hole punched therein.
2016.03.22 View 10091 -
 Non-Natural Biomedical Polymers Produced from Microorganisms
KAIST researchers have developed metabolically engineered Escherichia coli strains to synthesize non-natural, biomedically important polymers including poly(lactate-co-glycolate) (PLGA), previously considered impossible to obtain from biobased materials.
Renewable non-food biomass could potentially replace petrochemical raw materials to produce energy sources, useful chemicals, or a vast array of petroleum-based end products such as plastics, lubricants, paints, fertilizers, and vitamin capsules. In recent years, biorefineries which transform non-edible biomass into fuel, heat, power, chemicals, and materials have received a great deal of attention as a sustainable alternative to decreasing the reliance on fossil fuels.
A research team headed by Distinguished Professor Sang Yup Lee of the Chemical and Biomolecular Engineering Department at KAIST has established a biorefinery system to create non-natural polymers from natural sources, allowing various plastics to be made in an environmentally-friendly and sustainable manner. The research results were published online in Nature Biotechnology on March 7, 2016. The print version will be issued in April 2016.
The research team adopted a systems metabolic engineering approach to develop a microorganism that can produce diverse non-natural, biomedically important polymers and succeeded in synthesizing poly(lactate-co-glycolate) (PLGA), a copolymer of two different polymer monomers, lactic and glycolic acid. PLGA is biodegradable, biocompatible, and non-toxic, and has been widely used in biomedical and therapeutic applications such as surgical sutures, prosthetic devices, drug delivery, and tissue engineering.
Inspired by the biosynthesis process for polyhydroxyalkanoates (PHAs), biologically-derived polyesters produced in nature by the bacterial fermentation of sugar or lipids, the research team designed a metabolic pathway for the biosynthesis of PLGA through microbial fermentation directly from carbohydrates in Escherichia coli (E. coli) strains.
The team had previously reported a recombinant E. coli producing PLGA by using the glyoxylate shunt pathway for the generation of glycolate from glucose, which was disclosed in their patents KR10-1575585-0000 (filing date of March 11, 2011), US08883463 and JP5820363. However, they discovered that the polymer content and glycolate fraction of PLGA could not be significantly enhanced via further engineering techniques. Thus, in this research, the team introduced a heterologous pathway to produce glycolate from xylose and succeeded in developing the recombinant E. coli producing PLGA and various novel copolymers much more efficiently.
In order to produce PLGA by microbial fermentation directly from carbohydrates, the team incorporated external and engineered enzymes as catalysts to co-polymerize PLGA while establishing a few additional metabolic pathways for the biosynthesis to produce a range of different non-natural polymers, some for the first time. This bio-based synthetic process for PLGA and other polymers could substitute for the existing complicated chemical production that involves the preparation and purification of precursors, chemical polymerization processes, and the elimination of metal catalysts.
Professor Lee and his team performed in silico genome-scale metabolic simulations of the E. coli cell to predict and analyze changes in the metabolic fluxes of cells which were caused by the introduction of external metabolic pathways. Based on these results, genes are manipulated to optimize metabolic fluxes by eliminating the genes responsible for byproducts formation and enhancing the expression levels of certain genes, thereby achieving the effective production of target polymers as well as stimulating cell growth.
The team utilized the structural basis of broad substrate specificity of the key synthesizing enzyme, PHA synthase, to incorporate various co-monomers with main and side chains of different lengths. These monomers were produced inside the cell by metabolic engineering, and then copolymerized to improve the material properties of PLGA. As a result, a variety of PLGA copolymers with different monomer compositions such as the US Food and Drug Administration (FDA)-approved monomers, 3-hydroxyburate, 4-hydroxyburate, and 6-hydroxyhexanoate, were produced. Newly applied bioplastics such as 5-hydroxyvalerate and 2-hydroxyisovalerate were also made.
The team employed a systems metabolic engineering application which, according to the researchers, is the first successful example of biological production of PGLA and several novel copolymers from renewable biomass by one-step direct fermentation of metabolically engineered E.coli.
Professor Lee said, “We presented important findings that non-natural polymers, such as PLGA which is commonly used for drug delivery or biomedical devices, were produced by a metabolically engineered gut bacterium. Our research is meaningful in that it proposes a platform strategy in metabolic engineering, which can be further utilized in the development of numerous non-natural, useful polymers.”
Director Ilsub Baek at the Platform Technology Division of the Ministry of Science, ICT and Future Planning of Korea, who oversees the Technology Development Program to Solve Climate Change, said, “Professor Lee has led one of our research projects, the Systems Metabolic Engineering for Biorefineries, which began as part of the Ministry’s Technology Development Program to Solve Climate Change. He and his team have continuously achieved promising results and been attracting greater interest from the global scientific community. As climate change technology grows more important, this research on the biological production of non-natural, high value polymers will have a great impact on science and industry.”
The title of the research paper is “One-step Fermentative Production of Poly(lactate-co-glycolate) from Carbohydrates in Escherichia coli (DOI: 10.1038/nbt.3485).” The lead authors are So Young Choi, a Ph.D. candidate in the Department of Chemical and Biomolecular Engineering at KAIST, and Si Jae Park, Assistant Professor of the Environmental Engineering and Energy Department at Myongji University. Won Jun Kim and Jung Eun Yang, both doctoral students in the Department of Chemical and Biomolecular Engineering at KAIST, also participated in the research.
This research was supported by the Technology Development Program to Solve Climate Change’s research project titled “Systems Metabolic Engineering for Biorefineries” from the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea (NRF-2012M1A2A2026556).
Figure: Production of PLGA and Other Non-Natural Copolymers
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced a variety of PLGAs with different monomer compositions, proposing the chemosynthetic process of non-natural polymers from biomass. The non-natural polymer PLGA and its other copolymers, which were produced by engineered bacteria developed by taking a systems metabolic engineering approach, accumulate in granule forms within a cell.
2016.03.08 View 13369
Non-Natural Biomedical Polymers Produced from Microorganisms
KAIST researchers have developed metabolically engineered Escherichia coli strains to synthesize non-natural, biomedically important polymers including poly(lactate-co-glycolate) (PLGA), previously considered impossible to obtain from biobased materials.
Renewable non-food biomass could potentially replace petrochemical raw materials to produce energy sources, useful chemicals, or a vast array of petroleum-based end products such as plastics, lubricants, paints, fertilizers, and vitamin capsules. In recent years, biorefineries which transform non-edible biomass into fuel, heat, power, chemicals, and materials have received a great deal of attention as a sustainable alternative to decreasing the reliance on fossil fuels.
A research team headed by Distinguished Professor Sang Yup Lee of the Chemical and Biomolecular Engineering Department at KAIST has established a biorefinery system to create non-natural polymers from natural sources, allowing various plastics to be made in an environmentally-friendly and sustainable manner. The research results were published online in Nature Biotechnology on March 7, 2016. The print version will be issued in April 2016.
The research team adopted a systems metabolic engineering approach to develop a microorganism that can produce diverse non-natural, biomedically important polymers and succeeded in synthesizing poly(lactate-co-glycolate) (PLGA), a copolymer of two different polymer monomers, lactic and glycolic acid. PLGA is biodegradable, biocompatible, and non-toxic, and has been widely used in biomedical and therapeutic applications such as surgical sutures, prosthetic devices, drug delivery, and tissue engineering.
Inspired by the biosynthesis process for polyhydroxyalkanoates (PHAs), biologically-derived polyesters produced in nature by the bacterial fermentation of sugar or lipids, the research team designed a metabolic pathway for the biosynthesis of PLGA through microbial fermentation directly from carbohydrates in Escherichia coli (E. coli) strains.
The team had previously reported a recombinant E. coli producing PLGA by using the glyoxylate shunt pathway for the generation of glycolate from glucose, which was disclosed in their patents KR10-1575585-0000 (filing date of March 11, 2011), US08883463 and JP5820363. However, they discovered that the polymer content and glycolate fraction of PLGA could not be significantly enhanced via further engineering techniques. Thus, in this research, the team introduced a heterologous pathway to produce glycolate from xylose and succeeded in developing the recombinant E. coli producing PLGA and various novel copolymers much more efficiently.
In order to produce PLGA by microbial fermentation directly from carbohydrates, the team incorporated external and engineered enzymes as catalysts to co-polymerize PLGA while establishing a few additional metabolic pathways for the biosynthesis to produce a range of different non-natural polymers, some for the first time. This bio-based synthetic process for PLGA and other polymers could substitute for the existing complicated chemical production that involves the preparation and purification of precursors, chemical polymerization processes, and the elimination of metal catalysts.
Professor Lee and his team performed in silico genome-scale metabolic simulations of the E. coli cell to predict and analyze changes in the metabolic fluxes of cells which were caused by the introduction of external metabolic pathways. Based on these results, genes are manipulated to optimize metabolic fluxes by eliminating the genes responsible for byproducts formation and enhancing the expression levels of certain genes, thereby achieving the effective production of target polymers as well as stimulating cell growth.
The team utilized the structural basis of broad substrate specificity of the key synthesizing enzyme, PHA synthase, to incorporate various co-monomers with main and side chains of different lengths. These monomers were produced inside the cell by metabolic engineering, and then copolymerized to improve the material properties of PLGA. As a result, a variety of PLGA copolymers with different monomer compositions such as the US Food and Drug Administration (FDA)-approved monomers, 3-hydroxyburate, 4-hydroxyburate, and 6-hydroxyhexanoate, were produced. Newly applied bioplastics such as 5-hydroxyvalerate and 2-hydroxyisovalerate were also made.
The team employed a systems metabolic engineering application which, according to the researchers, is the first successful example of biological production of PGLA and several novel copolymers from renewable biomass by one-step direct fermentation of metabolically engineered E.coli.
Professor Lee said, “We presented important findings that non-natural polymers, such as PLGA which is commonly used for drug delivery or biomedical devices, were produced by a metabolically engineered gut bacterium. Our research is meaningful in that it proposes a platform strategy in metabolic engineering, which can be further utilized in the development of numerous non-natural, useful polymers.”
Director Ilsub Baek at the Platform Technology Division of the Ministry of Science, ICT and Future Planning of Korea, who oversees the Technology Development Program to Solve Climate Change, said, “Professor Lee has led one of our research projects, the Systems Metabolic Engineering for Biorefineries, which began as part of the Ministry’s Technology Development Program to Solve Climate Change. He and his team have continuously achieved promising results and been attracting greater interest from the global scientific community. As climate change technology grows more important, this research on the biological production of non-natural, high value polymers will have a great impact on science and industry.”
The title of the research paper is “One-step Fermentative Production of Poly(lactate-co-glycolate) from Carbohydrates in Escherichia coli (DOI: 10.1038/nbt.3485).” The lead authors are So Young Choi, a Ph.D. candidate in the Department of Chemical and Biomolecular Engineering at KAIST, and Si Jae Park, Assistant Professor of the Environmental Engineering and Energy Department at Myongji University. Won Jun Kim and Jung Eun Yang, both doctoral students in the Department of Chemical and Biomolecular Engineering at KAIST, also participated in the research.
This research was supported by the Technology Development Program to Solve Climate Change’s research project titled “Systems Metabolic Engineering for Biorefineries” from the Ministry of Science, ICT and Future Planning through the National Research Foundation of Korea (NRF-2012M1A2A2026556).
Figure: Production of PLGA and Other Non-Natural Copolymers
This schematic diagram shows the overall conceptualization of how metabolically engineered E. coli produced a variety of PLGAs with different monomer compositions, proposing the chemosynthetic process of non-natural polymers from biomass. The non-natural polymer PLGA and its other copolymers, which were produced by engineered bacteria developed by taking a systems metabolic engineering approach, accumulate in granule forms within a cell.
2016.03.08 View 13369 -
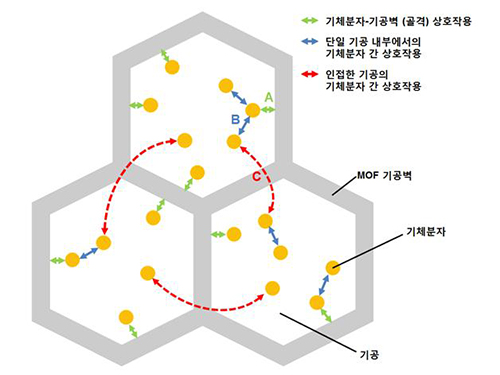 A New Way to Look at MOFs
An international research team composed of researchers from KAIST (led by Professors Osamu Terasaki and Jeung Ku Kang at the Graduate School of Energy, Environment, Water and Sustainability) and other universities, including UC Berkeley, has recently published research results on the adsorption process of metal-organic frameworks (MOFs) in Nature (November 9, 2015).
MOFs are porous three-dimensional crystals with a high internal surface area, which have a wide range of applications involving adsorption such as hydrogen, methane, or carbon dioxide storage. In the paper entitled “Extra Adsorption and Adsorbate Superlattice Formation in Metal-organic Frameworks,” the research team described their observation of a very specific interpore interaction process in MOFs.
For additional information, please see:
A New Way to Look at MOFs
International study challenges prevailing view on how metal organic frameworks store gases
EurekAlert, November 9, 2015
http://www.eurekalert.org/pub_releases/2015-11/dbnl-anw110915.php
(Courtesy of the US Department of Energy and Lawrence Berkeley National Laboratory news release)
2015.11.13 View 9317
A New Way to Look at MOFs
An international research team composed of researchers from KAIST (led by Professors Osamu Terasaki and Jeung Ku Kang at the Graduate School of Energy, Environment, Water and Sustainability) and other universities, including UC Berkeley, has recently published research results on the adsorption process of metal-organic frameworks (MOFs) in Nature (November 9, 2015).
MOFs are porous three-dimensional crystals with a high internal surface area, which have a wide range of applications involving adsorption such as hydrogen, methane, or carbon dioxide storage. In the paper entitled “Extra Adsorption and Adsorbate Superlattice Formation in Metal-organic Frameworks,” the research team described their observation of a very specific interpore interaction process in MOFs.
For additional information, please see:
A New Way to Look at MOFs
International study challenges prevailing view on how metal organic frameworks store gases
EurekAlert, November 9, 2015
http://www.eurekalert.org/pub_releases/2015-11/dbnl-anw110915.php
(Courtesy of the US Department of Energy and Lawrence Berkeley National Laboratory news release)
2015.11.13 View 9317 -
 Mapping the Folding Process of a Single Membrane Protein
KAIST and UCLA scientists were able to observe an individual membrane protein fold and unfold by pulling and releasing magnetically trapped protein molecules.
Proteins are huge molecules containing hundreds to thousands of atoms that adopt a unique three dimensional structure, placing chemical groups in just the right place to catalyze reactions or build cellular structures. How all those atoms manage to find the right location - the so-called folding problem - has fascinated molecular biologists since the first structures were seen in the 1950s. Moreover, folding has important medical implications because most genetic defects cause protein misfolding.
About a third of all proteins float around in the cell membrane where they ensure the right chemicals get in the cell in the right amounts. Membrane proteins also provide key information links between the cell and its environment. Indeed, most drugs target membrane proteins. Nevertheless, the folding of membrane proteins has been particularly difficult to study and has rarely been studied in natural environments, leaving the folding process for a large fraction of the protein universe still largely cloaked in mystery.
In a recent issue of Nature Chemical Biology, published on October 19, 2015, a research team led by Tae-Young Yoon of the Department of Physics at the Korea Advanced Institute of Science and Technology (KAIST) and James U. Bowie of the Department of Chemistry and Biochemistry at the University of California, Los Angeles (UCLA), report a new method for manipulating the folding of membrane proteins in a membrane environment using a tool called a magnetic tweezer.
Researchers first attach long DNA handles to the ends of the protein. One handle is attached to a glass surface and the other to a magnetic bead. Using a magnet, they can essentially grab the protein and pull on it, inducing it to unfold. By playing with the bead attached to the protein, they can force the protein to unfold or allow it to refold, and watch all this happening by 3D-tracking of the magnetic bead. With this novel strategy, they were able to quantitatively map the folding energy landscape, the folding kinetic rate, and folding intermediates of a membrane protein in a membrane environment for the first time.
“I have been dreaming about this experiment for a decade. To see it work so well is really gratifying,” said Dr. Bowie.
One of the major surprises in the study was that essentially all the atoms of the protein jump into the correct structure together. The researchers expected that the protein structure would come together in a more piecemeal fashion, with different parts of the structure forming separately, but that was not the case. It is possible that nature evolved such a smooth, highly cooperative folding process to prevent partially folded forms that could get into trouble in the crowded cell membrane. On the other hand, the cooperative folding seen here might not apply to other membrane proteins.
“We need to look at more proteins. The technique developed here may allow us to do just that,” said Dr. Yoon.
The single molecule mechanical manipulation technique could enable detailed folding studies of many other membrane proteins. A major barrier to the study of membrane proteins previously is that the proteins tend to stick together and get tangled up, as computer cords lying at your feet tend to do. With the tweezer technique used in this work, the protein cords are held apart from other cords so they can’t get knotted up. It is hoped that the new approach will open up an important part of the protein universe to scrutiny, including many proteins that become misfolded in disease states.
The title of the research paper is “Mapping the energy landscape for second-stage folding of a single membrane protein” (DOI: 10.1038/nchembio.1939).
Picture: Single-molecule magnetic tweezers that induce mechanical unfolding and refolding of a single membrane protein. Since the force applied is parallel to the biological lipid membrane, the unfolding and refolding processes occur within the membrane.
2015.10.20 View 11029
Mapping the Folding Process of a Single Membrane Protein
KAIST and UCLA scientists were able to observe an individual membrane protein fold and unfold by pulling and releasing magnetically trapped protein molecules.
Proteins are huge molecules containing hundreds to thousands of atoms that adopt a unique three dimensional structure, placing chemical groups in just the right place to catalyze reactions or build cellular structures. How all those atoms manage to find the right location - the so-called folding problem - has fascinated molecular biologists since the first structures were seen in the 1950s. Moreover, folding has important medical implications because most genetic defects cause protein misfolding.
About a third of all proteins float around in the cell membrane where they ensure the right chemicals get in the cell in the right amounts. Membrane proteins also provide key information links between the cell and its environment. Indeed, most drugs target membrane proteins. Nevertheless, the folding of membrane proteins has been particularly difficult to study and has rarely been studied in natural environments, leaving the folding process for a large fraction of the protein universe still largely cloaked in mystery.
In a recent issue of Nature Chemical Biology, published on October 19, 2015, a research team led by Tae-Young Yoon of the Department of Physics at the Korea Advanced Institute of Science and Technology (KAIST) and James U. Bowie of the Department of Chemistry and Biochemistry at the University of California, Los Angeles (UCLA), report a new method for manipulating the folding of membrane proteins in a membrane environment using a tool called a magnetic tweezer.
Researchers first attach long DNA handles to the ends of the protein. One handle is attached to a glass surface and the other to a magnetic bead. Using a magnet, they can essentially grab the protein and pull on it, inducing it to unfold. By playing with the bead attached to the protein, they can force the protein to unfold or allow it to refold, and watch all this happening by 3D-tracking of the magnetic bead. With this novel strategy, they were able to quantitatively map the folding energy landscape, the folding kinetic rate, and folding intermediates of a membrane protein in a membrane environment for the first time.
“I have been dreaming about this experiment for a decade. To see it work so well is really gratifying,” said Dr. Bowie.
One of the major surprises in the study was that essentially all the atoms of the protein jump into the correct structure together. The researchers expected that the protein structure would come together in a more piecemeal fashion, with different parts of the structure forming separately, but that was not the case. It is possible that nature evolved such a smooth, highly cooperative folding process to prevent partially folded forms that could get into trouble in the crowded cell membrane. On the other hand, the cooperative folding seen here might not apply to other membrane proteins.
“We need to look at more proteins. The technique developed here may allow us to do just that,” said Dr. Yoon.
The single molecule mechanical manipulation technique could enable detailed folding studies of many other membrane proteins. A major barrier to the study of membrane proteins previously is that the proteins tend to stick together and get tangled up, as computer cords lying at your feet tend to do. With the tweezer technique used in this work, the protein cords are held apart from other cords so they can’t get knotted up. It is hoped that the new approach will open up an important part of the protein universe to scrutiny, including many proteins that become misfolded in disease states.
The title of the research paper is “Mapping the energy landscape for second-stage folding of a single membrane protein” (DOI: 10.1038/nchembio.1939).
Picture: Single-molecule magnetic tweezers that induce mechanical unfolding and refolding of a single membrane protein. Since the force applied is parallel to the biological lipid membrane, the unfolding and refolding processes occur within the membrane.
2015.10.20 View 11029 -
 Establishment of System Metabolic Engineering Strategies
Although conventional petrochemical processes have generated chemicals and materials which have been useful to mankind, they have also triggered a variety of environmental problems including climate change and relied too much on nonrenewable natural resources. To ameliorate this, researchers have actively pursued the development of industrial microbial strains around the globe in order to overproduce industrially useful chemicals and materials from microbes using renewable biomass. This discipline is called metabolic engineering.
Thanks to advances in genetic engineering and our knowledge of cellular metabolism, conventional metabolic engineering efforts have succeeded to a certain extent in developing microbial strains that overproduce bioproducts at an industrial level. However, many metabolic engineering projects launched in academic labs do not reach commercial markets due to a failure to fully integrate industrial bioprocesses.
In response to this, Distinguished Professor Sang Yup Lee and Dr. Hyun Uk Kim, both from the Department of Chemical and Biomolecular Engineering at KAIST, have recently suggested ten general strategies of systems metabolic engineering to successfully develop industrial microbial strains. Systems metabolic engineering differs from conventional metabolic engineering by incorporating traditional metabolic engineering approaches along with tools of other fields, such as systems biology, synthetic biology, and molecular evolution.
The ten strategies of systems metabolic engineering have been featured in Nature Biotechnology released online in October 2015, which is entitled "Systems strategies for developing industrial microbial strains."
The strategies cover economic, state-of-the-art biological techniques and traditional bioprocess aspects. Specifically, they consist of: 1) project design including economic evaluation of a target bioproduct; 2) selection of host strains to be used for overproduction of a bioproduct; 3) metabolic pathway reconstruction for bioproducts that are not naturally produced in the selected host strains; 4) increasing tolerance of a host strain against the bioproduct; 5) removing negative regulatory circuits in the microbial host limiting overproduction of a bioproduct; 6) rerouting intracellular fluxes to optimize cofactor and precursor availability necessary for the bioproduct formation; 7) diagnosing and optimizing metabolic fluxes towards product formation; 8) diagnosis and optimization of microbial culture conditions including carbon sources; 9) system-wide gene manipulation to further increase the host strain's production performance using high-throughput genome-scale engineering and computational tools; and 10) scale-up fermentation of the developed strain and diagnosis for the reproducibility of the strain's production performance.
These ten strategies were articulated with successful examples of the production of L-arginine using Corynebacterium glutamicum, 1,4-butanediol using Escherichia coli, and L-lysine and bio-nylon using C. glutamicum.
Professor Sang Yup Lee said, "At the moment, the chance of commercializing microbial strains developed in academic labs is very low. The strategies of systems metabolic engineering outlined in this analysis can serve as guidelines when developing industrial microbial strains. We hope that these strategies contribute to improving opportunities to commercialize microbial strains developed in academic labs with drastically reduced costs and efforts, and that a large fraction of petroleum-based processes will be replaced with sustainable bioprocesses."
Lee S. Y. & Kim, H. U. Systems Strategies for Developing Industrial Microbial Strains. Nature Biotechnology (2015).
This work was supported by the Technology Development Program to Solve Climate Change on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556) and by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) from the Ministry of Science, ICT and Future Planning (MSIP), Korea, and through the National Research Foundation (NRF) of Korea. This work was also supported by the Novo Nordisk Foundation.
Picture: Concept of the Systems Metabolic Engineering Framework
(a) Three major bioprocess stages (b) Considerations in systems metabolic engineering to optimize the whole bioprocess. List of considerations for the strain development and fermentation contribute to improving microbial strain's production performance (red), whereas those for the separation and purification help in reducing overall operation costs by facilitating the downstream process (blue). Some of the considerations can be repeated in the course of systems metabolic engineering.
2015.10.19 View 12665
Establishment of System Metabolic Engineering Strategies
Although conventional petrochemical processes have generated chemicals and materials which have been useful to mankind, they have also triggered a variety of environmental problems including climate change and relied too much on nonrenewable natural resources. To ameliorate this, researchers have actively pursued the development of industrial microbial strains around the globe in order to overproduce industrially useful chemicals and materials from microbes using renewable biomass. This discipline is called metabolic engineering.
Thanks to advances in genetic engineering and our knowledge of cellular metabolism, conventional metabolic engineering efforts have succeeded to a certain extent in developing microbial strains that overproduce bioproducts at an industrial level. However, many metabolic engineering projects launched in academic labs do not reach commercial markets due to a failure to fully integrate industrial bioprocesses.
In response to this, Distinguished Professor Sang Yup Lee and Dr. Hyun Uk Kim, both from the Department of Chemical and Biomolecular Engineering at KAIST, have recently suggested ten general strategies of systems metabolic engineering to successfully develop industrial microbial strains. Systems metabolic engineering differs from conventional metabolic engineering by incorporating traditional metabolic engineering approaches along with tools of other fields, such as systems biology, synthetic biology, and molecular evolution.
The ten strategies of systems metabolic engineering have been featured in Nature Biotechnology released online in October 2015, which is entitled "Systems strategies for developing industrial microbial strains."
The strategies cover economic, state-of-the-art biological techniques and traditional bioprocess aspects. Specifically, they consist of: 1) project design including economic evaluation of a target bioproduct; 2) selection of host strains to be used for overproduction of a bioproduct; 3) metabolic pathway reconstruction for bioproducts that are not naturally produced in the selected host strains; 4) increasing tolerance of a host strain against the bioproduct; 5) removing negative regulatory circuits in the microbial host limiting overproduction of a bioproduct; 6) rerouting intracellular fluxes to optimize cofactor and precursor availability necessary for the bioproduct formation; 7) diagnosing and optimizing metabolic fluxes towards product formation; 8) diagnosis and optimization of microbial culture conditions including carbon sources; 9) system-wide gene manipulation to further increase the host strain's production performance using high-throughput genome-scale engineering and computational tools; and 10) scale-up fermentation of the developed strain and diagnosis for the reproducibility of the strain's production performance.
These ten strategies were articulated with successful examples of the production of L-arginine using Corynebacterium glutamicum, 1,4-butanediol using Escherichia coli, and L-lysine and bio-nylon using C. glutamicum.
Professor Sang Yup Lee said, "At the moment, the chance of commercializing microbial strains developed in academic labs is very low. The strategies of systems metabolic engineering outlined in this analysis can serve as guidelines when developing industrial microbial strains. We hope that these strategies contribute to improving opportunities to commercialize microbial strains developed in academic labs with drastically reduced costs and efforts, and that a large fraction of petroleum-based processes will be replaced with sustainable bioprocesses."
Lee S. Y. & Kim, H. U. Systems Strategies for Developing Industrial Microbial Strains. Nature Biotechnology (2015).
This work was supported by the Technology Development Program to Solve Climate Change on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556) and by the Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) from the Ministry of Science, ICT and Future Planning (MSIP), Korea, and through the National Research Foundation (NRF) of Korea. This work was also supported by the Novo Nordisk Foundation.
Picture: Concept of the Systems Metabolic Engineering Framework
(a) Three major bioprocess stages (b) Considerations in systems metabolic engineering to optimize the whole bioprocess. List of considerations for the strain development and fermentation contribute to improving microbial strain's production performance (red), whereas those for the separation and purification help in reducing overall operation costs by facilitating the downstream process (blue). Some of the considerations can be repeated in the course of systems metabolic engineering.
2015.10.19 View 12665 -
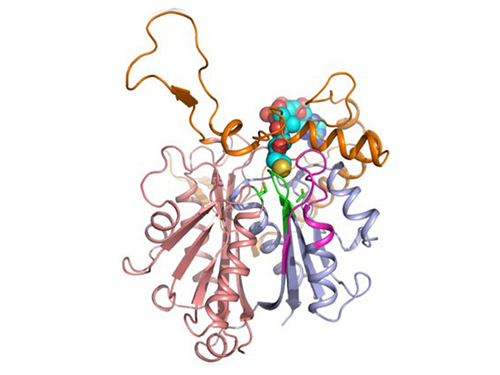 Discovery of Redox-Switch of KEenzyme Involved in N-Butanol Biosynthesis
Research teams at KAIST and Kyungpook National University (KNU) have succeeded in uncovering the redox-switch of thiolase, a key enzyme for n-butanol production in Clostridium acetobutylicum, one of the best known butanol-producing bacteria.
Biological n-butanol production was first reported by Louis Pasteur in 1861, and the bioprocess was industrialized usingClostridium acetobutylicum. The fermentation process by Clostridium strains has been known to be the most efficient one for n-butanol production. Due to growing world-wide issues such as energy security and climate change, the biological production of n-butanol has been receiving much renewed interest. This is because n-butanol possesses much better fuel characteristics compared to ethanol, such as higher energy content (29.2 MJ/L vs 19.6 MJ/L), less corrosiveness, less hygroscopy, and the ease with which it can be blended with gasoline and diesel.
In the paper published in Nature Communications, a broad-scope, online-only, and open access journal issued by the Nature Publishing Group (NPG), on September 22, 2015, Professor Kyung-Jin Kim at the School of Life Sciences, KNU, and Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering, KAIST, have proved that the redox-switch of thiolase plays a role in a regulation of metabolic flux in C. acetobutylicum by using in silico modeling and simulation tools.
The research team has redesigned thiolase with enhanced activity on the basis of the 3D structure of the wild-type enzyme. To reinforce a metabolic flux toward butanol production, the metabolic network of C. acetobutylicum strain was engineered with the redesigned enzyme. The combination of the discovery of 3D enzyme structure and systems metabolic engineering approaches resulted in increased n-butanol production in C. acetobutylicum, which allows the production of this important industrial chemical to be cost competitive.
Professors Kim and Lee said, "We have reported the 3D structure of C. acetobutylicum thiolase-a key enzyme involved in n-butanol biosynthesis, for the first time. Further study will be done to produce butanol more economically on the basis of the 3D structure of C. acetobutylicum thiolase."
This work was published online in Nature Communications on September 22, 2015.
Reference: Kim et al. "Redox-switch regulatory mechanism of thiolase from Clostridium acetobutylicum," Nature Communications
This research was supported by the Technology Development Program to Solve Climate Changes from the Ministry of Education, Science and Technology (MEST), Korea, the National Research Foundation of Korea, and the Advanced Biomass Center through the Global Frontier Research Program of the MEST, Korea.
For further information, contact Dr. Sang Yup Lee, Distinguished Professor, KAIST, Daejeon, Korea (leesy@kaist.ac.kr, +82-42-350-3930); and Dr. Kyung-Jin Kim, Professor, KNU, Daegu, Korea (kkim@knu.ac.kr, +82-53-950-6088).
Figure 1: A redox-switch of thiolase involves in butanol biosynthesis in Clostridium acetobutylicum. Thiolase condenses two acetyl-CoA molecules for initiating four carbon flux towards butanol.
Figure 2: Thiolase catalyzes the condensation reaction of acetyl-CoA to acetoacetyl-CoA. Two catalytic cysteine residues at 88th and 378th are oxidized and formed an intermolecular disulfide bond in an oxidized status, which results in inactivation of the enzyme for n-butanol biosynthesis. The intermolecular disulfide bond is broken enabling the n-butanol biosynthesis, when the environment status is reduced.
2015.09.23 View 12323
Discovery of Redox-Switch of KEenzyme Involved in N-Butanol Biosynthesis
Research teams at KAIST and Kyungpook National University (KNU) have succeeded in uncovering the redox-switch of thiolase, a key enzyme for n-butanol production in Clostridium acetobutylicum, one of the best known butanol-producing bacteria.
Biological n-butanol production was first reported by Louis Pasteur in 1861, and the bioprocess was industrialized usingClostridium acetobutylicum. The fermentation process by Clostridium strains has been known to be the most efficient one for n-butanol production. Due to growing world-wide issues such as energy security and climate change, the biological production of n-butanol has been receiving much renewed interest. This is because n-butanol possesses much better fuel characteristics compared to ethanol, such as higher energy content (29.2 MJ/L vs 19.6 MJ/L), less corrosiveness, less hygroscopy, and the ease with which it can be blended with gasoline and diesel.
In the paper published in Nature Communications, a broad-scope, online-only, and open access journal issued by the Nature Publishing Group (NPG), on September 22, 2015, Professor Kyung-Jin Kim at the School of Life Sciences, KNU, and Distinguished Professor Sang Yup Lee at the Department of Chemical and Biomolecular Engineering, KAIST, have proved that the redox-switch of thiolase plays a role in a regulation of metabolic flux in C. acetobutylicum by using in silico modeling and simulation tools.
The research team has redesigned thiolase with enhanced activity on the basis of the 3D structure of the wild-type enzyme. To reinforce a metabolic flux toward butanol production, the metabolic network of C. acetobutylicum strain was engineered with the redesigned enzyme. The combination of the discovery of 3D enzyme structure and systems metabolic engineering approaches resulted in increased n-butanol production in C. acetobutylicum, which allows the production of this important industrial chemical to be cost competitive.
Professors Kim and Lee said, "We have reported the 3D structure of C. acetobutylicum thiolase-a key enzyme involved in n-butanol biosynthesis, for the first time. Further study will be done to produce butanol more economically on the basis of the 3D structure of C. acetobutylicum thiolase."
This work was published online in Nature Communications on September 22, 2015.
Reference: Kim et al. "Redox-switch regulatory mechanism of thiolase from Clostridium acetobutylicum," Nature Communications
This research was supported by the Technology Development Program to Solve Climate Changes from the Ministry of Education, Science and Technology (MEST), Korea, the National Research Foundation of Korea, and the Advanced Biomass Center through the Global Frontier Research Program of the MEST, Korea.
For further information, contact Dr. Sang Yup Lee, Distinguished Professor, KAIST, Daejeon, Korea (leesy@kaist.ac.kr, +82-42-350-3930); and Dr. Kyung-Jin Kim, Professor, KNU, Daegu, Korea (kkim@knu.ac.kr, +82-53-950-6088).
Figure 1: A redox-switch of thiolase involves in butanol biosynthesis in Clostridium acetobutylicum. Thiolase condenses two acetyl-CoA molecules for initiating four carbon flux towards butanol.
Figure 2: Thiolase catalyzes the condensation reaction of acetyl-CoA to acetoacetyl-CoA. Two catalytic cysteine residues at 88th and 378th are oxidized and formed an intermolecular disulfide bond in an oxidized status, which results in inactivation of the enzyme for n-butanol biosynthesis. The intermolecular disulfide bond is broken enabling the n-butanol biosynthesis, when the environment status is reduced.
2015.09.23 View 12323 -
 Nature Biotechnology Nominates Sang Yup Lee of KAIST for Top 20 Translational Researchers of 2014
Nature Biotechnology, recognized as the most prestigious journal in the field of biotechnology, has released today its list of the Top 20 Translational Researchers of 2014. Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST (Korea Advanced Institute of Science and Technology) ranked seventh in the list. He is the only Asian researcher listed.
The journal, in partnership with IP Checkups, a patent analytics firm, presents an annual ranking of researchers based on their paper and patent output. The list includes, among others, each researcher’s most-cited patent in the past five years and their H index, a measurement to evaluate the impact of a researcher’s published work utilizing citation analysis. (More details can be found at http://www.nature.com/bioent/2015/150801/full/bioe.2015.9.html.)
American institutions made up the majority of the list, with 18 universities and research institutes, and the remainder was filled by KAIST in Korea and the Commonwealth Scientific and Industrial Research Organization (CSIRO) in Australia.
Globally known as a leading researcher in systems metabolic engineering, Professor Lee has published more than 500 journal papers and 580 patents. He has received many awards, including the Citation Classic Award, Elmer Gaden Award, Merck Metabolic Engineering Award, ACS Marvin Johnson Award, SIMB Charles Thom Award, POSCO TJ Park Prize, Amgen Biochemical Engineering Award, and the Ho Am Prize in Engineering.
2015.08.27 View 13353
Nature Biotechnology Nominates Sang Yup Lee of KAIST for Top 20 Translational Researchers of 2014
Nature Biotechnology, recognized as the most prestigious journal in the field of biotechnology, has released today its list of the Top 20 Translational Researchers of 2014. Distinguished Professor Sang Yup Lee of the Department of Chemical and Biomolecular Engineering at KAIST (Korea Advanced Institute of Science and Technology) ranked seventh in the list. He is the only Asian researcher listed.
The journal, in partnership with IP Checkups, a patent analytics firm, presents an annual ranking of researchers based on their paper and patent output. The list includes, among others, each researcher’s most-cited patent in the past five years and their H index, a measurement to evaluate the impact of a researcher’s published work utilizing citation analysis. (More details can be found at http://www.nature.com/bioent/2015/150801/full/bioe.2015.9.html.)
American institutions made up the majority of the list, with 18 universities and research institutes, and the remainder was filled by KAIST in Korea and the Commonwealth Scientific and Industrial Research Organization (CSIRO) in Australia.
Globally known as a leading researcher in systems metabolic engineering, Professor Lee has published more than 500 journal papers and 580 patents. He has received many awards, including the Citation Classic Award, Elmer Gaden Award, Merck Metabolic Engineering Award, ACS Marvin Johnson Award, SIMB Charles Thom Award, POSCO TJ Park Prize, Amgen Biochemical Engineering Award, and the Ho Am Prize in Engineering.
2015.08.27 View 13353