research
-
 High Resolution 3D Blood Vessel Endoscope System Developed
Professor Wangyeol Oh of KAIST’s Mechanical Engineering Department has succeeded in developing an optical imaging endoscope system that employs an imaging velocity, which is up to 3.5 times faster than the previous systems. Furthermore, he has utilized this endoscope to acquire the world’s first high-resolution 3D images of the insides of in vivo blood vessel.
Professor Oh’s work is Korea’s first development of blood vessel endoscope system, possessing an imaging speed, resolution, imaging quality, and image-capture area. The system can also simultaneously perform a functional imaging, such as polarized imaging, which is advantageous for identifying the vulnerability of the blood vessel walls.
The Endoscopic Optical Coherence Tomography (OCT) System provides the highest resolution that is used to diagnose cardiovascular diseases, represented mainly by myocardial infarction.
However, the previous system was not fast enough to take images inside of the vessels, and therefore it was often impossible to accurately identify and analyze the vessel condition. To achieve an in vivo blood vessel optical imaging in clinical trials, the endoscope needed to be inserted, after which a clear liquid flows instantly, and pictures can be taken in only a few seconds.
The KAIST research team proposed a solution for such problem by developing a high-speed, high-resolution optical tomographic imaging system, a flexible endoscope with a diameter of 0.8 mm, as well as a device that can scan the imaging light within the blood vessels at high speed. Then, these devices were combined to visualize the internal structure of the vessel wall.
Using the developed system, the researchers were able to obtain high-resolution images of about 7 cm blood vessels of a rabbit’s aorta, which is similar size to human’s coronary arteries. The tomography scan took only 5.8 seconds, at a speed of 350 scans per second in all three directions with a resolution of 10~35㎛.
If the images are taken every 200 ㎛, like the currently available commercial vascular imaging endoscopes, a 7cm length vessel can be imaged in only one second.
Professor Wangyeol Oh said, “Our newly developed blood vessel endoscope system was tested by imaging a live animal’s blood vessels, which is similar to human blood vessels. The result was very successful.”
“Collaborating closely with hospitals, we are preparing to produce the imaging of an animal’s coronary arteries, which is similar in size to the human heart,” commented Professor Oh on the future clinical application and commercialization of the endoscope system. He added, “After such procedures, the technique can be applied in clinical patients within a few years.”
Professor Oh’s research was supported by the National Research Foundation of Korea and the Global Frontier Project by the Korean government. The research results were published in the 2014 January’s edition of Biomedical Optics Express.
Figure 1: End portion of optical endoscope (upper left)
Figure 2: High-speed optical scanning unit of the endoscope (top right)
Figure 3: High-resolution images of the inside of in vivo animal blood vessels (in the direction of vascular circumference and length)
Figure 4: High-resolution images of the inside of in vivo animal blood vessels (in the direction of the vein depth)
2014.03.25 View 12948
High Resolution 3D Blood Vessel Endoscope System Developed
Professor Wangyeol Oh of KAIST’s Mechanical Engineering Department has succeeded in developing an optical imaging endoscope system that employs an imaging velocity, which is up to 3.5 times faster than the previous systems. Furthermore, he has utilized this endoscope to acquire the world’s first high-resolution 3D images of the insides of in vivo blood vessel.
Professor Oh’s work is Korea’s first development of blood vessel endoscope system, possessing an imaging speed, resolution, imaging quality, and image-capture area. The system can also simultaneously perform a functional imaging, such as polarized imaging, which is advantageous for identifying the vulnerability of the blood vessel walls.
The Endoscopic Optical Coherence Tomography (OCT) System provides the highest resolution that is used to diagnose cardiovascular diseases, represented mainly by myocardial infarction.
However, the previous system was not fast enough to take images inside of the vessels, and therefore it was often impossible to accurately identify and analyze the vessel condition. To achieve an in vivo blood vessel optical imaging in clinical trials, the endoscope needed to be inserted, after which a clear liquid flows instantly, and pictures can be taken in only a few seconds.
The KAIST research team proposed a solution for such problem by developing a high-speed, high-resolution optical tomographic imaging system, a flexible endoscope with a diameter of 0.8 mm, as well as a device that can scan the imaging light within the blood vessels at high speed. Then, these devices were combined to visualize the internal structure of the vessel wall.
Using the developed system, the researchers were able to obtain high-resolution images of about 7 cm blood vessels of a rabbit’s aorta, which is similar size to human’s coronary arteries. The tomography scan took only 5.8 seconds, at a speed of 350 scans per second in all three directions with a resolution of 10~35㎛.
If the images are taken every 200 ㎛, like the currently available commercial vascular imaging endoscopes, a 7cm length vessel can be imaged in only one second.
Professor Wangyeol Oh said, “Our newly developed blood vessel endoscope system was tested by imaging a live animal’s blood vessels, which is similar to human blood vessels. The result was very successful.”
“Collaborating closely with hospitals, we are preparing to produce the imaging of an animal’s coronary arteries, which is similar in size to the human heart,” commented Professor Oh on the future clinical application and commercialization of the endoscope system. He added, “After such procedures, the technique can be applied in clinical patients within a few years.”
Professor Oh’s research was supported by the National Research Foundation of Korea and the Global Frontier Project by the Korean government. The research results were published in the 2014 January’s edition of Biomedical Optics Express.
Figure 1: End portion of optical endoscope (upper left)
Figure 2: High-speed optical scanning unit of the endoscope (top right)
Figure 3: High-resolution images of the inside of in vivo animal blood vessels (in the direction of vascular circumference and length)
Figure 4: High-resolution images of the inside of in vivo animal blood vessels (in the direction of the vein depth)
2014.03.25 View 12948 -
 Box-shaped Pressure Vessel for LNG Developed by KAIST Research Team
Earlier today, Korean researchers successfully showcased the installation and operation of a box-shaped, high-pressure tank for the storage of liquefied natural gas in Pohang, Republic of Korea. The development was the first of its kind in the world. Pressure vessels have many applications and are widely used within the petrochemical, energy, and other industrial sectors where the transport and storage of many types of pressurized gases and fluids are essential. Pressure vessels must be designed, manufactured, installed, and operated strictly in accordance with the appropriate codes and standards since they can, in cases of leak or rupture, pose considerable health and safety hazards. Pressure vessels are normally designed in the form of a cylindrical or spherical tank. These shapes are, in principle, highly efficient in withstanding internal pressure, but rather inefficient in terms of space utilization. The tanks fit very poorly within a typically prismatic-shaped room. They cannot be packed closely together, so they do not efficiently utilize the overall space. Moreover, cylindrical or spherical tanks are not easily scalable to very large sizes because the wall thickness of the tank must increase proportionally to its overall radius. Therefore, a large pressure vessel unavoidably will have very thick walls, which are difficult and expensive to manufacture, requiring a great amount of thick-walled steel to be rolled, forged, and welded together. KAIST researchers, sponsored by POSCO, a multinational steel-making company based in Pohang, Republic of Korea, have taken a turnabout approach to construct a pressure vessel that is neither cylindrical nor spherical. Professors Pål G. Bergan and Daejun Chang and of Ocean Systems Engineering at KAIST developed a box-type, large size pressure vessel for the storage and transportation of liquids such as liquefied petroleum gas (LPG), compressed natural gas (CNG), or liquefied natural gas (LNG). The box-shaped pressure vessel has an internal, load-carrying lattice-type structure. The lattice pattern is modular in all three spatial directions, thereby effectively anchoring and balancing pressure forces on the external walls of the vessel. The modular lattice can easily be adapted to prescribed pressure levels as the overall volumetric dimensions are directly linked to the number of repetitive modules. A giant prismatic pressure vessel with a size of 20,000 m3 and a design pressure of 10 atmospheres (10 barg) can be built simply by scaling up a smaller size pressure vessel. It is interesting to note that the thickness of steel walls remains unchanged and that the weight of steel per unit storage volume goes down as the vessel size increases. Professor Chang explained the benefit of a prismatic or box-shaped pressure vessel.“If we use cylindrical pressure vessels to supply LNG fuel for a large container ship, for example, many fuel tanks will be needed. Those tanks will take up large and valuable space onboard because the cylinders have to be lined up. In our case, however, much less space is needed. The operation of a ship becomes simpler with one fuel tank rather than with many. Furthermore, our box-type pressure vessel can be designed with dimensions that precisely fit a ship. For a container ship, there may be room for a substantially higher number of containers to be loaded than when using cylindrical vessels. In a case study on a 13,000 TEU container ship, the value of the increased transport capacity tuned out USD 8.4 million for one year of operation for one ship.”The manufacturing cost of a pressure vessel has been reduced as well. Several types of special steel for cryogenic (low temperature) applications have been investigated in design and analysis studies, and this includes a new type of high-manganese steel that is being developed by POSCO. Regardless of materials, in any instance of large pressure vessels, the new lattice tank technology can offer significant savings of combined capital and operational costs. Professor Bergan was also upbeat regarding the impact of the KAIST technology innovation. “Our box-type pressure vessel represents ground-breaking research. This innovative technology will dramatically change the rules of the game for industry concerning production, transportation, and storage of fluids under high pressure and at low temperatures.”The showcased prismatic pressure vessel was a scale-down model with a volume size of 80 m3 and design pressure of 10 atmospheres. The vessel complies with the American Society of Mechanical Engineers (ASME) Boiler and Pressure Vessel Code (BPVC), the international standard for the appropriateness of design, fabrication, and inspection of boilers and pressure vessels. It passed the 15 pressure testing in January 2014 and received an accreditation from the ASME BPVC (ASME U2 Stamp). KAIST’s prismatic pressure vessel will be presented and displayed at Gastech 2014, the largest global conference and exhibition in the natural gas, LNG, and hydrocarbons industry. This event will take place on March 24-27 at KINTEX in Ilsan, Republic of Korea. Youtube: http://www.youtube.com/watch?v=woJwc5zisxk&list=TLGOLcI7L6_YYTn0lImPqNyeppQWRXqUt5Picture 1: The prototype of a prismatic pressure vesselPicture 2: A lattice pattern that is lined inside a prismatic pressure tankPicture 3: Above is a container ship having a box-shaped pressure vessel as a fuel tank, and below are traditional cylindrical fuel tanks.
2014.03.25 View 15136
Box-shaped Pressure Vessel for LNG Developed by KAIST Research Team
Earlier today, Korean researchers successfully showcased the installation and operation of a box-shaped, high-pressure tank for the storage of liquefied natural gas in Pohang, Republic of Korea. The development was the first of its kind in the world. Pressure vessels have many applications and are widely used within the petrochemical, energy, and other industrial sectors where the transport and storage of many types of pressurized gases and fluids are essential. Pressure vessels must be designed, manufactured, installed, and operated strictly in accordance with the appropriate codes and standards since they can, in cases of leak or rupture, pose considerable health and safety hazards. Pressure vessels are normally designed in the form of a cylindrical or spherical tank. These shapes are, in principle, highly efficient in withstanding internal pressure, but rather inefficient in terms of space utilization. The tanks fit very poorly within a typically prismatic-shaped room. They cannot be packed closely together, so they do not efficiently utilize the overall space. Moreover, cylindrical or spherical tanks are not easily scalable to very large sizes because the wall thickness of the tank must increase proportionally to its overall radius. Therefore, a large pressure vessel unavoidably will have very thick walls, which are difficult and expensive to manufacture, requiring a great amount of thick-walled steel to be rolled, forged, and welded together. KAIST researchers, sponsored by POSCO, a multinational steel-making company based in Pohang, Republic of Korea, have taken a turnabout approach to construct a pressure vessel that is neither cylindrical nor spherical. Professors Pål G. Bergan and Daejun Chang and of Ocean Systems Engineering at KAIST developed a box-type, large size pressure vessel for the storage and transportation of liquids such as liquefied petroleum gas (LPG), compressed natural gas (CNG), or liquefied natural gas (LNG). The box-shaped pressure vessel has an internal, load-carrying lattice-type structure. The lattice pattern is modular in all three spatial directions, thereby effectively anchoring and balancing pressure forces on the external walls of the vessel. The modular lattice can easily be adapted to prescribed pressure levels as the overall volumetric dimensions are directly linked to the number of repetitive modules. A giant prismatic pressure vessel with a size of 20,000 m3 and a design pressure of 10 atmospheres (10 barg) can be built simply by scaling up a smaller size pressure vessel. It is interesting to note that the thickness of steel walls remains unchanged and that the weight of steel per unit storage volume goes down as the vessel size increases. Professor Chang explained the benefit of a prismatic or box-shaped pressure vessel.“If we use cylindrical pressure vessels to supply LNG fuel for a large container ship, for example, many fuel tanks will be needed. Those tanks will take up large and valuable space onboard because the cylinders have to be lined up. In our case, however, much less space is needed. The operation of a ship becomes simpler with one fuel tank rather than with many. Furthermore, our box-type pressure vessel can be designed with dimensions that precisely fit a ship. For a container ship, there may be room for a substantially higher number of containers to be loaded than when using cylindrical vessels. In a case study on a 13,000 TEU container ship, the value of the increased transport capacity tuned out USD 8.4 million for one year of operation for one ship.”The manufacturing cost of a pressure vessel has been reduced as well. Several types of special steel for cryogenic (low temperature) applications have been investigated in design and analysis studies, and this includes a new type of high-manganese steel that is being developed by POSCO. Regardless of materials, in any instance of large pressure vessels, the new lattice tank technology can offer significant savings of combined capital and operational costs. Professor Bergan was also upbeat regarding the impact of the KAIST technology innovation. “Our box-type pressure vessel represents ground-breaking research. This innovative technology will dramatically change the rules of the game for industry concerning production, transportation, and storage of fluids under high pressure and at low temperatures.”The showcased prismatic pressure vessel was a scale-down model with a volume size of 80 m3 and design pressure of 10 atmospheres. The vessel complies with the American Society of Mechanical Engineers (ASME) Boiler and Pressure Vessel Code (BPVC), the international standard for the appropriateness of design, fabrication, and inspection of boilers and pressure vessels. It passed the 15 pressure testing in January 2014 and received an accreditation from the ASME BPVC (ASME U2 Stamp). KAIST’s prismatic pressure vessel will be presented and displayed at Gastech 2014, the largest global conference and exhibition in the natural gas, LNG, and hydrocarbons industry. This event will take place on March 24-27 at KINTEX in Ilsan, Republic of Korea. Youtube: http://www.youtube.com/watch?v=woJwc5zisxk&list=TLGOLcI7L6_YYTn0lImPqNyeppQWRXqUt5Picture 1: The prototype of a prismatic pressure vesselPicture 2: A lattice pattern that is lined inside a prismatic pressure tankPicture 3: Above is a container ship having a box-shaped pressure vessel as a fuel tank, and below are traditional cylindrical fuel tanks.
2014.03.25 View 15136 -
 Spillover Phenomenon Identified Using Model Catalyst System
Researchers at KAIST have identified spillover phenomenon, which has remained controversial since its discovery in the early 1960s.
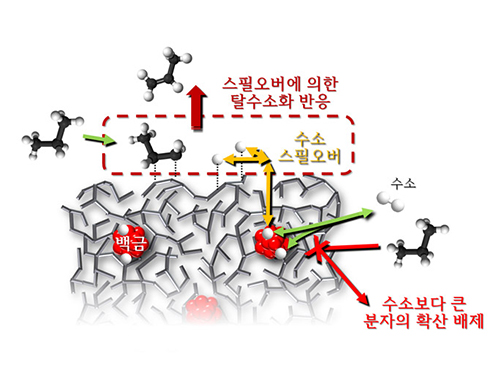
KAIST Department of Chemical and Biomolecular Engineering’s Professor Min-Gi Choi and his team has explained the "spillover phenomenon," using their own model catalyst system where platinum is selectively located within the amorphous aluminosilicate.
The research results were published on the 25th February online edition of Nature Communications.
Spillover refers to a phenomenon that occurs when hydrogen atoms that have been activated on the surface of metals, such as platinum, move to the surface of the catalyst. It was predicted that this phenomenon can be used to design a catalyst with high activity and stability, and thus has been actively studied over the last 50 years.
However, many cases of the known catalysts involved competing reactions on the exposed metal surface, which made it impossible to directly identify the presence and formation mechanism of spillover.
The catalysts developed by the researchers at KAIST used platinum nanoparticles covered with aluminosilicate. This only allowed the hydrogen molecules to pass through and has effectively blocked the competing reactions, enabling the research team to study the spillover phenomenon.
Through various catalyst structure and reactivity analysis, as well as computer modeling, the team has discovered that Brönsted acid sites present on the aluminosilicate plays a crucial role in spillover phenomenon.
In addition, the spillover-based hydrogenation catalyst proposed by the research team showed very high hydrogenation and dehydrogenation activity. The ability of the catalyst to significantly inhibit unwanted hydrogenolysis reaction during the petrochemical processes also suggested a large industrial potential.
Professor Min-Gi Choi said, “This particular catalyst, which can trigger the reaction only by spillover phenomenon, can be properly designed to exceed the capacity of the conventional metal catalysts. The future goal is to make a catalyst with much higher activity and selectivity.”
The research was conducted through funds subsidized by SK Innovation and Ministry of Science, ICT and Future Planning.
The senior research fellow of SK Innovation Seung-Hun Oh said, “SK Innovation will continue to develop a new commercial catalyst based on the technology from this research.”
Juh-Wan Lim and Hye-Yeong Shin led the research as joint first authors under supervision of Professor Min-Gi Choi and computer modeling works were conducted by KAIST EEWS (environment, energy, water, and sustainability) graduate school’s Professor Hyeong-Jun Kim.
2014.03.03 View 11388
Spillover Phenomenon Identified Using Model Catalyst System
Researchers at KAIST have identified spillover phenomenon, which has remained controversial since its discovery in the early 1960s.
KAIST Department of Chemical and Biomolecular Engineering’s Professor Min-Gi Choi and his team has explained the "spillover phenomenon," using their own model catalyst system where platinum is selectively located within the amorphous aluminosilicate.
The research results were published on the 25th February online edition of Nature Communications.
Spillover refers to a phenomenon that occurs when hydrogen atoms that have been activated on the surface of metals, such as platinum, move to the surface of the catalyst. It was predicted that this phenomenon can be used to design a catalyst with high activity and stability, and thus has been actively studied over the last 50 years.
However, many cases of the known catalysts involved competing reactions on the exposed metal surface, which made it impossible to directly identify the presence and formation mechanism of spillover.
The catalysts developed by the researchers at KAIST used platinum nanoparticles covered with aluminosilicate. This only allowed the hydrogen molecules to pass through and has effectively blocked the competing reactions, enabling the research team to study the spillover phenomenon.
Through various catalyst structure and reactivity analysis, as well as computer modeling, the team has discovered that Brönsted acid sites present on the aluminosilicate plays a crucial role in spillover phenomenon.
In addition, the spillover-based hydrogenation catalyst proposed by the research team showed very high hydrogenation and dehydrogenation activity. The ability of the catalyst to significantly inhibit unwanted hydrogenolysis reaction during the petrochemical processes also suggested a large industrial potential.
Professor Min-Gi Choi said, “This particular catalyst, which can trigger the reaction only by spillover phenomenon, can be properly designed to exceed the capacity of the conventional metal catalysts. The future goal is to make a catalyst with much higher activity and selectivity.”
The research was conducted through funds subsidized by SK Innovation and Ministry of Science, ICT and Future Planning.
The senior research fellow of SK Innovation Seung-Hun Oh said, “SK Innovation will continue to develop a new commercial catalyst based on the technology from this research.”
Juh-Wan Lim and Hye-Yeong Shin led the research as joint first authors under supervision of Professor Min-Gi Choi and computer modeling works were conducted by KAIST EEWS (environment, energy, water, and sustainability) graduate school’s Professor Hyeong-Jun Kim.
2014.03.03 View 11388 -
 KAIST developed an extremely low-powered, high-performance head-mounted display embedding an augmented reality chip
Walking around the streets searching for a place to eat will be no hassle when a head-mounted display (HMD) becomes affordable and ubiquitous. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) developed K-Glass, a wearable, hands-free HMD that enables users to find restaurants while checking out their menus. If the user of K-Glass walks up to a restaurant and looks at the name of the restaurant, today’s menu and a 3D image of food pop up. The Glass can even show the number of tables available inside the restaurant. K-Glass makes this possible because of its built-in augmented reality (AR) processor. Unlike virtual reality which replaces the real world with a computer-simulated environment, AR incorporates digital data generated by the computer into the reality of a user. With the computer-made sensory inputs such as sound, video, graphics or GPS data, the user’s real and physical world becomes live and interactive. Augmentation takes place in real-time and in semantic context with surrounding environments, such as a menu list overlain on the signboard of a restaurant when the user passes by it, not an airplane flight schedule, which is irrelevant information, displayed. Most commonly, location-based or computer-vision services are used in order to generate AR effects. Location-based services activate motion sensors to identify the user’s surroundings, whereas computer-vision uses algorithms such as facial, pattern, and optical character recognition, or object and motion tracking to distinguish images and objects. Many of the current HMDs deliver augmented reality experiences employing location-based services by scanning the markers or barcodes printed on the back of objects. The AR system tracks the codes or markers to identify objects and then align them with virtual reality. However, this AR algorithm is difficult to use for the objects or spaces which do not have barcodes, QR codes, or markers, particularly those in outdoor environments and thus cannot be recognized. To solve this problem, Hoi-Jun Yoo, Professor of Electrical Engineering at KAIST and his team developed, for the first time in the world, an AR chip that works just like human vision. This processor is based on the Visual Attention Model (VAM) that duplicates the ability of human brain to process visual data. VAM, almost unconsciously or automatically, disentangles the most salient and relevant information about the environment in which human vision operates, thereby eliminating unnecessary data unless they must be processed. In return, the processor can dramatically speed up the computation of complex AR algorithms. The AR processor has a data processing network similar to that of a human brain’s central nervous system. When the human brain perceives visual data, different sets of neurons, all connected, work concurrently on each fragment of a decision-making process; one group’s work is relayed to other group of neurons for the next round of the process, which continues until a set of decider neurons determines the character of the data. Likewise, the artificial neural network allows parallel data processing, alleviating data congestion and reducing power consumption significantly. KAIST’s AR processor, which is produced using the 65 nm (nanometers) manufacturing process with the area of 32 mm2, delivers 1.22 TOPS (tera-operations per second) peak performance when running at 250 MHz and consumes 778 miliWatts on a 1.2V power supply. The ultra-low power processor shows 1.57 TOPS/W high efficiency rate of energy consumption under the real-time operation of 30fps/720p video camera, a 76% improvement in power conservation over other devices. The HMDs, available on the market including the Project Glass whose battery lasts only for two hours, have revealed so far poor performance. Professor Yoo said, “Our processor can work for long hours without sacrificing K-Glass’s high performance, an ideal mobile gadget or wearable computer, which users can wear for almost the whole day.” He further commented:“HMDs will become the next mobile device, eventually taking over smartphones. Their markets have been growing fast, and it’s really a matter of time before mobile users will eventually embrace an optical see-through HMD as part of their daily use. Through augmented reality, we will have richer, deeper, and more powerful reality in all aspects of our life from education, business, and entertainment to art and culture.” The KAIST team presented a research paper at the International Solid-State Circuits Conference (ISSCC) held on February 9-13, 2014 in San Francisco, CA, which is entitled “1.22TOPS and 1.52mW/MHz Augmented Reality Multi-Core Processor with Neural Network NoC for HMD Applications.”Youtube Link: http://www.youtube.com/watch?v=wSqY30FOu2s&feature=c4-overview&list=UUirZA3OFhxP4YFreIJkTtXw
2014.02.20 View 18253
KAIST developed an extremely low-powered, high-performance head-mounted display embedding an augmented reality chip
Walking around the streets searching for a place to eat will be no hassle when a head-mounted display (HMD) becomes affordable and ubiquitous. Researchers at the Korea Advanced Institute of Science and Technology (KAIST) developed K-Glass, a wearable, hands-free HMD that enables users to find restaurants while checking out their menus. If the user of K-Glass walks up to a restaurant and looks at the name of the restaurant, today’s menu and a 3D image of food pop up. The Glass can even show the number of tables available inside the restaurant. K-Glass makes this possible because of its built-in augmented reality (AR) processor. Unlike virtual reality which replaces the real world with a computer-simulated environment, AR incorporates digital data generated by the computer into the reality of a user. With the computer-made sensory inputs such as sound, video, graphics or GPS data, the user’s real and physical world becomes live and interactive. Augmentation takes place in real-time and in semantic context with surrounding environments, such as a menu list overlain on the signboard of a restaurant when the user passes by it, not an airplane flight schedule, which is irrelevant information, displayed. Most commonly, location-based or computer-vision services are used in order to generate AR effects. Location-based services activate motion sensors to identify the user’s surroundings, whereas computer-vision uses algorithms such as facial, pattern, and optical character recognition, or object and motion tracking to distinguish images and objects. Many of the current HMDs deliver augmented reality experiences employing location-based services by scanning the markers or barcodes printed on the back of objects. The AR system tracks the codes or markers to identify objects and then align them with virtual reality. However, this AR algorithm is difficult to use for the objects or spaces which do not have barcodes, QR codes, or markers, particularly those in outdoor environments and thus cannot be recognized. To solve this problem, Hoi-Jun Yoo, Professor of Electrical Engineering at KAIST and his team developed, for the first time in the world, an AR chip that works just like human vision. This processor is based on the Visual Attention Model (VAM) that duplicates the ability of human brain to process visual data. VAM, almost unconsciously or automatically, disentangles the most salient and relevant information about the environment in which human vision operates, thereby eliminating unnecessary data unless they must be processed. In return, the processor can dramatically speed up the computation of complex AR algorithms. The AR processor has a data processing network similar to that of a human brain’s central nervous system. When the human brain perceives visual data, different sets of neurons, all connected, work concurrently on each fragment of a decision-making process; one group’s work is relayed to other group of neurons for the next round of the process, which continues until a set of decider neurons determines the character of the data. Likewise, the artificial neural network allows parallel data processing, alleviating data congestion and reducing power consumption significantly. KAIST’s AR processor, which is produced using the 65 nm (nanometers) manufacturing process with the area of 32 mm2, delivers 1.22 TOPS (tera-operations per second) peak performance when running at 250 MHz and consumes 778 miliWatts on a 1.2V power supply. The ultra-low power processor shows 1.57 TOPS/W high efficiency rate of energy consumption under the real-time operation of 30fps/720p video camera, a 76% improvement in power conservation over other devices. The HMDs, available on the market including the Project Glass whose battery lasts only for two hours, have revealed so far poor performance. Professor Yoo said, “Our processor can work for long hours without sacrificing K-Glass’s high performance, an ideal mobile gadget or wearable computer, which users can wear for almost the whole day.” He further commented:“HMDs will become the next mobile device, eventually taking over smartphones. Their markets have been growing fast, and it’s really a matter of time before mobile users will eventually embrace an optical see-through HMD as part of their daily use. Through augmented reality, we will have richer, deeper, and more powerful reality in all aspects of our life from education, business, and entertainment to art and culture.” The KAIST team presented a research paper at the International Solid-State Circuits Conference (ISSCC) held on February 9-13, 2014 in San Francisco, CA, which is entitled “1.22TOPS and 1.52mW/MHz Augmented Reality Multi-Core Processor with Neural Network NoC for HMD Applications.”Youtube Link: http://www.youtube.com/watch?v=wSqY30FOu2s&feature=c4-overview&list=UUirZA3OFhxP4YFreIJkTtXw
2014.02.20 View 18253 -
 Is it possible to identify rumors on SNS?
Rumors
sporadically spread with people with fewer followers in the centerResearched
over 100 rumors in the US from 2006 to 2009 Is it
possible to filter information on SNS such as Twitter and Facebook? A research
team led by Professor Mee-Young Cha from the Department of Cultural Technology
Graduate School at KAIST, Professor Kyo-Min Jung of Seoul National University,
Doctor Wei Chen and Yajun Wang of Microsoft Asia, has developed a technology
that can accurately filter out information on Twitter to 90% accuracy. The
research not only deduced a new mathematical model, network structure, and
linguistic characteristics on rumors from SNS data, but is also expected to
enhance the effort to make secure technology to regulate Internet rumors. The team
analysed the characteristics of rumors in over 100 widespread cases in the US
from 2006 to 2009 on Twitter. The team gathered data, which included a range of
areas such as politics, IT, health and celebrity gossips, and their analysis
could identify rumors to 90% accuracy. The filtering was more accurate in
rumors that included slanders or insults. The
research team identified three characteristics of the spread of rumors. Firstly,
rumors spread continuously. Normal news spreads widely once and is mentioned
rarely again on media, but rumors tend to continue for years. Secondly,
rumors spread through sporadic participation of random users with no
connections. Rumors start from people with fewer followers and spread to the
more popular. This phenomenon is often observed in rumors concerning
celebrities or politicians. Lastly,
rumors have unique linguistic characteristics. Rumors frequently include words
(such as “it may be true,” “although not certain, I think,” “although I cannot
fully remember”) related to psychological processes that question, deny, or
infer the reliability of the information. Professor
Cha said, “This research deduced not only a statistical and mathematical model
but also is an integrated research on social psychological theory on the
characteristics of rumors that attract great attention from the society based
on ample data.” The results
were made public in IEEE International Conference on Data Mining last December
in Texas, USA.
2014.02.03 View 11537
Is it possible to identify rumors on SNS?
Rumors
sporadically spread with people with fewer followers in the centerResearched
over 100 rumors in the US from 2006 to 2009 Is it
possible to filter information on SNS such as Twitter and Facebook? A research
team led by Professor Mee-Young Cha from the Department of Cultural Technology
Graduate School at KAIST, Professor Kyo-Min Jung of Seoul National University,
Doctor Wei Chen and Yajun Wang of Microsoft Asia, has developed a technology
that can accurately filter out information on Twitter to 90% accuracy. The
research not only deduced a new mathematical model, network structure, and
linguistic characteristics on rumors from SNS data, but is also expected to
enhance the effort to make secure technology to regulate Internet rumors. The team
analysed the characteristics of rumors in over 100 widespread cases in the US
from 2006 to 2009 on Twitter. The team gathered data, which included a range of
areas such as politics, IT, health and celebrity gossips, and their analysis
could identify rumors to 90% accuracy. The filtering was more accurate in
rumors that included slanders or insults. The
research team identified three characteristics of the spread of rumors. Firstly,
rumors spread continuously. Normal news spreads widely once and is mentioned
rarely again on media, but rumors tend to continue for years. Secondly,
rumors spread through sporadic participation of random users with no
connections. Rumors start from people with fewer followers and spread to the
more popular. This phenomenon is often observed in rumors concerning
celebrities or politicians. Lastly,
rumors have unique linguistic characteristics. Rumors frequently include words
(such as “it may be true,” “although not certain, I think,” “although I cannot
fully remember”) related to psychological processes that question, deny, or
infer the reliability of the information. Professor
Cha said, “This research deduced not only a statistical and mathematical model
but also is an integrated research on social psychological theory on the
characteristics of rumors that attract great attention from the society based
on ample data.” The results
were made public in IEEE International Conference on Data Mining last December
in Texas, USA.
2014.02.03 View 11537 -
 A Molecular Switch Controlling Self-Assembly of Protein Nanotubes Discovered
International collaborative research among South Korea, United States, and Israel research institutionsThe key to the treatment of cancer and brain disease mechanism The molecular switch that controls the self-assembly structure of the protein nanotubes, which plays crucial role in cell division and intracellular transport of materials, has been discovered. KAIST Bio and Brain Engineering Department’s Professor Myeong-Cheol Choi and Professor Chae-Yeon Song conducted the research, in collaboration with the University of California in Santa Barbara, U.S., and Hebrew University in Israel. The findings of the research were published in Nature Materials on the 19th. Microtubules are tube shaped and composed of protein that plays a key role in cell division, cytoskeleton, and intercellular material transport and is only 25nm in diameter (1/100,000 thickness of a human hair). Conventionally, cancer treatment focused on disrupting the formation of microtubules to suppress the division of cancer cells. In addition Alzheimer’s is known to be caused by the diminishing of structural integrity of microtubules responsible for intercellular material transport which leads to failure in signal transfer. The research team utilized synchrotron x-ray scattering and transmission electron microscope to analyze the self assemble structure of protein nanotubes to subnanometer accuracy. As a result, the microtubules were found to assemble into 25nm thickness tubules by stacking protein blocks 4 x 5 x 8nm in dimension. In the process, the research team discovered the molecular switch that controls the shape of these protein blocks. In addition the research team was successful in creating a new protein tube structure. Professor Choi commented that they were successful in introducing a new paradigm that suggests the possibility of controlling the complex biological functions of human’s biological system with the simple use of physical principles. He commented further that it is anticipated that the findings will allow for the application of bio nanotubes in engineering and that this is a small step in finding the mechanism behind cancer treatment and neural diseases.
2014.02.03 View 11508
A Molecular Switch Controlling Self-Assembly of Protein Nanotubes Discovered
International collaborative research among South Korea, United States, and Israel research institutionsThe key to the treatment of cancer and brain disease mechanism The molecular switch that controls the self-assembly structure of the protein nanotubes, which plays crucial role in cell division and intracellular transport of materials, has been discovered. KAIST Bio and Brain Engineering Department’s Professor Myeong-Cheol Choi and Professor Chae-Yeon Song conducted the research, in collaboration with the University of California in Santa Barbara, U.S., and Hebrew University in Israel. The findings of the research were published in Nature Materials on the 19th. Microtubules are tube shaped and composed of protein that plays a key role in cell division, cytoskeleton, and intercellular material transport and is only 25nm in diameter (1/100,000 thickness of a human hair). Conventionally, cancer treatment focused on disrupting the formation of microtubules to suppress the division of cancer cells. In addition Alzheimer’s is known to be caused by the diminishing of structural integrity of microtubules responsible for intercellular material transport which leads to failure in signal transfer. The research team utilized synchrotron x-ray scattering and transmission electron microscope to analyze the self assemble structure of protein nanotubes to subnanometer accuracy. As a result, the microtubules were found to assemble into 25nm thickness tubules by stacking protein blocks 4 x 5 x 8nm in dimension. In the process, the research team discovered the molecular switch that controls the shape of these protein blocks. In addition the research team was successful in creating a new protein tube structure. Professor Choi commented that they were successful in introducing a new paradigm that suggests the possibility of controlling the complex biological functions of human’s biological system with the simple use of physical principles. He commented further that it is anticipated that the findings will allow for the application of bio nanotubes in engineering and that this is a small step in finding the mechanism behind cancer treatment and neural diseases.
2014.02.03 View 11508 -
 Space Observatory Video by Science & Technology Satellite No. 3 Released
Images of the Andromeda Galaxy, the Orion Nebula, and the Rosetta Nebula taken by the Science & Technology Satellite No. 3, which was built by the KAIST Satellite Technology Research Center and launched at the Yasny launch site in Russia, were released on December 17, 21 st and 22 nd , 2013.
The Andromeda Galaxy (M31) is the nearest spiral galaxy and is located about two million light years away from the earth. The first image received was an infrared image recorded by the space telescope loaded in the satellite.
Research using the satellite’s infrared camera and imaging spectrometer for observing the Earth will also be conducted until February, 2014. After that, the satellite will be collecting images on infrared cosmic background radiation and exploring the galactic plane at a height of 600 km for two years. The infrared and spectrometer images from the Earth observation can be utilized for disaster monitoring and applied to basic research for the detection of wildfires and urban heat island effect as well as flood damage observation and water quality prediction.
Infrared Light Observed in the Universe, Andromeda Galaxy
2014.01.13 View 9454
Space Observatory Video by Science & Technology Satellite No. 3 Released
Images of the Andromeda Galaxy, the Orion Nebula, and the Rosetta Nebula taken by the Science & Technology Satellite No. 3, which was built by the KAIST Satellite Technology Research Center and launched at the Yasny launch site in Russia, were released on December 17, 21 st and 22 nd , 2013.
The Andromeda Galaxy (M31) is the nearest spiral galaxy and is located about two million light years away from the earth. The first image received was an infrared image recorded by the space telescope loaded in the satellite.
Research using the satellite’s infrared camera and imaging spectrometer for observing the Earth will also be conducted until February, 2014. After that, the satellite will be collecting images on infrared cosmic background radiation and exploring the galactic plane at a height of 600 km for two years. The infrared and spectrometer images from the Earth observation can be utilized for disaster monitoring and applied to basic research for the detection of wildfires and urban heat island effect as well as flood damage observation and water quality prediction.
Infrared Light Observed in the Universe, Andromeda Galaxy
2014.01.13 View 9454 -
 Materials Developed for Sodium Rechargeable Battery by EEWS
The research group of Professor William Goddard III, You-Sung Jung, and Jang-Wook Choi from the Graduate School of Energy, Environment, Water, and Sustainability (EEWS) at KAIST has developed a new sodium-ion rechargeable battery which operates at a high voltage, can be charged, and stably discharges over 10,000 cycles. The research results were published in the online version of the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on December 30, 2013.
Since the material costs of sodium rechargeable batteries is 30 to 40 times lower than lithium batteries, it has received attention as an energy saving tool for smart grids and as the next generation of lithium rechargeable batteries. Until now, sodium-ion rechargeable batteries have had issues with stability when charging and discharging. The research group developed a vanadium-based electrode to solve these problems.
The group said follow-up research will be continued to develop advanced technology on sodium rechargeable batteries as it is still currently in the beginning stages.
The research team: From left to right is Professors William Goddard, You-Sung Jung, and Jang-Wook Choi
2014.01.13 View 12389
Materials Developed for Sodium Rechargeable Battery by EEWS
The research group of Professor William Goddard III, You-Sung Jung, and Jang-Wook Choi from the Graduate School of Energy, Environment, Water, and Sustainability (EEWS) at KAIST has developed a new sodium-ion rechargeable battery which operates at a high voltage, can be charged, and stably discharges over 10,000 cycles. The research results were published in the online version of the Proceedings of the National Academy of Sciences of the United States of America (PNAS) on December 30, 2013.
Since the material costs of sodium rechargeable batteries is 30 to 40 times lower than lithium batteries, it has received attention as an energy saving tool for smart grids and as the next generation of lithium rechargeable batteries. Until now, sodium-ion rechargeable batteries have had issues with stability when charging and discharging. The research group developed a vanadium-based electrode to solve these problems.
The group said follow-up research will be continued to develop advanced technology on sodium rechargeable batteries as it is still currently in the beginning stages.
The research team: From left to right is Professors William Goddard, You-Sung Jung, and Jang-Wook Choi
2014.01.13 View 12389 -
 Mechanism in regulation of cancer-related key enzyme, ATM, for DNA damage and repair revealed
Professor Kwang-Wook Choi
A research team led by Professor Kwang-Wook Choi and Dr. Seong-Tae Hong from the Department of Biological Sciences at KAIST has successfully investigated the operational mechanism of the protein Ataxia Telangiectasia Mutated (ATM), an essential protein to the function of a crucial key enzyme that repairs the damaged DNA which stores biometric information. The results were published on December 19th Nature Communications online edition.
All organisms, including humans, constantly strive to protect the information within their DNA from damages posed by a number of factors, such as carbonized materials in our daily food intake, radioactive materials such as radon emitting from the cement of buildings or ultraviolet of the sunlight, which could be a trigger for cancer.
In order to keep the DNA information safe, the organisms are always carrying out complex and sophisticated DNA repair work, which involves the crucial DNA damage repair protein ATM. Consequently, a faulty ATM leads to higher risks of cancer.
Until now, academia predicted that the Translationally Controlled Tumor Protein (TCTP) will play an important role in regulating the function of ATM. However, since most of main research regarding TCTP has only been conducted in cultured cells, it was unable to identify exactly what mechanisms TCTP employs to control ATM.
The KAIST research team identified that TCTP can combine with ATM or increase the enzymatic activity of ATM. In addition, Drosophilia, one of the most widely used model organisms for molecular genetics, has been used to identify that TCTP and ATM play a very important role in repairing the DNA damaged by radiation. This information has allowed the researchers to establish TCTP’s essential function in maintaining the DNA information in cell cultures and even in higher organisms, and to provide specific and important clues to the regulation of ATM by TCTP.
Professor Kwang-Wook Choi said, “Our research is a good example that basic research using Drosophilia can make important contributions to understanding the process of diseases, such as cancer, and to developing adequate treatment.”
The research has been funded by the Ministry of Science, ICT and Future Planning, Republic of Korea, and the National Research Foundation of Korea.
Figure 1. When the amount of TCTP protein is reduced, cells of the Drosophila's eye are abnormally deformed by radiation. Scale bars = 200mm
Figure 2. When the amount of TCTP protein is reduced, the chromosomes of Drosophilia are easily broken by radiation. Scale bars = 10 mm.
Figure 3. When gene expressions of TCTP and ATM are reduced, large defects occur in the normal development of the eye. (Left: normal Drosophilia's eye, right: development-deficient eye)
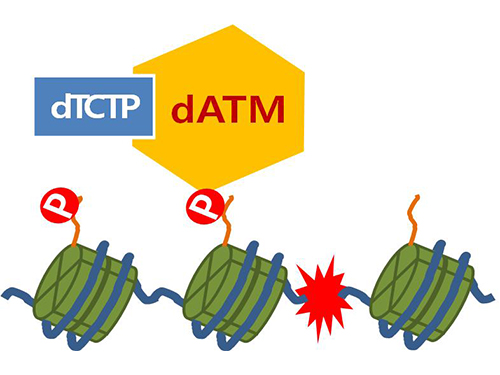
Figure 4. ATM marks the position of the broken DNA, with TCTP helping to facilitate this reaction. DNA (blue line) within the cell nucleus is coiled around the histone protein (green cylinder). When DNA is broken, ATM protein attaches a phosphate group (P). Multiple DNA repair protein recognizes the phosphate as a signal that requires repair and gathers at the site.
2014.01.07 View 15148
Mechanism in regulation of cancer-related key enzyme, ATM, for DNA damage and repair revealed
Professor Kwang-Wook Choi
A research team led by Professor Kwang-Wook Choi and Dr. Seong-Tae Hong from the Department of Biological Sciences at KAIST has successfully investigated the operational mechanism of the protein Ataxia Telangiectasia Mutated (ATM), an essential protein to the function of a crucial key enzyme that repairs the damaged DNA which stores biometric information. The results were published on December 19th Nature Communications online edition.
All organisms, including humans, constantly strive to protect the information within their DNA from damages posed by a number of factors, such as carbonized materials in our daily food intake, radioactive materials such as radon emitting from the cement of buildings or ultraviolet of the sunlight, which could be a trigger for cancer.
In order to keep the DNA information safe, the organisms are always carrying out complex and sophisticated DNA repair work, which involves the crucial DNA damage repair protein ATM. Consequently, a faulty ATM leads to higher risks of cancer.
Until now, academia predicted that the Translationally Controlled Tumor Protein (TCTP) will play an important role in regulating the function of ATM. However, since most of main research regarding TCTP has only been conducted in cultured cells, it was unable to identify exactly what mechanisms TCTP employs to control ATM.
The KAIST research team identified that TCTP can combine with ATM or increase the enzymatic activity of ATM. In addition, Drosophilia, one of the most widely used model organisms for molecular genetics, has been used to identify that TCTP and ATM play a very important role in repairing the DNA damaged by radiation. This information has allowed the researchers to establish TCTP’s essential function in maintaining the DNA information in cell cultures and even in higher organisms, and to provide specific and important clues to the regulation of ATM by TCTP.
Professor Kwang-Wook Choi said, “Our research is a good example that basic research using Drosophilia can make important contributions to understanding the process of diseases, such as cancer, and to developing adequate treatment.”
The research has been funded by the Ministry of Science, ICT and Future Planning, Republic of Korea, and the National Research Foundation of Korea.
Figure 1. When the amount of TCTP protein is reduced, cells of the Drosophila's eye are abnormally deformed by radiation. Scale bars = 200mm
Figure 2. When the amount of TCTP protein is reduced, the chromosomes of Drosophilia are easily broken by radiation. Scale bars = 10 mm.
Figure 3. When gene expressions of TCTP and ATM are reduced, large defects occur in the normal development of the eye. (Left: normal Drosophilia's eye, right: development-deficient eye)
Figure 4. ATM marks the position of the broken DNA, with TCTP helping to facilitate this reaction. DNA (blue line) within the cell nucleus is coiled around the histone protein (green cylinder). When DNA is broken, ATM protein attaches a phosphate group (P). Multiple DNA repair protein recognizes the phosphate as a signal that requires repair and gathers at the site.
2014.01.07 View 15148 -
 Success in Measuring Protein Interaction at the Molecular Level
Professor Tae Young Yoon
- Live observation of two protein interaction in molecular level successful- The limit in measurement and time resolution of immunoprecipitation technique improved by a hundred thousand fold
KAIST Department of Physics Professor Tae Young Yoon’s research team has successfully observed the interaction of two proteins live on molecular level and the findings were published in the October edition of Nature Protocols.
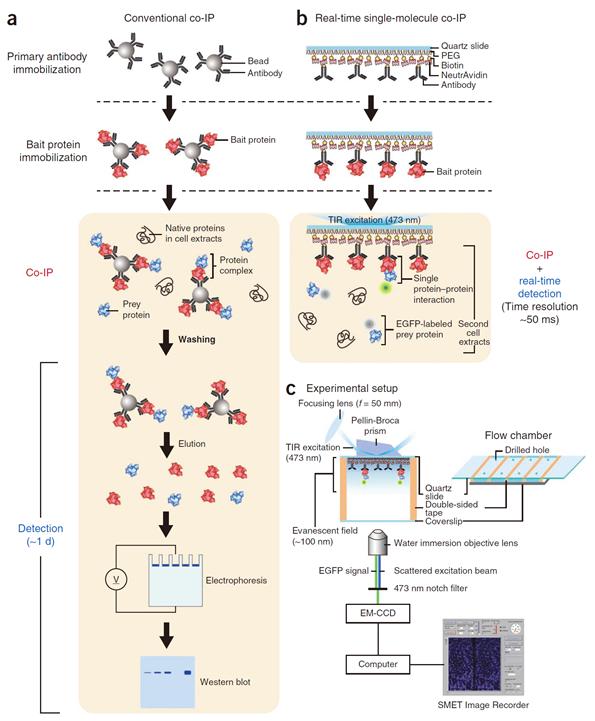
Professor Yoon’s research team developed a fluorescent microscope that can observe a single molecule. The team grafted the immunoprecipitation technique, traditionally used in protein interaction analysis, to the microscope to develop a “live molecular level immunoprecipitation technique”. The team successfully and accurately measured the reaction between two proteins by repeated momentary interactions in the unit of tens of milliseconds.
The existing immunoprecipitation technique required at least one day to detect interaction between two proteins. There were limitations in detecting momentary or weak interactions. Also, quantitative analysis of the results was difficult since the image was measured by protein-band strength. The technique could not be used for live observation.
The team aimed to drastically improve the existing technique and to develop accurate method of measurement on molecular level. The newly developed technology can enable observation of protein interaction within one hour. Also, the interaction can be measured live, thus the protein interaction phenomenon can be measured in depth.
Moreover, every programme used in the experiment was developed and distributed by the research team so source energy is secured and created the foundation for global infra.
Professor Tae Young Yoon said, “The newly developed technology does not require additional protein expression or purification. Hence, a very small sample of protein is enough to accurately analyse protein interaction on a kinetic level.” He continued, “Even cancerous protein from the tissue of a cancer patient can be analysed. Thus a platform for customised anti-cancer medicine in the future has been prepared, as well.”
Figure 1. Mimetic diagram comparing the existing immunoprecipitation technique and the newly developed live molecular level immunoprecipitation technique
2013.12.11 View 8859
Success in Measuring Protein Interaction at the Molecular Level
Professor Tae Young Yoon
- Live observation of two protein interaction in molecular level successful- The limit in measurement and time resolution of immunoprecipitation technique improved by a hundred thousand fold
KAIST Department of Physics Professor Tae Young Yoon’s research team has successfully observed the interaction of two proteins live on molecular level and the findings were published in the October edition of Nature Protocols.
Professor Yoon’s research team developed a fluorescent microscope that can observe a single molecule. The team grafted the immunoprecipitation technique, traditionally used in protein interaction analysis, to the microscope to develop a “live molecular level immunoprecipitation technique”. The team successfully and accurately measured the reaction between two proteins by repeated momentary interactions in the unit of tens of milliseconds.
The existing immunoprecipitation technique required at least one day to detect interaction between two proteins. There were limitations in detecting momentary or weak interactions. Also, quantitative analysis of the results was difficult since the image was measured by protein-band strength. The technique could not be used for live observation.
The team aimed to drastically improve the existing technique and to develop accurate method of measurement on molecular level. The newly developed technology can enable observation of protein interaction within one hour. Also, the interaction can be measured live, thus the protein interaction phenomenon can be measured in depth.
Moreover, every programme used in the experiment was developed and distributed by the research team so source energy is secured and created the foundation for global infra.
Professor Tae Young Yoon said, “The newly developed technology does not require additional protein expression or purification. Hence, a very small sample of protein is enough to accurately analyse protein interaction on a kinetic level.” He continued, “Even cancerous protein from the tissue of a cancer patient can be analysed. Thus a platform for customised anti-cancer medicine in the future has been prepared, as well.”
Figure 1. Mimetic diagram comparing the existing immunoprecipitation technique and the newly developed live molecular level immunoprecipitation technique
2013.12.11 View 8859 -
 The key to Alzheimer disease, PET-MRI made in Korea
Professor Kyu-Sung Cho
- Simultaneous PET-MRI imaging system commercialization technology developed purely from domestic technology - - Inspiring achievement by KAIST, National NanoFab Center, Sogang University, Seoul National University Hospital –
Hopes are high for the potential of producing domestic products in the field of state-of-the-art medical imaging equipment that used to rely on imported products.
The joint research team (KAIST, Sogang University and Seoul National University) with KAIST Department of Nuclear and Quantum Engineering Professor Kyu-Sung Cho in charge, together with National Nanofab Institution (NNFC; Director Jae-Young Lee), has developed PET-MRI simultaneous imaging system with domestic technology only. The team successfully acquired brain images of 3 volunteers with the newly developed system.
PET-MRI is integrated state-of-the-art medical imaging equipment that combines the advantages of Magnetic Resonance Imaging (MRI) that shows anatomical images of the body and Position Emission Tomography (PET) that analyses cell activity and metabolism. Since the anatomical information and functional information can be seen simultaneously, the device can be used to diagnose early onset Alzheimer’s disease and is essential in biological science research, such as new medicine development.
The existing equipment used to take MRI and PET images separately due to the strong magnetic field generated by MRI and combine the images. Hence, it was time consuming and error-prone due to patient’s movement. There was a need to develop PET that functions within a magnetic field to create a simultaneous imaging system.
The newly developed integral PET-MRI has 3 technical characteristics: 1. PET detector without magnetic interference, 2. PET-MRI integration system, 3.PET-MRI imaging processing.
The PET detector is the most important factor and accounts for half the cost of the whole system. KAIST Professor Cho and NNFC Doctor Woo-Suk Seol’s team successfully developed the Silicon Photomultiplier (amplifies light coming into the radiation detector) that can be used in strong magnetic fields. The developed sensor has a global competitive edge since it optimises semiconductor processing to yield over 95% productivity and around 10% gamma radiation energy resolving power.
Sogang University Department and Electrical Engineering Professor Yong Choi developed cutting edge PET system using a new concept of electric charge signal transmission method and imaging location distinction circuit. The creativity and excellence of the research findings were recognised and hence published on the cover of Medical Physics in June.
Seoul National University Hospital Department of Nuclear Medicine Professor Jae-Sung Lee developed the Silicon Photomultiplier sensor based PET imaging reconstitution programme, MRI imaging based PET imaging revision technology and PET-MRI imaging integration software.
Furthermore, KAIST Department of Electrical Engineering Professor Hyun-Wook Park was responsible for the development of RF Shielding technology that enables simultaneous installation of PET and MRI and using this technology, he developed a head coil for the brain that can be connected to PET for installation.
Based on the technology describe above, the joint research team successfully developed PET-MRI system for brains and acquired PET-MRI integrated brain images from 3 volunteers last June.
In particular, this system has the distinct feature of a detachable PET module and MRI head coil to the existing whole body MRI, so that PET-MRI simultaneous imaging is possible with low installation cost.
Professor Cho said, “We have prepared the foundation of domestic commercial PET and the system has a competitive edge in the global market of PET-MRI system technology.” He continued, “It can reduce the cost of the increasing brain related disease diagnosis, including Alzheimer’s, dramatically.”
Funded by Ministry of Trade, Industry and Energy as an Industrial Foundation Technology Development Project (98 billion won in 7 years), the research applied for over 20 patents and 20 CSI theses.
Figure 1.Brain phantom images from developed PET-MRI system
Figure 2. Brain images from developed PET-MRI system
Figure 3. Domestic PET-MRI clinical trial
Figure 4. Head RF coil and PET detector inserted in MRI
Figure 5. Insertion type PET detector module
Figure 6. Silicon Photomultiplier sensor (Left) and flash crystal block (right)
Figure7. Silicon Photomultiplier sensor
Figure 8. PET detection principle
2013.11.28 View 15248
The key to Alzheimer disease, PET-MRI made in Korea
Professor Kyu-Sung Cho
- Simultaneous PET-MRI imaging system commercialization technology developed purely from domestic technology - - Inspiring achievement by KAIST, National NanoFab Center, Sogang University, Seoul National University Hospital –
Hopes are high for the potential of producing domestic products in the field of state-of-the-art medical imaging equipment that used to rely on imported products.
The joint research team (KAIST, Sogang University and Seoul National University) with KAIST Department of Nuclear and Quantum Engineering Professor Kyu-Sung Cho in charge, together with National Nanofab Institution (NNFC; Director Jae-Young Lee), has developed PET-MRI simultaneous imaging system with domestic technology only. The team successfully acquired brain images of 3 volunteers with the newly developed system.
PET-MRI is integrated state-of-the-art medical imaging equipment that combines the advantages of Magnetic Resonance Imaging (MRI) that shows anatomical images of the body and Position Emission Tomography (PET) that analyses cell activity and metabolism. Since the anatomical information and functional information can be seen simultaneously, the device can be used to diagnose early onset Alzheimer’s disease and is essential in biological science research, such as new medicine development.
The existing equipment used to take MRI and PET images separately due to the strong magnetic field generated by MRI and combine the images. Hence, it was time consuming and error-prone due to patient’s movement. There was a need to develop PET that functions within a magnetic field to create a simultaneous imaging system.
The newly developed integral PET-MRI has 3 technical characteristics: 1. PET detector without magnetic interference, 2. PET-MRI integration system, 3.PET-MRI imaging processing.
The PET detector is the most important factor and accounts for half the cost of the whole system. KAIST Professor Cho and NNFC Doctor Woo-Suk Seol’s team successfully developed the Silicon Photomultiplier (amplifies light coming into the radiation detector) that can be used in strong magnetic fields. The developed sensor has a global competitive edge since it optimises semiconductor processing to yield over 95% productivity and around 10% gamma radiation energy resolving power.
Sogang University Department and Electrical Engineering Professor Yong Choi developed cutting edge PET system using a new concept of electric charge signal transmission method and imaging location distinction circuit. The creativity and excellence of the research findings were recognised and hence published on the cover of Medical Physics in June.
Seoul National University Hospital Department of Nuclear Medicine Professor Jae-Sung Lee developed the Silicon Photomultiplier sensor based PET imaging reconstitution programme, MRI imaging based PET imaging revision technology and PET-MRI imaging integration software.
Furthermore, KAIST Department of Electrical Engineering Professor Hyun-Wook Park was responsible for the development of RF Shielding technology that enables simultaneous installation of PET and MRI and using this technology, he developed a head coil for the brain that can be connected to PET for installation.
Based on the technology describe above, the joint research team successfully developed PET-MRI system for brains and acquired PET-MRI integrated brain images from 3 volunteers last June.
In particular, this system has the distinct feature of a detachable PET module and MRI head coil to the existing whole body MRI, so that PET-MRI simultaneous imaging is possible with low installation cost.
Professor Cho said, “We have prepared the foundation of domestic commercial PET and the system has a competitive edge in the global market of PET-MRI system technology.” He continued, “It can reduce the cost of the increasing brain related disease diagnosis, including Alzheimer’s, dramatically.”
Funded by Ministry of Trade, Industry and Energy as an Industrial Foundation Technology Development Project (98 billion won in 7 years), the research applied for over 20 patents and 20 CSI theses.
Figure 1.Brain phantom images from developed PET-MRI system
Figure 2. Brain images from developed PET-MRI system
Figure 3. Domestic PET-MRI clinical trial
Figure 4. Head RF coil and PET detector inserted in MRI
Figure 5. Insertion type PET detector module
Figure 6. Silicon Photomultiplier sensor (Left) and flash crystal block (right)
Figure7. Silicon Photomultiplier sensor
Figure 8. PET detection principle
2013.11.28 View 15248 -
 Green Technology for Data Centers: Ultra-low Power 100 Gbps Ethernet Integrated Circuit Developed
A new integrated circuit (IC), consuming only 0.75W of electricity, will reduce the power usage of data chips installed at data centers by one-third.
Each day, billions of people surf the Internet for information, entertainment, and educational content. The Internet contains an immeasurable amount of information and knowledge generated every minute all around the world that is readily available to everyone with a click of a computer mouse. The real magic of the Internet, however, lies in data centers, where hundreds of billions of data are stored and distributed to designated users around the clock.
Today, almost every business or organization either has its own data centers or outsources data center services to a third party. These centers house highly specialized equipment responsible for the support of computers, networks, data storage, and business security. Accordingly, the operational cost of data centers is tremendous because they consume a large amount of electricity.
Data centers can consume up to 100 times more energy than a standard office building. Data center energy consumption doubled from 2000 to 2006, reaching more than 60 billion kilowatt hours per year. If the current usage and technology trends continue, the energy consumption of data centers in the US will reach 8% of the country’s total electric power consumption by 2020.
A research team at the Korea Advanced Institute of Science and Technology (KAIST) and Terasquare, Inc. (
http://www.terasquare.co.kr
), a spin-off company of the university,
developed an extremely low-powered integrated circuit for Ethernet that consumes less than 0.75W of electricity but is able to send and receive data at the high speed of 100 gigabits per second (Gbps). The research team, headed by Hyeon-Min Bae, assistant professor of electrical engineering at KAIST, claims that the new microchip uses only one-third of the electricity consumed by the currently installed chips at data centers, thereby helping the centers to save energy.
Integrated circuits are embedded on communication modules that are inserted into a line card. Data centers have numerous line cards to build a network including routers and switches. Currently, 8W ICs are the most common in the market, and they consume a lot of energy and require the largest modules (112 cm
2
of CFP), decreasing the port density of line cards and, thus, limiting the amount of data transmission.
The ultra-low-power-circuit, 100-gigabit, full-transceiver CDR, is the world’s first solution that can be loaded to the smallest communication modules (20 cm
2
of CFP4 or 16 cm
2
of
QSFP28), the next-generation chips for data centers. Compared with other chip producers, the 100 Gbps CDR is a greener version of the technology that improves the energy efficiency of data centers while maintaining the high speed of data transmission.
Professor Hyeon-Min Bae said, “When we demonstrate our chip in September of this year at one of the leading companies that manufacture optical communication components and systems, they said that our product is two years ahead of those of our competitors. We plan to produce the chip from 2014 and expect that it will lead the 100 Gbps Ethernet IC market, which is expected to grow to USD 1 billion by 2017.”
The commercial model of the IC was first introduced at the 39
th
European Conference and Exhibition on Optical Communication (ECOC), the largest optical communication forum for new results and developments in Europe, held from September 22-26 at ExCeL London, an international exhibition and convention center.
Professor Bae added, “We received positive responses to our ultra-low-power 100-Gbps Ethernet IC at the ECOC. The chip will be used not only for a particular industry but also for many of next-generation, super-high-speed information communications technologies, such as high-speed USB, high-definition multimedia interface (HDMI), and TV interface.”
Before joining KAIST, Hyeon-Min Bae worked for many years at Finisar as a researcher who designed and developed the world’s first super-high-speed circuit, the 100 Gbps Ethernet IC.
2013.11.25 View 10425
Green Technology for Data Centers: Ultra-low Power 100 Gbps Ethernet Integrated Circuit Developed
A new integrated circuit (IC), consuming only 0.75W of electricity, will reduce the power usage of data chips installed at data centers by one-third.
Each day, billions of people surf the Internet for information, entertainment, and educational content. The Internet contains an immeasurable amount of information and knowledge generated every minute all around the world that is readily available to everyone with a click of a computer mouse. The real magic of the Internet, however, lies in data centers, where hundreds of billions of data are stored and distributed to designated users around the clock.
Today, almost every business or organization either has its own data centers or outsources data center services to a third party. These centers house highly specialized equipment responsible for the support of computers, networks, data storage, and business security. Accordingly, the operational cost of data centers is tremendous because they consume a large amount of electricity.
Data centers can consume up to 100 times more energy than a standard office building. Data center energy consumption doubled from 2000 to 2006, reaching more than 60 billion kilowatt hours per year. If the current usage and technology trends continue, the energy consumption of data centers in the US will reach 8% of the country’s total electric power consumption by 2020.
A research team at the Korea Advanced Institute of Science and Technology (KAIST) and Terasquare, Inc. (
http://www.terasquare.co.kr
), a spin-off company of the university,
developed an extremely low-powered integrated circuit for Ethernet that consumes less than 0.75W of electricity but is able to send and receive data at the high speed of 100 gigabits per second (Gbps). The research team, headed by Hyeon-Min Bae, assistant professor of electrical engineering at KAIST, claims that the new microchip uses only one-third of the electricity consumed by the currently installed chips at data centers, thereby helping the centers to save energy.
Integrated circuits are embedded on communication modules that are inserted into a line card. Data centers have numerous line cards to build a network including routers and switches. Currently, 8W ICs are the most common in the market, and they consume a lot of energy and require the largest modules (112 cm
2
of CFP), decreasing the port density of line cards and, thus, limiting the amount of data transmission.
The ultra-low-power-circuit, 100-gigabit, full-transceiver CDR, is the world’s first solution that can be loaded to the smallest communication modules (20 cm
2
of CFP4 or 16 cm
2
of
QSFP28), the next-generation chips for data centers. Compared with other chip producers, the 100 Gbps CDR is a greener version of the technology that improves the energy efficiency of data centers while maintaining the high speed of data transmission.
Professor Hyeon-Min Bae said, “When we demonstrate our chip in September of this year at one of the leading companies that manufacture optical communication components and systems, they said that our product is two years ahead of those of our competitors. We plan to produce the chip from 2014 and expect that it will lead the 100 Gbps Ethernet IC market, which is expected to grow to USD 1 billion by 2017.”
The commercial model of the IC was first introduced at the 39
th
European Conference and Exhibition on Optical Communication (ECOC), the largest optical communication forum for new results and developments in Europe, held from September 22-26 at ExCeL London, an international exhibition and convention center.
Professor Bae added, “We received positive responses to our ultra-low-power 100-Gbps Ethernet IC at the ECOC. The chip will be used not only for a particular industry but also for many of next-generation, super-high-speed information communications technologies, such as high-speed USB, high-definition multimedia interface (HDMI), and TV interface.”
Before joining KAIST, Hyeon-Min Bae worked for many years at Finisar as a researcher who designed and developed the world’s first super-high-speed circuit, the 100 Gbps Ethernet IC.
2013.11.25 View 10425